-
Citation: S. Nouranian, M.A. Tschopp, S.R. Gwaltney, M.I. Baskes, and M.F. Horstemeyer (2014), "An interatomic potential for saturated hydrocarbons based on the modified embedded-atom method", Physical Chemistry Chemical Physics 16(13), 6233-6249. DOI: 10.1039/c4cp00027g.Abstract: In this work, we developed an interatomic potential for saturated hydrocarbons using the modified embedded-atom method (MEAM), a reactive semi-empirical many-body potential based on density functional theory and pair potentials. We parameterized the potential by fitting to a large experimental and first-principles (FP) database consisting of (1) bond distances, bond angles, and atomization energies at 0 K of a homologous series of alkanes and their select isomers from methane to n-octane, (2) the potential energy curves of H2, CH, and C2 diatomics, (3) the potential energy curves of hydrogen, methane, ethane, and propane dimers, i.e., (H2)2, (CH4)2, (C2H6)2, and (C3H8)2, respectively, and (4) pressure–volume–temperature (PVT) data of a dense high-pressure methane system with the density of 0.5534 g cc−1. We compared the atomization energies and geometries of a range of linear alkanes, cycloalkanes, and free radicals calculated from the MEAM potential to those calculated by other commonly used reactive potentials for hydrocarbons, i.e., second-generation reactive empirical bond order (REBO) and reactive force field (ReaxFF). MEAM reproduced the experimental and/or FP data with accuracy comparable to or better than REBO or ReaxFF. The experimental PVT data for a relatively large series of methane, ethane, propane, and butane systems with different densities were predicted reasonably well by the MEAM potential. Although the MEAM formalism has been applied to atomic systems with predominantly metallic bonding in the past, the current work demonstrates the promising extension of the MEAM potential to covalently bonded molecular systems, specifically saturated hydrocarbons and saturated hydrocarbon-based polymers. The MEAM potential has already been parameterized for a large number of metallic unary, binary, ternary, carbide, nitride, and hydride systems, and extending it to saturated hydrocarbons provides a reliable and transferable potential for atomistic/molecular studies of complex material phenomena involving hydrocarbon–metal or polymer–metal interfaces, polymer–metal nanocomposites, fracture and failure in hydrocarbon-based polymers, etc. The latter is especially true since MEAM is a reactive potential that allows for dynamic bond formation and bond breaking during simulation. Our results show that MEAM predicts the energetics of two major chemical reactions for saturated hydrocarbons, i.e., breaking a C–C and a C–H bond, reasonably well. However, the current parameterization does not accurately reproduce the energetics and structures of unsaturated hydrocarbons and, therefore, should not be applied to such systems.
Notes: Dr. Sasan Nouranian (Center for Advanced Vehicular Systems, Mississippi State Univ.) noted: "These MEAM parameters for elements C and H as well as the diatomic CH are appropriate for energy minimization and reactive molecular dynamics simulations of SATURATED hydrocarbons, where all carbon atoms have the sp3 hybridization (single C-C bonds). At the current state, MEAM cannot handle unsaturated compounds with great accuracy. Furthermore, these C and H parameters are not appropriate for diamond and graphite systems. For the first time, MEAM can be used to simulate hydrocarbons and hydrocarbon/metal systems, since it has a large parameter database for major metals in the periodic table of elements. Since MEAM is a reactive potential, it can also be used to simulate fracture and fatigue in hydrocarbon-based polymers, such as polyethylene and polypropylene and their composites with nanometals as well as polymer/metal interfaces."
Related Models: -
LAMMPS pair_style meam (2014--Nouranian-S--CH--ipr1)Notes: These files were contributed by Sasan Nouranian (Center for Advanced Vehicular Systems, Mississippi State Univ.) on 1 Jul. 2014. An example of energy minimization for an isobutane molecule using the MEAM potential in LAMMPS is also included (Isobutane.in and Isobutane.dat).
File(s):
Implementation Information
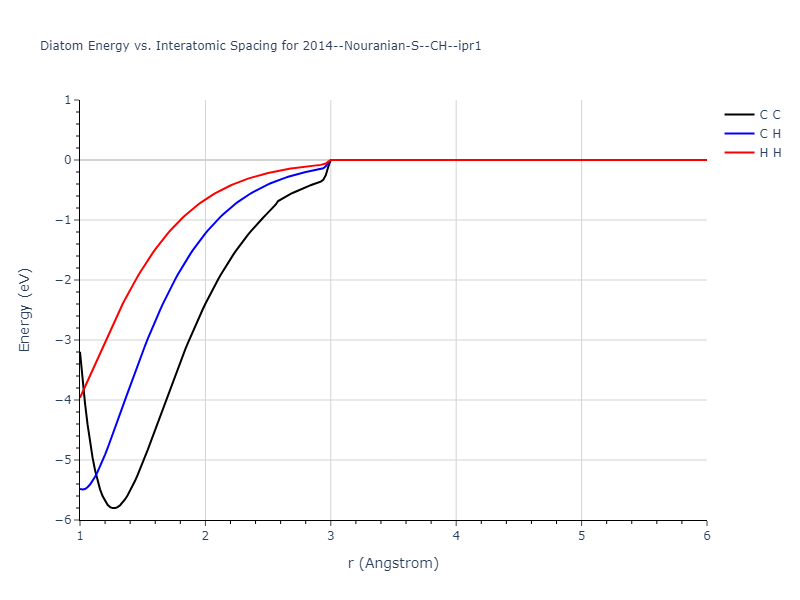
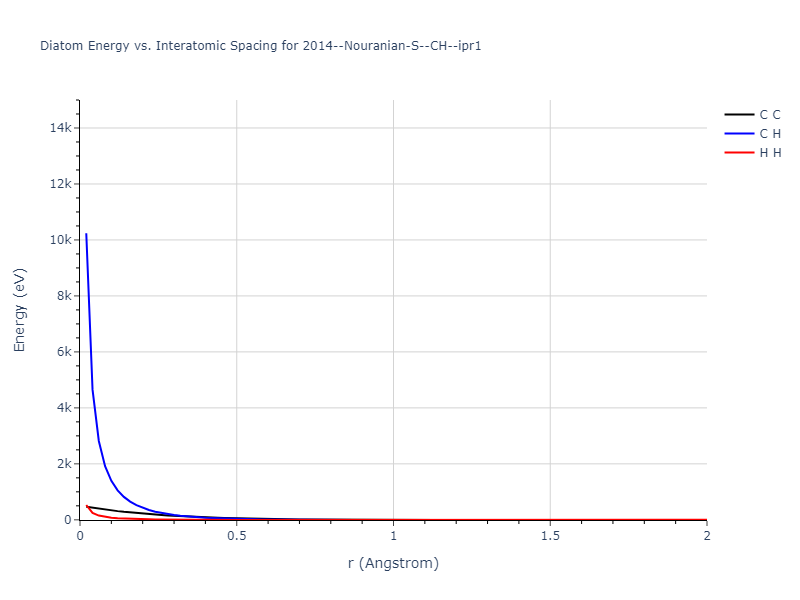
Diatom Energy vs. Interatomic Spacing
Plots of the potential energy vs interatomic spacing, r, are shown below for all diatom sets associated with the interatomic potential. This calculation provides insights into the functional form of the potential's two-body interactions. A system consisting of only two atoms is created, and the potential energy is evaluated for the atoms separated by 0.02 Å <= r <= 6.0> Å in intervals of 0.02 Å. Two plots are shown: one for the "standard" interaction distance range, and one for small values of r. The small r plot is useful for determining whether the potential is suitable for radiation studies.
The calculation method used is available as the iprPy diatom_scan calculation method.
Clicking on the image of a plot will open an interactive version of it in a new tab. The underlying data for the plots can be downloaded by clicking on the links above each plot.
Notes and Disclaimers:
- These values are meant to be guidelines for comparing potentials, not the absolute values for any potential's properties. Values listed here may change if the calculation methods are updated due to improvements/corrections. Variations in the values may occur for variations in calculation methods, simulation software and implementations of the interatomic potentials.
- As this calculation only involves two atoms, it neglects any multi-body interactions that may be important in molecules, liquids and crystals.
- NIST disclaimer
Version Information:
- 2019-11-14. Maximum value range on the shortrange plots are now limited to "expected" levels as details are otherwise lost.
- 2019-08-07. Plots added.
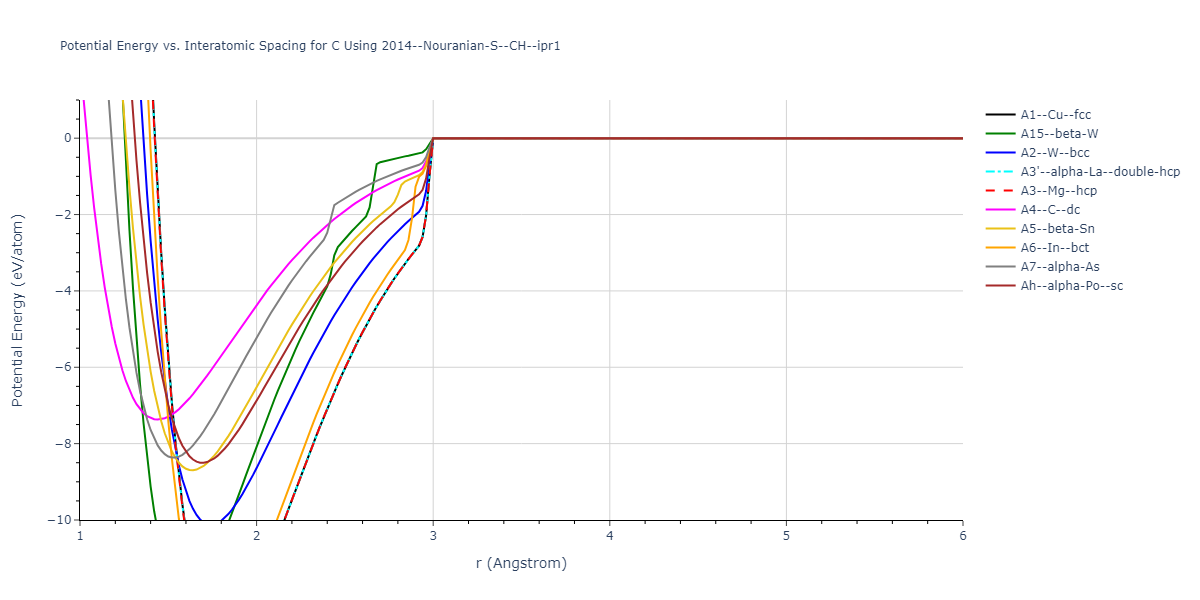
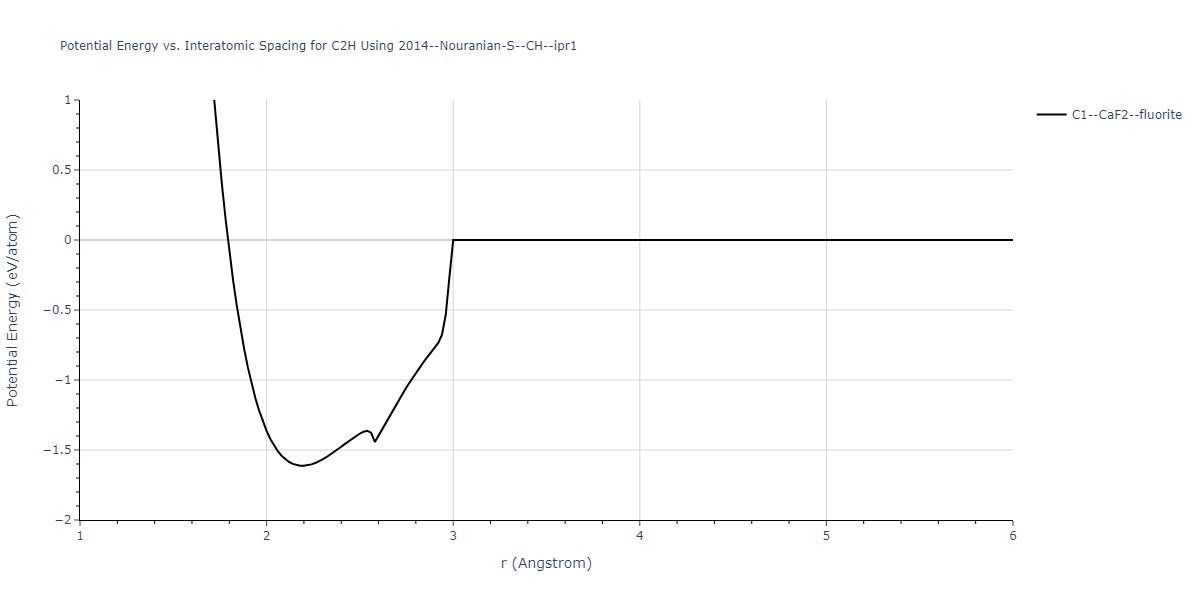
Cohesive Energy vs. Interatomic Spacing
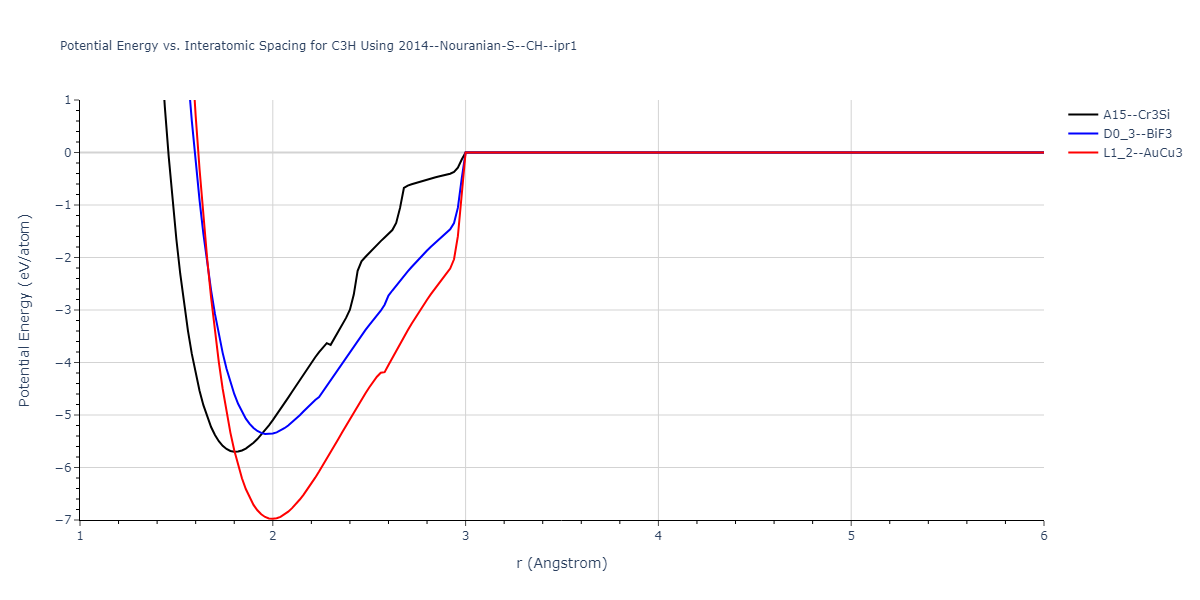
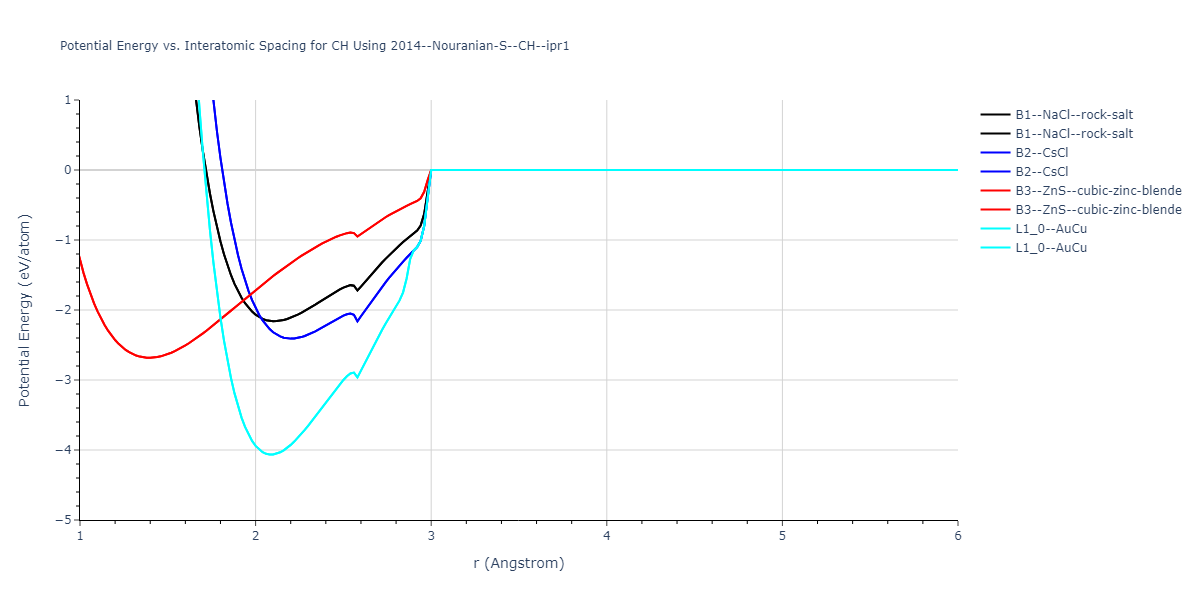
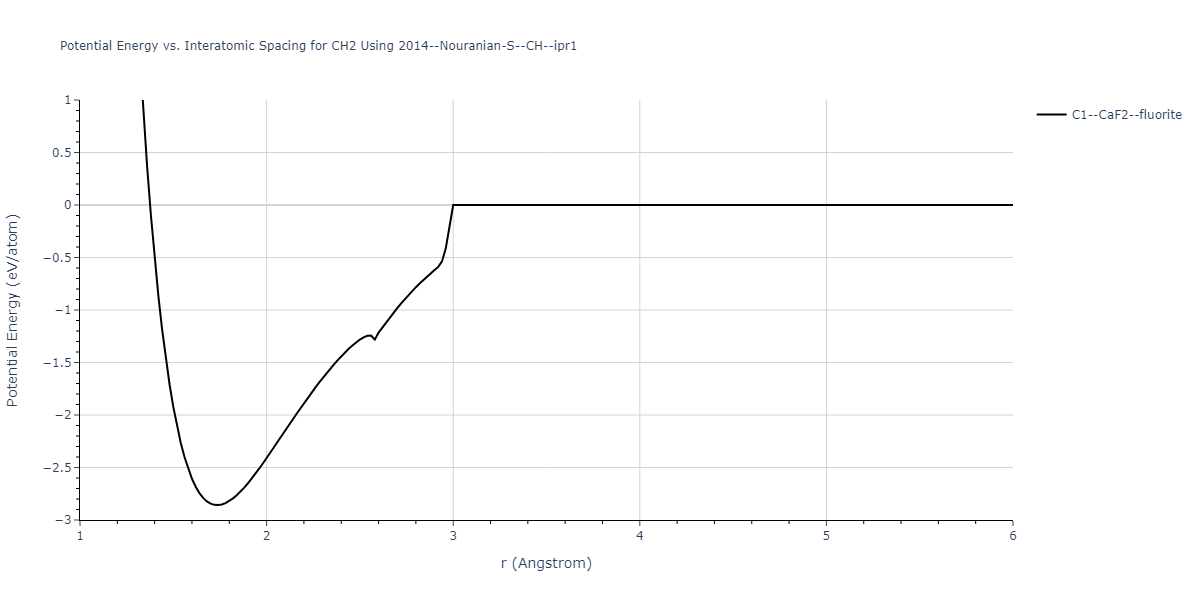
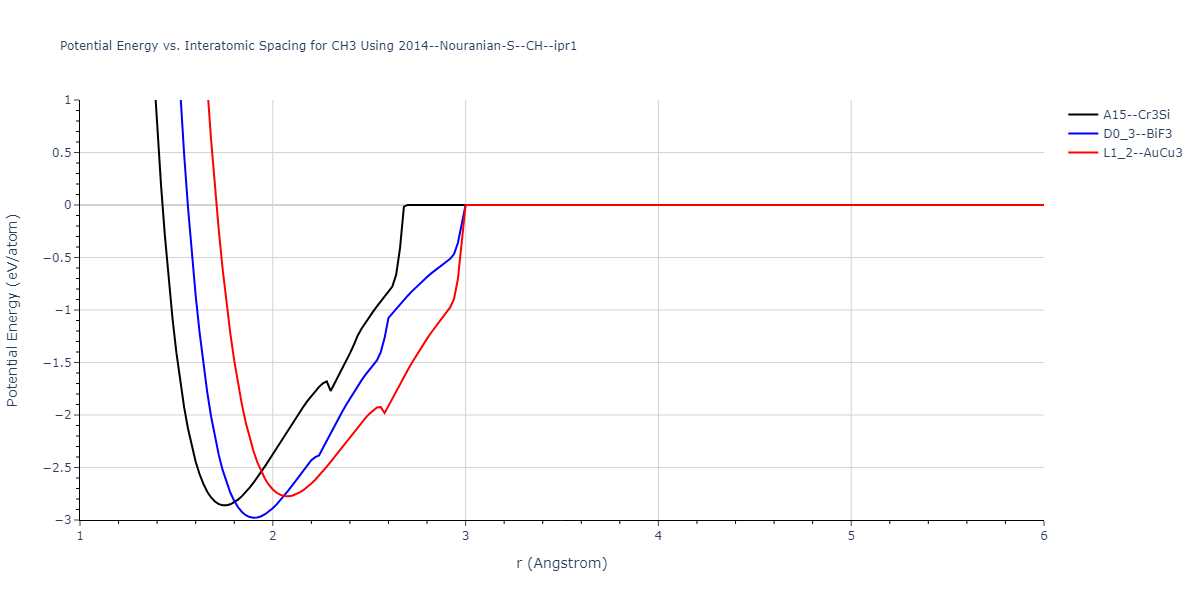
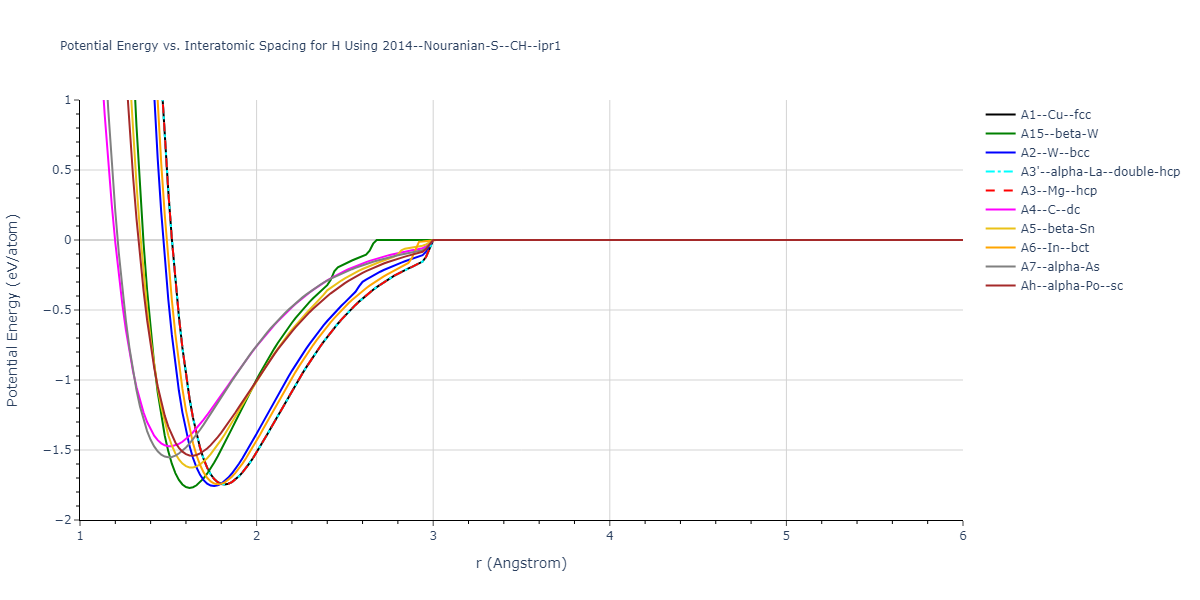
Plots of potential energy vs interatomic spacing, r, are shown below for a number of crystal structures. The structures are generated based on the ideal atomic positions and b/a and c/a lattice parameter ratios for a given crystal prototype. The size of the system is then uniformly scaled, and the energy calculated without relaxing the system. To obtain these plots, values of r are evaluated every 0.02 Å up to 6 Å.
The calculation method used is available as the iprPy E_vs_r_scan calculation method.
Clicking on the image of a plot will open an interactive version of it in a new tab. The underlying data for the plots can be downloaded by clicking on the links above each plot.
Notes and Disclaimers:
- These values are meant to be guidelines for comparing potentials, not the absolute values for any potential's properties. Values listed here may change if the calculation methods are updated due to improvements/corrections. Variations in the values may occur for variations in calculation methods, simulation software and implementations of the interatomic potentials.
- The minima identified by this calculation do not guarantee that the associated crystal structures will be stable since no relaxation is performed.
- NIST disclaimer
Version Information:
- 2020-12-18. Descriptions, tables and plots updated to reflect that the energy values are the measuredper atom potential energy rather than cohesive energy as some potentials have non-zero isolated atom energies.
- 2019-02-04. Values regenerated with even r spacings of 0.02 Å, and now include values less than 2 Å when possible. Updated calculation method and parameters enhance compatibility with more potential styles.
- 2019-04-26. Results for hcp, double hcp, α-As and L10 prototypes regenerated from different unit cell representations. Only α-As results show noticable (>1e-5 eV) difference due to using a different coordinate for Wykoff site c position.
- 2018-06-13. Values for MEAM potentials corrected. Dynamic versions of the plots moved to separate pages to improve page loading. Cosmetic changes to how data is shown and updates to the documentation.
- 2017-01-11. Replaced png pictures with interactive Bokeh plots. Data regenerated with 200 values of r instead of 300.
- 2016-09-28. Plots for binary structures added. Data and plots for elemental structures regenerated. Data values match the values of the previous version. Data table formatting slightly changed to increase precision and ensure spaces between large values. Composition added to plot title and structure names made longer.
- 2016-04-07. Plots for elemental structures added.
Crystal Structure Predictions
Computed lattice constants and cohesive/potential energies are displayed for a variety of crystal structures. The values displayed here are obtained using the following process.
- Initial crystal structure guesses are taken from:
- The iprPy E_vs_r_scan calculation results (shown above) by identifying all energy minima along the measured curves for a given crystal prototype + composition.
- Structures in the Materials Project and OQMD DFT databases.
- All initial guesses are relaxed using three independent methods using a 10x10x10 supercell:
- "box": The system's lattice constants are adjusted to zero pressure without internal relaxations using the iprPy relax_box calculation with a strainrange of 1e-6.
- "static": The system's lattice and atomic positions are statically relaxed using the iprPy relax_static calculation with a minimization force tolerance of 1e-10 eV/Angstrom.
- "dynamic": The system's lattice and atomic positions are dynamically relaxed for 10000 timesteps of 0.01 ps using the iprPy relax_dynamic calculation with an nph integration plus Langevin thermostat. The final configuration is then used as input in running an iprPy relax_static calculation with a minimization force tolerance of 1e-10 eV/Angstrom.
- The relaxed structures obtained from #2 are then evaluated using the spglib package to identify an ideal crystal unit cell based on the results.
- The space group information of the ideal unit cells is compared to the space group information of the corresponding reference structures to identify which structures transformed upon relaxation. The structures that did not transform to a different structure are listed in the table(s) below. The "method" field indicates the most rigorous relaxation method where the structure did not transform. The space group information is also used to match the DFT reference structures to the used prototype, where possible.
- The cohesive energy, Ecoh, is calculated from the measured potential energy per atom, Epot$, by subtracting the isolated energy averaged across all atoms in the unit cell. The isolated atom energies of each species model is obtained either by evaluating a single atom atomic configuration, or by identifying the first energy plateau from the diatom scan calculations for r > 2 Å.
The calculation methods used are implemented into iprPy as the following calculation styles
Notes and Disclaimers:
- These values are meant to be guidelines for comparing potentials, not the absolute values for any potential's properties. Values listed here may change if the calculation methods are updated due to improvements/corrections. Variations in the values may occur for variations in calculation methods, simulation software and implementations of the interatomic potentials.
- The presence of any structures in this list does not guarantee that those structures are stable. Also, the lowest energy structure may not be included in this list.
- Multiple values for the same crystal structure but different lattice constants are possible. This is because multiple energy minima are possible for a given structure and interatomic potential. Having multiple energy minima for a structure does not necessarily make the potential "bad" as unwanted configurations may be unstable or correspond to conditions that may not be relevant to the problem of interest (eg. very high strains).
- NIST disclaimer
Version Information:
- 2025-07-02. All "mp-" reference structure calculations were re-relaxed using the updated Materials Project database rather than the original database structures. Also, a bug was fixed that caused the "static" relaxations to occasionally throw unnecessary errors. This was fixed and all affected calculations were reset and performed again.
- 2022-05-27. The "box" method results have all been redone with an updated methodology more suited for non-orthogonal systems.
- 2020-12-18. Cohesive energies have been corrected by making them relative to the energies of the isolated atoms. The previous cohesive energy values are now listed as the potential energies.
- 2019-06-07. Structures with positive or near zero cohesive energies removed from the display tables. All values still present in the raw data files.
- 2019-04-26. Calculations now computed for each implementation. Results for hcp, double hcp, α-As and L10 prototypes regenerated from different unit cell representations.
- 2018-06-14. Methodology completely changed affecting how the information is displayed. Calculations involving MEAM potentials corrected.
- 2016-09-28. Values for simple compounds added. All identified energy minima for each structure are listed. The existing elemental data was regenerated. Most values are consistent with before, but some differences have been noted. Specifically, variations are seen with some values for potentials where the elastic constants don't vary smoothly near the equilibrium state. Additionally, the inclusion of some high-energy structures has changed based on new criteria for identifying when structures have relaxed to another structure.
- 2016-04-07. Values for elemental crystal structures added. Only values for the global energy minimum of each unique structure given.
Download raw data (including filtered results)
Reference structure matches:
A1--Cu--fcc = oqmd-1214512
A15--beta-W = oqmd-1214957
A2--W--bcc = oqmd-1215135
A3'--alpha-La--double-hcp = oqmd-1215403
A3--Mg--hcp = oqmd-1215313
A4--C--dc = mp-66, oqmd-675640
A5--beta-Sn = oqmd-1215581
A6--In--bct = mp-1181996, oqmd-1215670
A7--alpha-As = oqmd-594528
Ah--alpha-Po--sc = mp-998866, oqmd-636457, oqmd-1214601
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| A3--Mg--hcp | dynamic | -13.0504 | -13.0504 | 1.8116 | 1.8116 | 2.9284 | 90.0 | 90.0 | 120.0 |
| A3'--alpha-La--double-hcp | dynamic | -13.0386 | -13.0386 | 1.8092 | 1.8092 | 5.8801 | 90.0 | 90.0 | 120.0 |
| A1--Cu--fcc | dynamic | -13.0273 | -13.0273 | 2.5555 | 2.5555 | 2.5555 | 90.0 | 90.0 | 90.0 |
| A15--beta-W | dynamic | -11.9826 | -11.9826 | 3.1905 | 3.1905 | 3.1905 | 90.0 | 90.0 | 90.0 |
| mp-1096869 | static | -10.4285 | -10.4285 | 26.4627 | 14.7346 | 1.7324 | 90.0 | 90.1 | 90.0 |
| A2--W--bcc | static | -10.1289 | -10.1289 | 2.0104 | 2.0104 | 2.0104 | 90.0 | 90.0 | 90.0 |
| A3'--alpha-La--double-hcp | static | -9.4794 | -9.4794 | 2.5114 | 2.5114 | 3.4747 | 90.0 | 90.0 | 120.0 |
| A5--beta-Sn | static | -8.6976 | -8.6976 | 3.1531 | 3.1531 | 1.7344 | 90.0 | 90.0 | 90.0 |
| Ah--alpha-Po--sc | static | -8.5041 | -8.5041 | 1.6926 | 1.6926 | 1.6926 | 90.0 | 90.0 | 90.0 |
| mp-24 | box | -7.7149 | -7.7149 | 4.1795 | 4.1795 | 4.1795 | 90.0 | 90.0 | 90.0 |
| mp-611426 | dynamic | -7.6683 | -7.6683 | 2.5224 | 2.5224 | 7.0376 | 90.0 | 90.0 | 120.0 |
| mp-1008374 | box | -7.5298 | -7.5298 | 1.7874 | 9.1119 | 2.3978 | 90.0 | 90.0 | 90.0 |
| oqmd-599492 | dynamic | -7.37 | -7.37 | 2.3511 | 2.3511 | 15.3575 | 90.0 | 90.0 | 120.0 |
| oqmd-599490 | dynamic | -7.37 | -7.37 | 2.3511 | 2.3511 | 7.6788 | 90.0 | 90.0 | 120.0 |
| oqmd-599489 | dynamic | -7.37 | -7.37 | 2.3511 | 2.3511 | 3.8394 | 90.0 | 90.0 | 120.0 |
| A4--C--dc | dynamic | -7.37 | -7.37 | 3.325 | 3.325 | 3.325 | 90.0 | 90.0 | 90.0 |
| oqmd-599493 | dynamic | -7.37 | -7.37 | 2.3511 | 2.3511 | 28.7953 | 90.0 | 90.0 | 120.0 |
| oqmd-599494 | dynamic | -7.37 | -7.37 | 2.3511 | 2.3511 | 40.3135 | 90.0 | 90.0 | 120.0 |
| mp-611448 | dynamic | -7.37 | -7.37 | 2.3511 | 2.3511 | 11.5181 | 90.0 | 90.0 | 120.0 |
| oqmd-599492 | box | -7.3697 | -7.3697 | 2.3506 | 2.3506 | 15.3663 | 90.0 | 90.0 | 120.0 |
| mp-616440 | box | -7.3696 | -7.3696 | 2.3508 | 2.3508 | 15.3632 | 90.0 | 90.0 | 120.0 |
| oqmd-599491 | box | -7.3696 | -7.3696 | 2.3506 | 2.3506 | 11.5242 | 90.0 | 90.0 | 120.0 |
| mp-569567 | box | -7.3695 | -7.3695 | 2.3509 | 2.3509 | 40.3261 | 90.0 | 90.0 | 120.0 |
| mp-611448 | box | -7.3695 | -7.3695 | 2.3508 | 2.3508 | 11.523 | 90.0 | 90.0 | 120.0 |
| oqmd-599493 | box | -7.3695 | -7.3695 | 2.3509 | 2.3509 | 28.8051 | 90.0 | 90.0 | 120.0 |
| mp-569517 | box | -7.3694 | -7.3694 | 2.3507 | 2.3507 | 28.8102 | 90.0 | 90.0 | 120.0 |
| mp-611426 | box | -7.3694 | -7.3694 | 2.3504 | 2.3504 | 7.6846 | 90.0 | 90.0 | 120.0 |
| oqmd-599490 | box | -7.3693 | -7.3693 | 2.3508 | 2.3508 | 7.682 | 90.0 | 90.0 | 120.0 |
| oqmd-599489 | box | -7.3611 | -7.3611 | 2.3897 | 2.3897 | 3.7278 | 90.0 | 90.0 | 120.0 |
| mp-47 | box | -7.3607 | -7.3607 | 2.3903 | 2.3903 | 3.7265 | 90.0 | 90.0 | 120.0 |
| oqmd-595714 | static | -7.3516 | -7.3516 | 5.962 | 9.8491 | 14.3091 | 90.0 | 90.0 | 90.0 |
| mp-1078845 | dynamic | -7.3504 | -7.3504 | 3.9449 | 8.1248 | 2.3955 | 90.0 | 90.0 | 90.0 |
| mp-1078845 | box | -7.347 | -7.347 | 3.9529 | 8.187 | 2.3823 | 90.0 | 90.0 | 90.0 |
| oqmd-637353 | dynamic | -7.337 | -7.337 | 4.0834 | 4.0834 | 2.403 | 90.0 | 90.0 | 90.0 |
| oqmd-637353 | box | -7.337 | -7.337 | 4.0854 | 4.0854 | 2.4013 | 90.0 | 90.0 | 90.0 |
| mp-1008395 | box | -7.337 | -7.337 | 4.0868 | 4.0868 | 2.4001 | 90.0 | 90.0 | 90.0 |
| mp-1190171 | dynamic | -7.3222 | -7.3222 | 8.3483 | 2.4043 | 3.9594 | 90.0 | 90.0 | 90.0 |
| oqmd-637352 | dynamic | -7.3201 | -7.3201 | 8.4597 | 2.4055 | 3.9506 | 90.0 | 97.9 | 90.0 |
| oqmd-637130 | box | -7.3149 | -7.3149 | 8.4462 | 2.3623 | 4.0025 | 90.0 | 90.0 | 90.0 |
| mp-1190171 | box | -7.3147 | -7.3147 | 8.447 | 2.362 | 4.003 | 90.0 | 90.0 | 90.0 |
| oqmd-637352 | box | -7.3107 | -7.3107 | 8.5178 | 2.3651 | 3.9956 | 90.0 | 96.8 | 90.0 |
| mp-1080826 | box | -7.3103 | -7.3103 | 8.5172 | 2.365 | 3.9962 | 90.0 | 96.8 | 90.0 |
| oqmd-610557 | box | -7.2161 | -7.2161 | 4.2179 | 4.8176 | 4.4309 | 90.0 | 90.0 | 90.0 |
| oqmd-610556 | box | -7.2112 | -7.2112 | 4.2244 | 4.8164 | 4.4298 | 90.0 | 90.0 | 90.0 |
| mp-568410 | box | -7.2072 | -7.2072 | 4.228 | 4.8143 | 4.4321 | 90.0 | 90.0 | 90.0 |
| oqmd-585286 | dynamic | -7.1435 | -7.1435 | 4.7701 | 4.7701 | 4.7701 | 90.0 | 90.0 | 90.0 |
| mp-570002 | box | -7.1394 | -7.1394 | 4.7521 | 4.7521 | 4.7521 | 90.0 | 90.0 | 90.0 |
| oqmd-585286 | box | -7.1393 | -7.1393 | 4.752 | 4.752 | 4.752 | 90.0 | 90.0 | 90.0 |
| mp-630227 | static | -7.1033 | -7.1033 | 4.6551 | 10.2817 | 17.4762 | 90.0 | 90.0 | 90.0 |
| mp-1188817 | box | -6.7618 | -6.7618 | 4.9362 | 4.9362 | 4.9362 | 90.0 | 90.0 | 90.0 |
| oqmd-637443 | box | -6.7615 | -6.7615 | 4.9365 | 4.9365 | 4.9365 | 90.0 | 90.0 | 90.0 |
| oqmd-677227 | dynamic | -6.7099 | -6.7099 | 12.0366 | 12.0366 | 12.0366 | 90.0 | 90.0 | 90.0 |
| oqmd-610556 | static | -6.6509 | -6.6509 | 4.7922 | 5.2437 | 3.9901 | 90.0 | 90.0 | 90.0 |
| A6--In--bct | static | -6.638 | -6.638 | 1.6314 | 1.6314 | 6.3803 | 90.0 | 90.0 | 90.0 |
| oqmd-1216026 | static | -6.638 | -6.638 | 2.3072 | 2.3072 | 9.1654 | 90.0 | 93.2 | 90.0 |
| mp-1056957 | static | -6.638 | -6.638 | 2.3072 | 2.3072 | 6.5533 | 90.0 | 90.0 | 90.0 |
| mp-1097832 | static | -6.638 | -6.638 | 2.3072 | 2.3072 | 9.8799 | 90.0 | 90.0 | 90.0 |
| A6--In--bct | static | -6.638 | -6.638 | 1.6314 | 1.6314 | 6.5525 | 90.0 | 90.0 | 90.0 |
| oqmd-684315 | static | -6.5497 | -6.5497 | 7.8344 | 15.0709 | 8.7742 | 90.0 | 90.0 | 90.0 |
| mp-1245190 | static | -6.5493 | -6.5493 | 8.7906 | 8.8019 | 10.2839 | 96.3 | 91.1 | 105.6 |
| mp-1244964 | static | -6.4821 | -6.4821 | 7.883 | 9.8414 | 10.1299 | 83.9 | 85.4 | 83.8 |
| mp-1244913 | static | -6.2818 | -6.2818 | 8.7896 | 9.9474 | 10.0225 | 93.6 | 91.3 | 90.6 |
| oqmd-614417 | box | -6.1917 | -6.1917 | 8.8052 | 8.8052 | 23.8774 | 90.0 | 90.0 | 120.0 |
| mp-680372 | box | -6.1892 | -6.1892 | 8.8173 | 8.8173 | 25.0219 | 90.0 | 90.0 | 120.0 |
| oqmd-684627 | static | -6.1487 | -6.1487 | 8.8046 | 8.8046 | 15.1142 | 90.0 | 90.0 | 90.0 |
| mp-568028 | box | -6.1283 | -6.1283 | 7.8977 | 13.5431 | 9.5834 | 90.0 | 90.0 | 90.0 |
| mp-630227 | box | -6.1052 | -6.1052 | 8.8041 | 8.8672 | 15.11 | 90.0 | 90.0 | 90.0 |
| oqmd-684627 | box | -6.0982 | -6.0982 | 8.825 | 8.825 | 14.8684 | 90.0 | 90.0 | 90.0 |
| mp-1147718 | static | -6.0908 | -6.0908 | 8.732 | 14.3165 | 9.5228 | 90.0 | 90.0 | 90.0 |
| oqmd-677227 | static | -6.0805 | -6.0805 | 13.3651 | 13.3651 | 13.3651 | 90.0 | 90.0 | 90.0 |
| mp-1147718 | dynamic | -6.0578 | -6.0578 | 10.2047 | 14.641 | 8.8708 | 90.0 | 90.0 | 90.0 |
| mp-1196583 | static | -6.0421 | -6.0421 | 13.5588 | 13.5588 | 13.5588 | 90.0 | 90.0 | 90.0 |
| mp-1205283 | box | -6.0412 | -6.0412 | 9.4839 | 12.7945 | 9.1686 | 90.0 | 90.0 | 90.0 |
| oqmd-603085 | box | -6.0257 | -6.0257 | 13.4741 | 13.4741 | 13.4741 | 90.0 | 90.0 | 90.0 |
| mp-1147718 | box | -6.019 | -6.019 | 9.9173 | 14.2745 | 8.949 | 90.0 | 90.0 | 90.0 |
| mp-683919 | box | -6.0092 | -6.0092 | 9.9591 | 16.907 | 17.4367 | 90.0 | 90.0 | 90.0 |
| oqmd-689315 | box | -6.0011 | -6.0011 | 9.8795 | 16.8853 | 17.5366 | 90.0 | 90.0 | 90.0 |
| mp-632329 | dynamic | -5.9666 | -5.9666 | 3.946 | 2.6352 | 3.7369 | 90.0 | 93.7 | 90.0 |
| mp-632329 | static | -5.9666 | -5.9666 | 4.5861 | 2.2661 | 3.7293 | 90.0 | 90.8 | 90.0 |
| mp-169 | dynamic | -5.9666 | -5.9666 | 3.946 | 2.6352 | 3.7294 | 90.0 | 110.1 | 90.0 |
| mp-169 | static | -5.9666 | -5.9666 | 3.946 | 2.6352 | 3.893 | 90.0 | 115.9 | 90.0 |
| mp-937760 | static | -5.9666 | -5.9666 | 2.2661 | 4.5861 | 7.1733 | 90.0 | 90.0 | 90.0 |
| mp-568363 | static | -5.9666 | -5.9666 | 2.6352 | 3.946 | 8.1499 | 90.0 | 90.0 | 90.0 |
| mp-568363 | dynamic | -5.9666 | -5.9666 | 2.2661 | 4.5861 | 8.1499 | 90.0 | 90.0 | 90.0 |
| mp-579909 | static | -5.9666 | -5.9666 | 4.3325 | 5.8541 | 6.5039 | 90.0 | 90.0 | 90.0 |
| oqmd-683820 | static | -5.9666 | -5.9666 | 2.6352 | 3.946 | 6.8141 | 90.0 | 90.0 | 90.0 |
| oqmd-683819 | static | -5.9666 | -5.9666 | 2.6352 | 3.946 | 6.6655 | 90.0 | 90.0 | 90.0 |
| mp-568286 | static | -5.9666 | -5.9666 | 2.6352 | 3.946 | 7.0361 | 90.0 | 90.0 | 90.0 |
| oqmd-610552 | static | -5.9666 | -5.9666 | 4.3325 | 5.854 | 6.1066 | 90.0 | 90.0 | 90.0 |
| mp-579909 | box | -5.9649 | -5.9649 | 4.3559 | 5.6926 | 6.5596 | 90.0 | 90.0 | 90.0 |
| oqmd-610552 | box | -5.9646 | -5.9646 | 4.3679 | 5.6718 | 6.1589 | 90.0 | 90.0 | 90.0 |
| mp-624889 | box | -5.9509 | -5.9509 | 4.6753 | 4.9087 | 5.4352 | 90.0 | 90.0 | 90.0 |
| mp-1196583 | box | -5.9379 | -5.9379 | 13.9544 | 13.9544 | 13.9544 | 90.0 | 90.0 | 90.0 |
| mp-667273 | box | -5.937 | -5.937 | 14.0143 | 14.0143 | 14.0143 | 90.0 | 90.0 | 90.0 |
| oqmd-677227 | box | -5.932 | -5.932 | 13.7438 | 13.7438 | 13.7438 | 90.0 | 90.0 | 90.0 |
| mp-1018088 | static | -5.9273 | -5.9273 | 3.9397 | 3.9397 | 3.9397 | 90.0 | 90.0 | 90.0 |
| oqmd-684315 | box | -5.8259 | -5.8259 | 9.3954 | 14.4953 | 8.8453 | 90.0 | 90.0 | 90.0 |
| mp-1244964 | box | -5.5162 | -5.5162 | 10.2052 | 10.581 | 11.621 | 92.5 | 97.1 | 95.3 |
| mp-1197903 | static | -5.4266 | -5.4266 | 5.3028 | 9.0076 | 19.2811 | 86.5 | 84.4 | 73.8 |
| oqmd-1214868 | static | -5.3379 | -5.3379 | 7.2121 | 7.2121 | 7.2121 | 90.0 | 90.0 | 90.0 |
| mp-1095633 | box | -5.2538 | -5.2538 | 11.4359 | 11.4359 | 4.4502 | 90.0 | 90.0 | 90.0 |
| mp-1196857 | dynamic | -4.9863 | -4.9863 | 9.5768 | 17.6939 | 5.9759 | 90.0 | 90.0 | 90.0 |
| oqmd-1214868 | box | -4.9828 | -4.9828 | 6.8435 | 6.8435 | 6.8435 | 90.0 | 90.0 | 90.0 |
| mp-1182684 | static | -4.9401 | -4.9401 | 10.5338 | 14.4085 | 15.5294 | 90.0 | 90.0 | 90.0 |
| oqmd-1214779 | box | -4.9391 | -4.9391 | 9.245 | 9.245 | 9.245 | 90.0 | 90.0 | 90.0 |
| mp-1197903 | box | -4.9294 | -4.9294 | 8.1201 | 9.1935 | 25.2732 | 89.4 | 83.7 | 73.5 |
| mp-1192619 | box | -4.9236 | -4.9236 | 9.649 | 9.649 | 9.649 | 90.0 | 90.0 | 90.0 |
| mp-1182684 | box | -4.7739 | -4.7739 | 10.7727 | 11.997 | 16.1626 | 90.0 | 90.0 | 90.0 |
| mp-1205417 | box | -4.768 | -4.768 | 12.9953 | 12.9953 | 7.6011 | 90.0 | 90.0 | 90.0 |
| oqmd-1215848 | static | -4.4583 | -4.4583 | 3.4043 | 3.4043 | 1.524 | 90.0 | 90.0 | 120.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-30168 | static | -4.3437 | -4.3437 | 10.5568 | 7.2557 | 10.8754 | 90.0 | 104.0 | 90.0 |
| mp-866659 | box | -4.3365 | -4.3365 | 10.5649 | 7.2598 | 10.8176 | 90.0 | 106.2 | 90.0 |
| mp-30168 | box | -4.3308 | -4.3308 | 11.0407 | 7.6927 | 11.2829 | 90.0 | 104.9 | 90.0 |
| oqmd-10260 | box | -4.327 | -4.327 | 9.7617 | 6.9635 | 10.5193 | 90.0 | 105.2 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-603334 | static | -4.9578 | -4.9578 | 10.6249 | 4.6089 | 14.5809 | 90.0 | 108.9 | 90.0 |
| oqmd-20106 | static | -4.9437 | -4.9437 | 9.5508 | 5.0172 | 14.8506 | 90.0 | 110.9 | 90.0 |
| mp-995184 | box | -4.8769 | -4.8769 | 12.5827 | 2.5367 | 3.2738 | 90.0 | 98.5 | 90.0 |
| mp-603334 | box | -4.8731 | -4.8731 | 10.211 | 4.7315 | 16.3925 | 90.0 | 111.0 | 90.0 |
| oqmd-20106 | box | -4.8448 | -4.8448 | 9.4595 | 5.3978 | 13.5653 | 90.0 | 108.5 | 90.0 |
| C1--CaF2--fluorite | static | -1.6131 | -1.6131 | 5.0596 | 5.0596 | 5.0596 | 90.0 | 90.0 | 90.0 |
| C1--CaF2--fluorite | static | -1.4609 | -1.4609 | 5.9393 | 5.9393 | 5.9393 | 90.0 | 90.0 | 90.0 |
| C1--CaF2--fluorite | box | -1.3635 | -1.3635 | 5.8664 | 5.8664 | 5.8664 | 90.0 | 90.0 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
D0_3--BiF3 = oqmd-313359
L1_2--AuCu3 = oqmd-346564
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| L1_2--AuCu3 | static | -6.9783 | -6.9783 | 2.8254 | 2.8254 | 2.8254 | 90.0 | 90.0 | 90.0 |
| A15--Cr3Si | static | -5.7006 | -5.7006 | 3.6113 | 3.6113 | 3.6113 | 90.0 | 90.0 | 90.0 |
| D0_3--BiF3 | static | -5.3639 | -5.3639 | 4.5606 | 4.5606 | 4.5606 | 90.0 | 90.0 | 90.0 |
| oqmd-302817 | static | -5.1371 | -5.1371 | 2.8774 | 2.8774 | 8.524 | 90.0 | 90.0 | 90.0 |
| A15--Cr3Si | static | -3.6643 | -3.6643 | 4.5998 | 4.5998 | 4.5998 | 90.0 | 90.0 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-995217 | static | -4.7299 | -4.7299 | 9.7565 | 9.552 | 4.7293 | 90.0 | 114.3 | 90.0 |
| mp-995217 | box | -4.6638 | -4.6638 | 9.2697 | 11.0734 | 4.8743 | 90.0 | 109.3 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-1195106 | static | -4.4734 | -4.4734 | 6.5196 | 6.5196 | 9.1534 | 90.0 | 90.0 | 90.0 |
| mp-1195106 | dynamic | -4.4734 | -4.4734 | 6.5199 | 6.5199 | 9.1522 | 90.0 | 90.0 | 90.0 |
| mp-1195106 | box | -4.4696 | -4.4696 | 6.7802 | 6.7802 | 9.261 | 90.0 | 90.0 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-995202 | static | -5.5118 | -5.5118 | 2.5145 | 3.9304 | 12.9453 | 90.2 | 90.6 | 105.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-1097016 | static | -4.8442 | -4.8442 | 9.4075 | 11.0006 | 14.134 | 90.0 | 95.2 | 90.0 |
| mp-1097016 | box | -4.7777 | -4.7777 | 8.9444 | 11.5131 | 15.1784 | 90.0 | 92.7 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
B1--NaCl--rock-salt = oqmd-1106288
B2--CsCl = oqmd-307103
L1_0--AuCu = oqmd-338645
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| L1_0--AuCu | static | -5.4861 | -5.4861 | 1.6575 | 1.6575 | 4.5787 | 90.0 | 90.0 | 90.0 |
| L1_0--AuCu | static | -5.3194 | -5.3194 | 1.6842 | 1.6842 | 4.0482 | 90.0 | 90.0 | 90.0 |
| mp-1096986 | dynamic | -4.9868 | -4.9868 | 2.4613 | 2.4613 | 4.6056 | 90.0 | 90.0 | 120.0 |
| mp-1096986 | box | -4.9766 | -4.9766 | 2.4706 | 2.4706 | 4.9052 | 90.0 | 90.0 | 120.0 |
| mp-1079612 | static | -4.6009 | -4.6009 | 4.5999 | 4.5999 | 4.5999 | 90.0 | 90.0 | 90.0 |
| mp-995197 | static | -4.5789 | -4.5789 | 5.3171 | 9.6824 | 5.6997 | 90.0 | 110.0 | 90.0 |
| mp-1079612 | box | -4.5511 | -4.5511 | 4.4961 | 4.4961 | 4.4961 | 90.0 | 90.0 | 90.0 |
| mp-995197 | box | -4.5048 | -4.5048 | 5.8895 | 10.5745 | 4.8975 | 90.0 | 105.1 | 90.0 |
| L1_0--AuCu | box | -4.2797 | -4.2797 | 3.0688 | 3.0688 | 1.5821 | 90.0 | 90.0 | 90.0 |
| B3--ZnS--cubic-zinc-blende | box | -2.6829 | -2.6829 | 3.2216 | 3.2216 | 3.2216 | 90.0 | 90.0 | 90.0 |
| B2--CsCl | static | -2.4089 | -2.4089 | 2.5422 | 2.5422 | 2.5422 | 90.0 | 90.0 | 90.0 |
| B2--CsCl | static | -2.1919 | -2.1919 | 2.9696 | 2.9696 | 2.9696 | 90.0 | 90.0 | 90.0 |
| oqmd-1228602 | box | -2.1743 | -2.1743 | 2.4258 | 2.4258 | 3.1517 | 90.0 | 90.0 | 120.0 |
| B1--NaCl--rock-salt | static | -2.1602 | -2.1602 | 4.1975 | 4.1975 | 4.1975 | 90.0 | 90.0 | 90.0 |
| B2--CsCl | box | -2.0509 | -2.0509 | 2.9342 | 2.9342 | 2.9342 | 90.0 | 90.0 | 90.0 |
| B1--NaCl--rock-salt | static | -1.7447 | -1.7447 | 5.1435 | 5.1435 | 5.1435 | 90.0 | 90.0 | 90.0 |
| B1--NaCl--rock-salt | box | -1.6438 | -1.6438 | 5.0923 | 5.0923 | 5.0923 | 90.0 | 90.0 | 90.0 |
| B4--ZnS--wurtzite | static | -0.9641 | -0.9641 | 4.1917 | 4.1917 | 6.8839 | 90.0 | 90.0 | 120.0 |
| B4--ZnS--wurtzite | static | -0.9641 | -0.9641 | 4.1921 | 4.1921 | 6.8826 | 90.0 | 90.0 | 120.0 |
| B3--ZnS--cubic-zinc-blende | box | -0.8943 | -0.8943 | 5.8758 | 5.8758 | 5.8758 | 90.0 | 90.0 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-985782 | dynamic | -4.2508 | -4.2508 | 7.0594 | 2.5392 | 4.6922 | 90.0 | 90.0 | 90.0 |
| mp-985782 | box | -4.2475 | -4.2475 | 7.1496 | 2.5558 | 4.9179 | 90.0 | 90.0 | 90.0 |
| C1--CaF2--fluorite | static | -2.8583 | -2.8583 | 4.0071 | 4.0071 | 4.0071 | 90.0 | 90.0 | 90.0 |
| C1--CaF2--fluorite | static | -1.3133 | -1.3133 | 5.9388 | 5.9388 | 5.9388 | 90.0 | 90.0 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
D0_3--BiF3 = oqmd-311536
L1_2--AuCu3 = oqmd-348387
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| D0_3--BiF3 | static | -2.979 | -2.979 | 4.3964 | 4.3964 | 4.3964 | 90.0 | 90.0 | 90.0 |
| A15--Cr3Si | static | -2.8614 | -2.8614 | 3.5 | 3.5 | 3.5 | 90.0 | 90.0 | 90.0 |
| oqmd-323901 | box | -2.8055 | -2.8055 | 4.1767 | 4.1767 | 3.1395 | 90.0 | 90.0 | 120.0 |
| L1_2--AuCu3 | static | -2.7743 | -2.7743 | 2.9348 | 2.9348 | 2.9348 | 90.0 | 90.0 | 90.0 |
| L1_2--AuCu3 | static | -2.0092 | -2.0092 | 3.6369 | 3.6369 | 3.6369 | 90.0 | 90.0 | 90.0 |
| A15--Cr3Si | static | -1.7715 | -1.7715 | 4.6004 | 4.6004 | 4.6004 | 90.0 | 90.0 | 90.0 |
| A15--Cr3Si | static | -0.0 | -0.0 | 6.0 | 6.0 | 6.0 | 90.0 | 90.0 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| mp-1021328 | static | -3.6892 | -3.6892 | 4.4338 | 4.4338 | 4.4338 | 90.0 | 90.0 | 90.0 |
| mp-1021328 | box | -3.6639 | -3.6639 | 4.9589 | 4.9589 | 4.9589 | 90.0 | 90.0 | 90.0 |
Download raw data (including filtered results)
Reference structure matches:
A1--Cu--fcc = mp-634659, oqmd-5497, oqmd-5499, oqmd-1214529
A15--beta-W = oqmd-1214974
A2--W--bcc = mp-632250, oqmd-5517, oqmd-1215152
A3'--alpha-La--double-hcp = oqmd-1215420
A3--Mg--hcp = mp-23907, mp-570752, oqmd-40867, oqmd-676625, oqmd-1215330
A4--C--dc = oqmd-1215509
A5--beta-Sn = oqmd-1215598
A6--In--bct = oqmd-1215687
A7--alpha-As = oqmd-1215776
| prototype | method | Ecoh (eV/atom) | Epot (eV/atom) | a0 (Å) | b0 (Å) | c0 (Å) | α (degrees) | β (degrees) | γ (degrees) |
|---|---|---|---|---|---|---|---|---|---|
| oqmd-1215063 | static | -2.3875 | -2.3875 | 4.5474 | 4.4615 | 3.5002 | 90.0 | 90.0 | 90.0 |
| oqmd-5539 | box | -2.386 | -2.386 | 3.8858 | 3.8858 | 2.8007 | 90.0 | 90.0 | 90.0 |
| mp-850274 | dynamic | -2.3851 | -2.3851 | 4.3181 | 4.3181 | 2.9326 | 90.0 | 90.0 | 90.0 |
| mp-632172 | dynamic | -2.3851 | -2.3851 | 4.4568 | 4.4568 | 2.9326 | 90.0 | 90.0 | 90.0 |
| oqmd-1214707 | box | -2.3798 | -2.3798 | 4.1591 | 4.1535 | 4.2112 | 90.0 | 90.0 | 90.0 |
| mp-1188177 | box | -2.3788 | -2.3788 | 5.6388 | 6.0404 | 5.7114 | 90.0 | 90.0 | 90.0 |
| mp-632291 | box | -2.3788 | -2.3788 | 2.9459 | 2.9459 | 4.1728 | 90.0 | 90.0 | 90.0 |
| oqmd-11544 | box | -2.3727 | -2.3727 | 3.9755 | 3.9755 | 3.9526 | 90.0 | 90.0 | 120.0 |
| oqmd-1215063 | box | -2.3698 | -2.3698 | 4.8238 | 4.231 | 3.1962 | 90.0 | 90.0 | 90.0 |
| mp-1181265 | static | -2.3661 | -2.3661 | 5.3174 | 7.4234 | 10.0682 | 77.9 | 78.8 | 73.8 |
| oqmd-1214796 | static | -2.2146 | -2.2146 | 7.8101 | 7.8101 | 7.8101 | 90.0 | 90.0 | 90.0 |
| oqmd-1214796 | box | -2.1745 | -2.1745 | 7.2435 | 7.2435 | 7.2435 | 90.0 | 90.0 | 90.0 |
| A15--beta-W | static | -1.7705 | -1.7705 | 3.2417 | 3.2417 | 3.2417 | 90.0 | 90.0 | 90.0 |
| A2--W--bcc | dynamic | -1.7577 | -1.7577 | 2.0303 | 2.0303 | 2.0303 | 90.0 | 90.0 | 90.0 |
| oqmd-82345 | box | -1.7577 | -1.7577 | 2.0303 | 2.8715 | 2.8715 | 90.0 | 90.0 | 90.0 |
| A3--Mg--hcp | dynamic | -1.7459 | -1.7459 | 1.8175 | 1.8175 | 2.968 | 90.0 | 90.0 | 120.0 |
| A3'--alpha-La--double-hcp | dynamic | -1.7459 | -1.7459 | 1.8175 | 1.8175 | 5.936 | 90.0 | 90.0 | 120.0 |
| A1--Cu--fcc | dynamic | -1.7459 | -1.7459 | 2.5704 | 2.5704 | 2.5704 | 90.0 | 90.0 | 90.0 |
| oqmd-57238 | box | -1.7449 | -1.7449 | 2.6755 | 2.3669 | 2.6753 | 90.0 | 90.0 | 90.0 |
| A5--beta-Sn | static | -1.6306 | -1.6306 | 3.0359 | 3.0359 | 1.8646 | 90.0 | 90.0 | 90.0 |
| oqmd-1215865 | box | -1.5664 | -1.5664 | 2.1611 | 2.1611 | 1.1922 | 90.0 | 90.0 | 120.0 |
| A3--Mg--hcp | box | -1.5599 | -1.5599 | 3.7564 | 3.7564 | 1.1561 | 90.0 | 90.0 | 120.0 |
| A5--beta-Sn | box | -1.556 | -1.556 | 4.4157 | 4.4157 | 1.1514 | 90.0 | 90.0 | 90.0 |
| A7--alpha-As | box | -1.5489 | -1.5489 | 2.3076 | 2.3076 | 5.8675 | 90.0 | 90.0 | 120.0 |
| mp-1096977 | box | -1.5434 | -1.5434 | 2.2991 | 2.2991 | 1.1315 | 90.0 | 90.0 | 90.0 |
| Ah--alpha-Po--sc | static | -1.5408 | -1.5408 | 1.6348 | 1.6348 | 1.6348 | 90.0 | 90.0 | 90.0 |
| oqmd-1215954 | box | -1.5404 | -1.5404 | 2.556 | 2.556 | 3.326 | 90.0 | 90.0 | 120.0 |
| A4--C--dc | static | -1.474 | -1.474 | 3.4877 | 3.4877 | 3.4877 | 90.0 | 90.0 | 90.0 |
| A15--beta-W | static | -0.0 | -0.0 | 6.0 | 6.0 | 6.0 | 90.0 | 90.0 | 90.0 |
| A2--W--bcc | dynamic | 0.0 | 0.0 | 5.21 | 5.21 | 5.21 | 90.0 | 90.0 | 90.0 |
| A3--Mg--hcp | dynamic | 0.0 | 0.0 | 3.9004 | 3.9004 | 6.3557 | 90.0 | 90.0 | 120.0 |
| A1--Cu--fcc | dynamic | 0.0 | 0.0 | 5.4286 | 5.4286 | 5.4286 | 90.0 | 90.0 | 90.0 |
| A3--Mg--hcp | dynamic | 0.0 | 0.0 | 5.1702 | 5.1702 | 3.8495 | 90.0 | 90.0 | 120.0 |
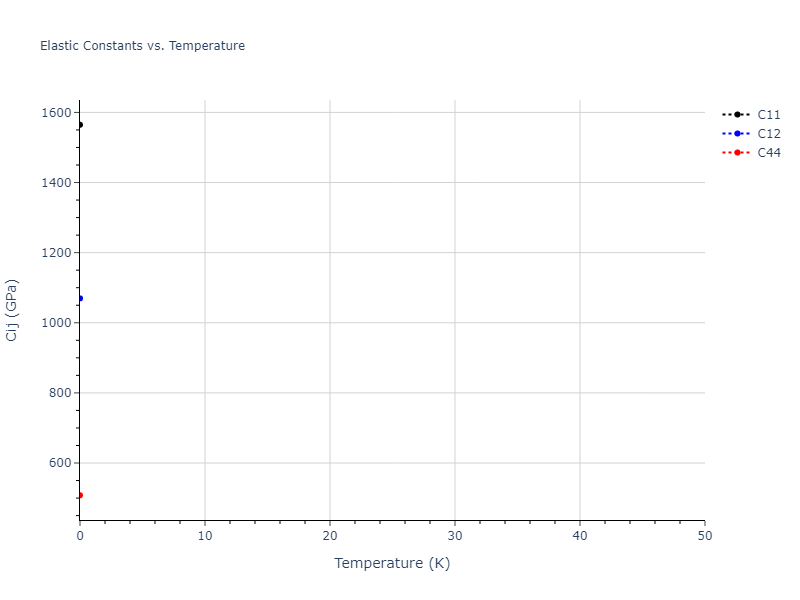
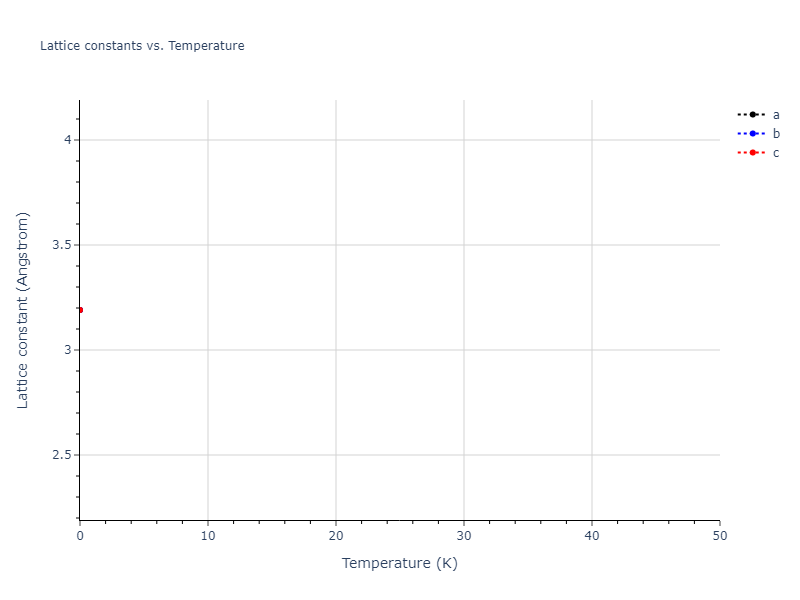
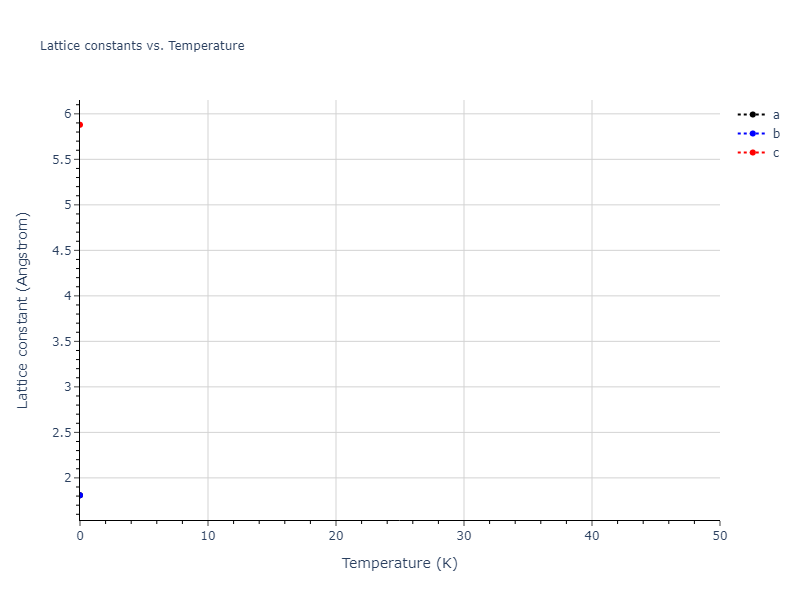
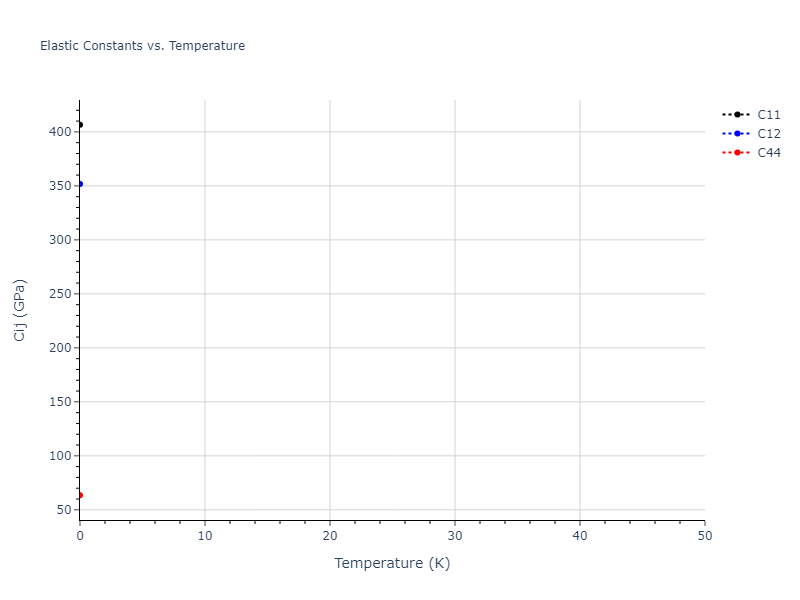
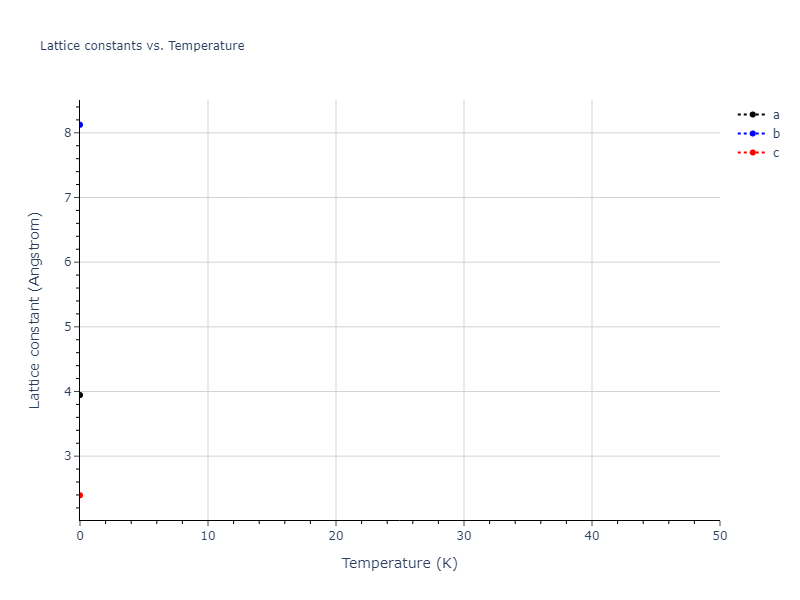
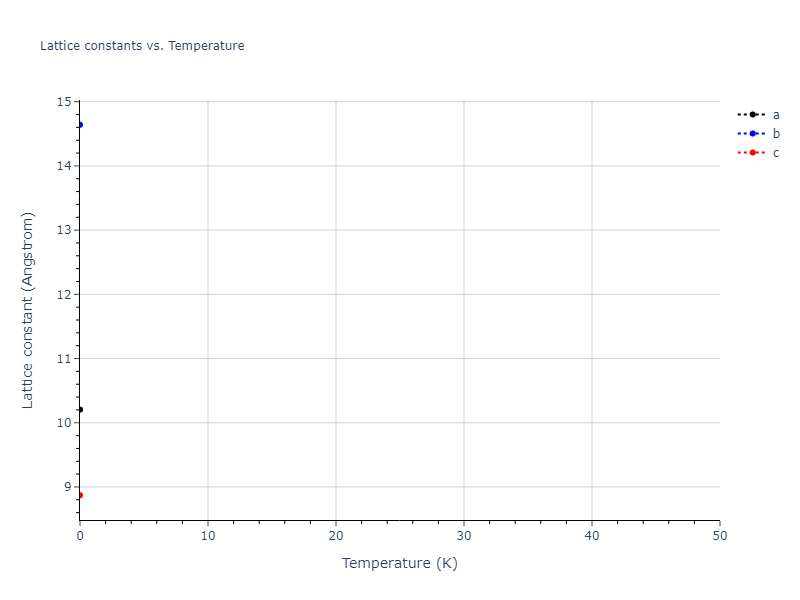
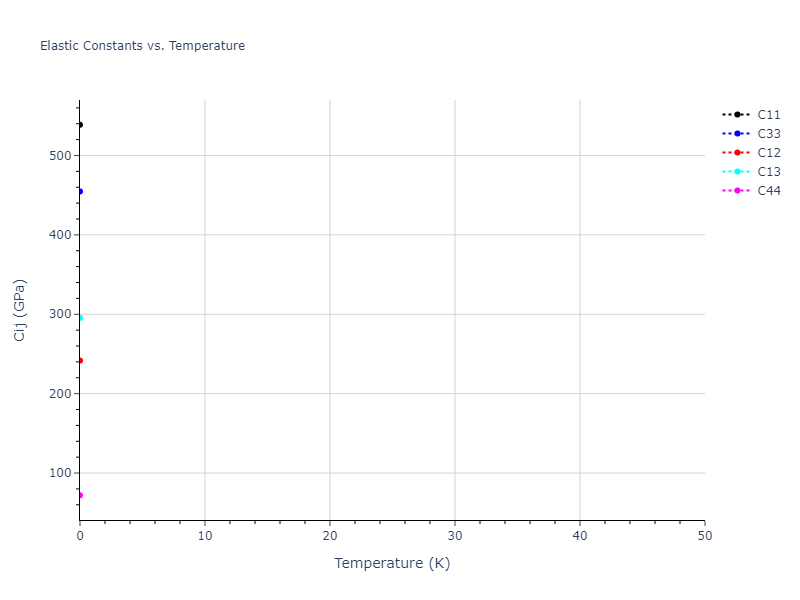
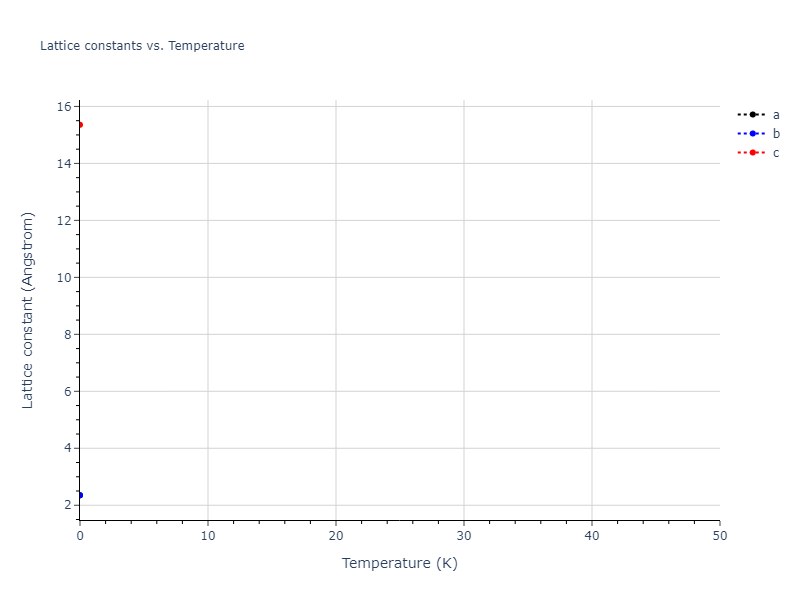
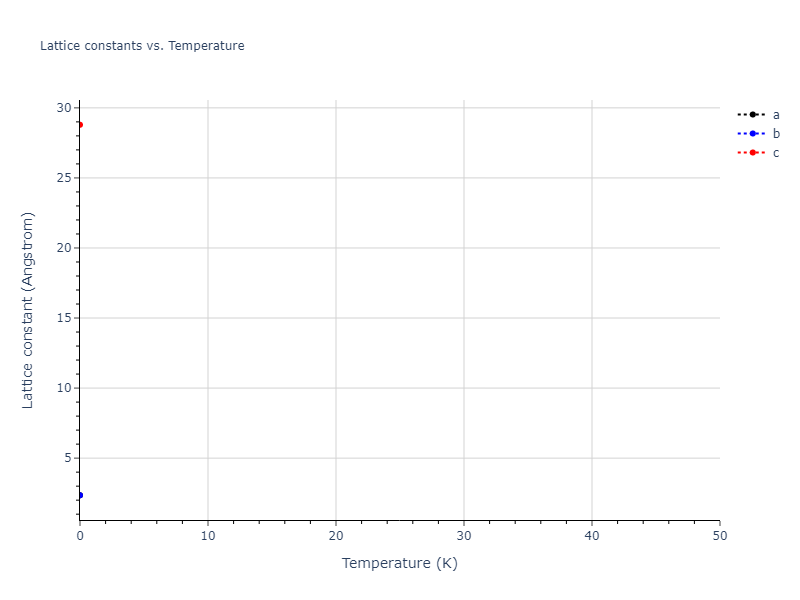
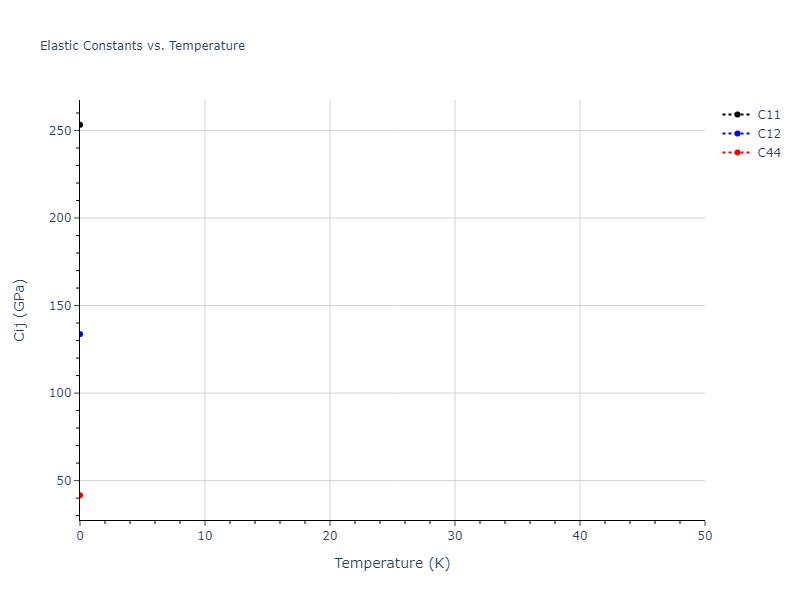
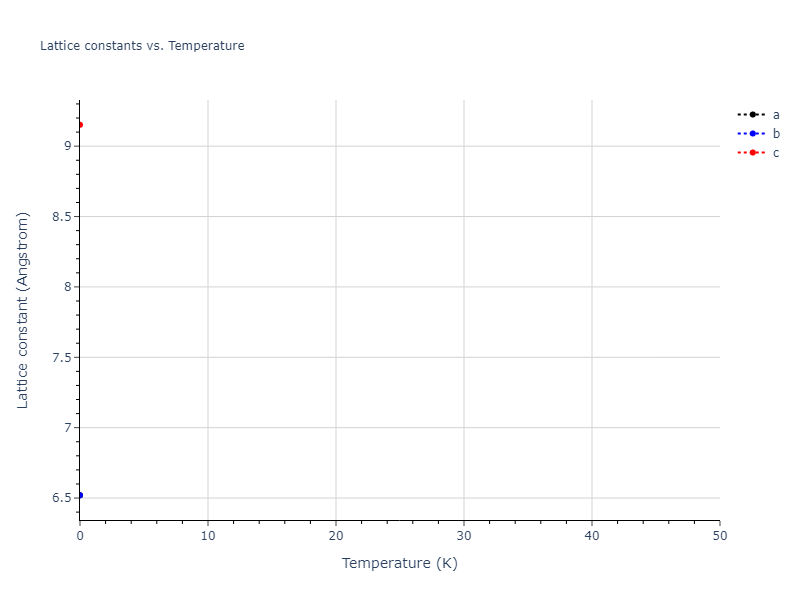
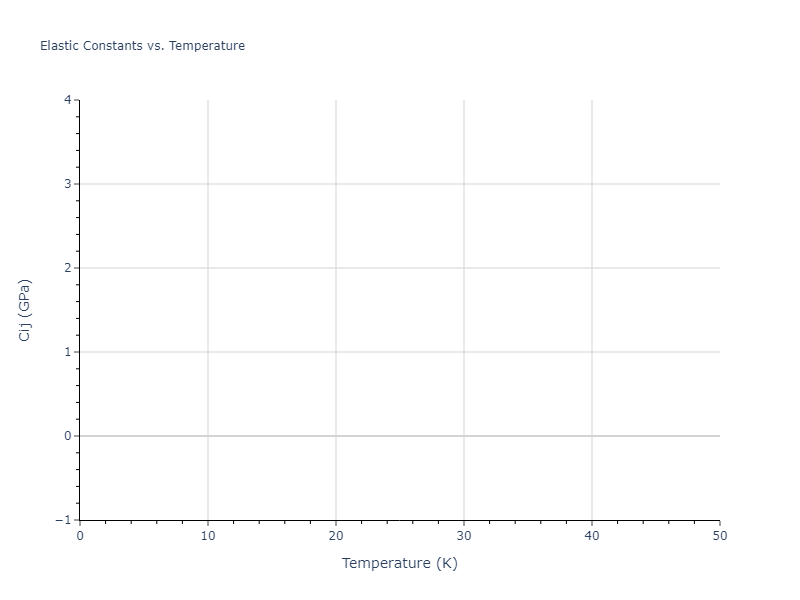
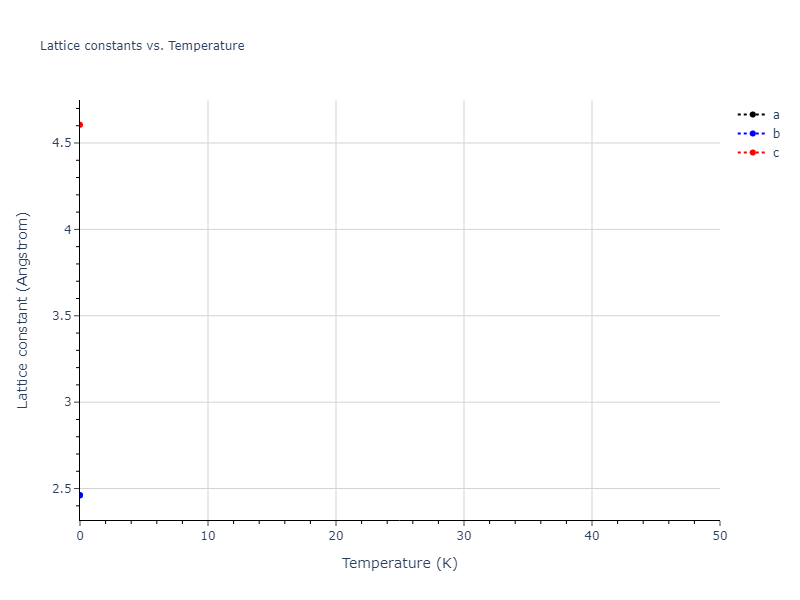
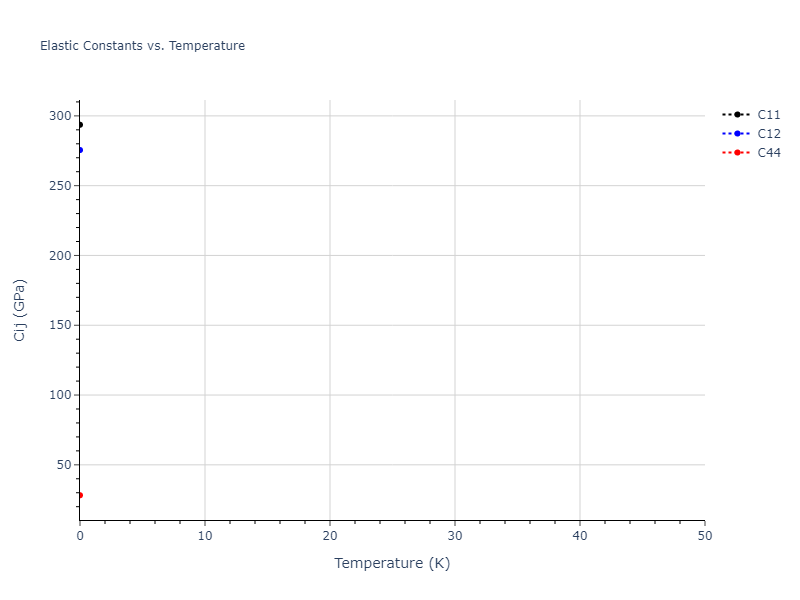
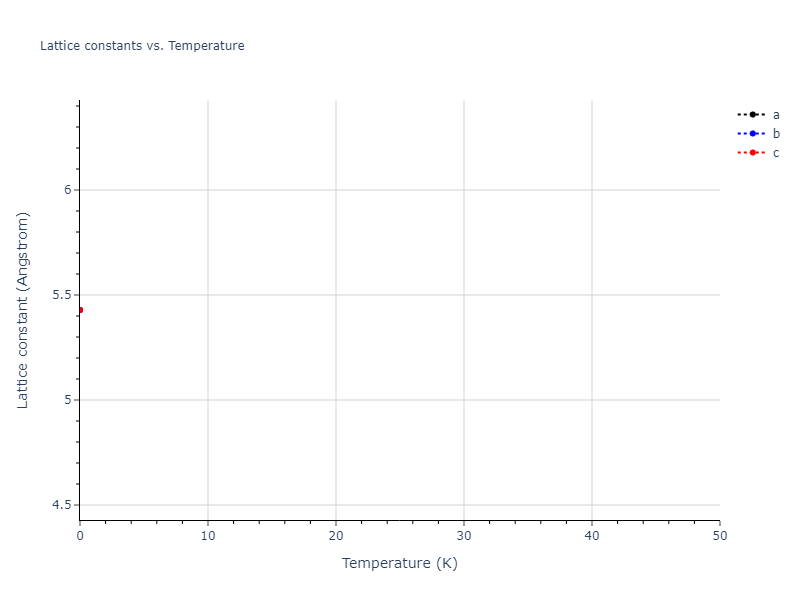
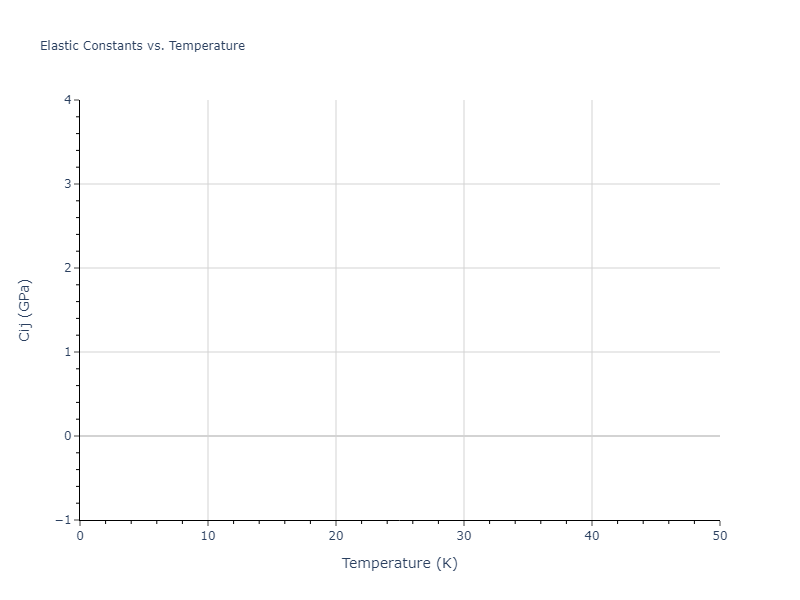
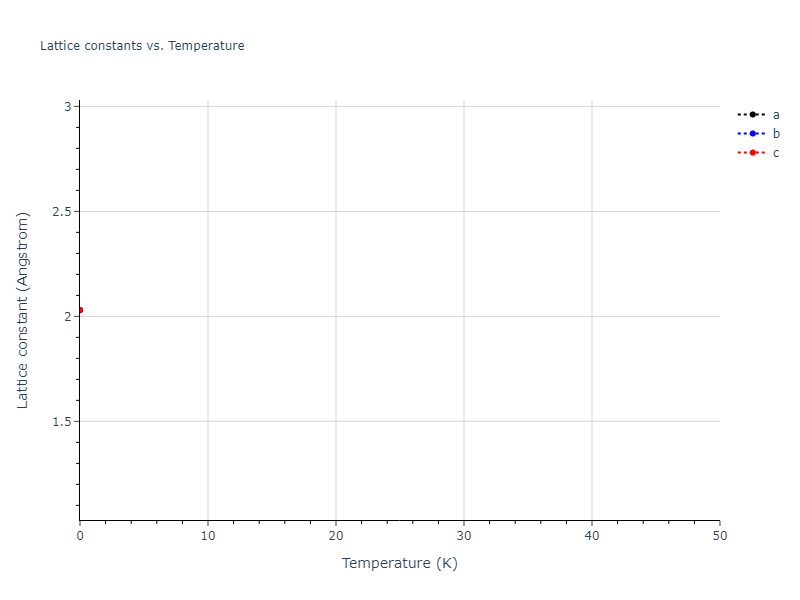
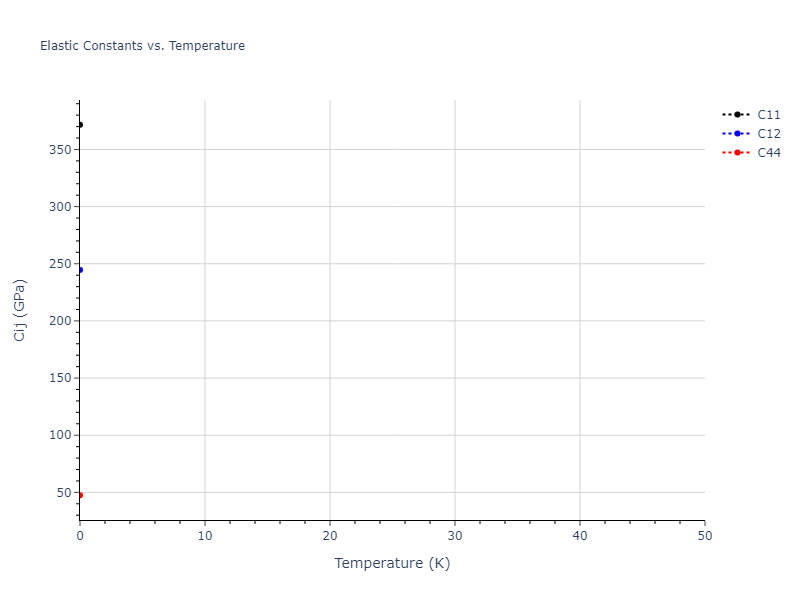
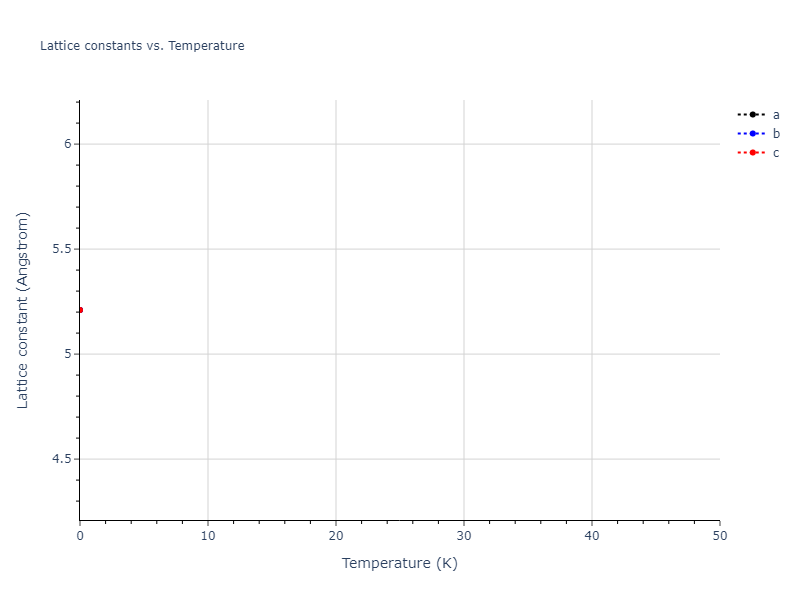
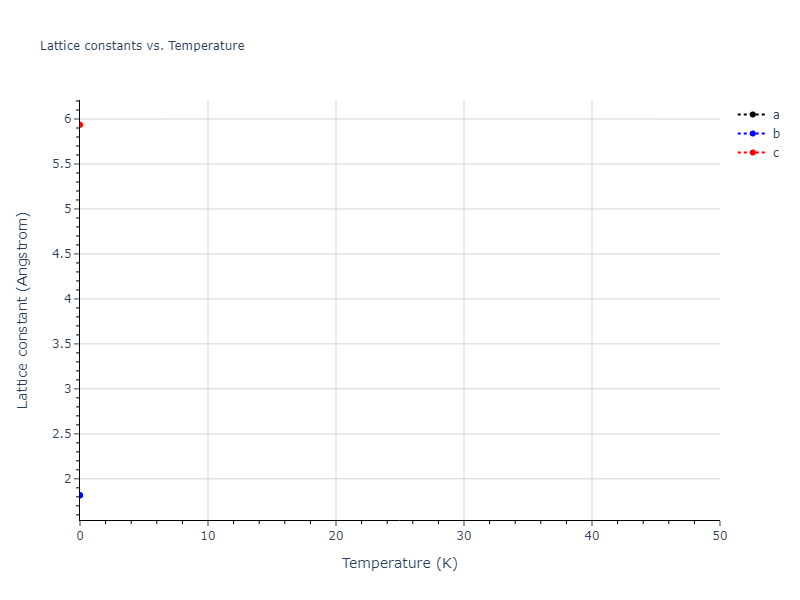
Elastic Constants Predictions
Static elastic constants are displayed for the unique structures identified in Crystal Structure Predictions above. The values displayed here are obtained by measuring the change in virial stresses due to applying small strains to the relaxed crystals. The initial structure and the strained states are all relaxed using force minimization.
The calculation method used is available as the iprPy elastic_constants_static calculation method.
Notes and Disclaimers:
- These values are meant to be guidelines for comparing potentials, not the absolute values for any potential's properties. Values listed here may change if the calculation methods are updated due to improvements/corrections. Variations in the values may occur for variations in calculation methods, simulation software and implementations of the interatomic potentials.
- The presence of any structures in this list does not guarantee that those structures are stable.
- The elastic constants have been computed for a variety of strains, and in some cases for slightly different lattice constant values. The static nature of this calculation can give poor predictions if the evaluated states straddle a functional discontinuity in the potential's third derivative. Be sure to compare the elastic constants for the different strains (positive and negative).
- NIST disclaimer
Version Information:
- 2019-08-07. Data added.
| 1564.791 | 1069.621 | 1069.621 | -0.000 | 0.000 | 0.000 |
| 1069.621 | 1564.791 | 1069.621 | 0.000 | -0.000 | 0.000 |
| 1069.621 | 1069.621 | 1564.791 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 508.069 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 508.069 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 508.069 |
| 1564.791 | 1069.621 | 1069.621 | 0.000 | -0.000 | -0.000 |
| 1069.621 | 1564.791 | 1069.621 | -0.000 | -0.000 | -0.000 |
| 1069.621 | 1069.621 | 1564.791 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 508.069 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 508.069 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 508.069 |
| 1564.792 | 1069.621 | 1069.621 | -0.000 | 0.000 | 0.000 |
| 1069.621 | 1564.792 | 1069.621 | 0.000 | -0.000 | 0.000 |
| 1069.621 | 1069.621 | 1564.792 | -0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 508.069 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 508.069 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 508.069 |
| 1564.791 | 1069.621 | 1069.621 | 0.000 | -0.000 | -0.000 |
| 1069.621 | 1564.791 | 1069.621 | -0.000 | -0.000 | -0.000 |
| 1069.621 | 1069.621 | 1564.791 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 508.069 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 508.069 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 508.069 |
| 1564.794 | 1069.624 | 1069.624 | -0.000 | 0.000 | 0.000 |
| 1069.624 | 1564.794 | 1069.624 | 0.000 | -0.000 | -0.000 |
| 1069.624 | 1069.624 | 1564.794 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.001 | 508.069 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.001 | -0.000 | 508.069 | 0.000 |
| 0.000 | 0.001 | -0.000 | 0.000 | 0.000 | 508.069 |
| 1564.789 | 1069.618 | 1069.618 | 0.000 | -0.000 | -0.000 |
| 1069.618 | 1564.789 | 1069.618 | -0.000 | -0.000 | -0.000 |
| 1069.618 | 1069.618 | 1564.789 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.001 | 508.069 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.001 | 0.000 | 508.069 | -0.000 |
| -0.000 | -0.001 | 0.000 | -0.000 | -0.000 | 508.069 |
| 5715.354 | -1162.198 | -1162.198 | -0.000 | 0.000 | 0.000 |
| -1162.198 | 5715.354 | -1162.199 | -0.000 | 0.000 | 0.000 |
| -1162.198 | -1162.198 | 5715.354 | 0.000 | 0.001 | 0.000 |
| -0.000 | 0.000 | -0.002 | 11911.777 | 0.001 | 0.000 |
| 0.000 | -0.000 | -0.001 | -0.000 | 11911.778 | 0.000 |
| 0.000 | 0.000 | -0.001 | 0.000 | 0.001 | 11911.778 |
| 5702.318 | -1155.680 | -1155.679 | 0.001 | -0.001 | -0.000 |
| -1155.680 | 5702.318 | -1155.679 | 0.000 | -0.001 | 0.000 |
| -1155.680 | -1155.680 | 5702.320 | 0.001 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.002 | 11911.777 | -0.001 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 11911.777 | 0.000 |
| -0.000 | -0.000 | 0.001 | -0.000 | -0.001 | 11911.777 |
| 5702.320 | -1155.680 | -1155.680 | -0.000 | 0.000 | 0.000 |
| -1155.680 | 5702.320 | -1155.680 | 0.000 | 0.000 | 0.000 |
| -1155.680 | -1155.680 | 5702.320 | -0.000 | -0.000 | 0.000 |
| -0.005 | 0.002 | 0.003 | 11939.774 | 0.000 | 0.000 |
| 0.002 | -0.005 | 0.004 | -0.000 | 11939.774 | 0.000 |
| 0.002 | 0.004 | -0.005 | -0.000 | 0.000 | 11939.774 |
| 5702.316 | -1155.679 | -1155.679 | 0.000 | 0.000 | -0.000 |
| -1155.679 | 5702.316 | -1155.679 | 0.000 | -0.000 | 0.000 |
| -1155.679 | -1155.679 | 5702.316 | 0.000 | -0.000 | -0.000 |
| 0.005 | -0.002 | -0.004 | 11939.774 | -0.000 | 0.000 |
| -0.002 | 0.005 | -0.004 | 0.000 | 11939.774 | -0.000 |
| -0.002 | -0.004 | 0.005 | 0.000 | -0.000 | 11939.774 |
| 5702.337 | -1155.683 | -1155.683 | 0.000 | 0.000 | -0.000 |
| -1155.683 | 5702.337 | -1155.683 | 0.000 | 0.000 | 0.000 |
| -1155.683 | -1155.683 | 5702.337 | 0.000 | 0.000 | 0.000 |
| -0.048 | 0.014 | 0.034 | 11911.778 | 0.000 | -0.000 |
| 0.014 | -0.048 | 0.035 | -0.000 | 11911.778 | -0.000 |
| 0.014 | 0.035 | -0.048 | -0.000 | 0.000 | 11911.778 |
| 5702.299 | -1155.676 | -1155.676 | 0.000 | -0.000 | -0.000 |
| -1155.676 | 5702.299 | -1155.676 | -0.000 | -0.000 | -0.000 |
| -1155.676 | -1155.676 | 5702.299 | -0.000 | 0.000 | -0.000 |
| 0.048 | -0.014 | -0.034 | 11911.778 | -0.000 | -0.000 |
| -0.014 | 0.048 | -0.034 | 0.000 | 11911.778 | -0.000 |
| -0.014 | -0.035 | 0.048 | -0.000 | -0.000 | 11911.778 |
| 931.202 | 907.073 | 907.073 | -0.000 | -0.000 | -0.000 |
| 907.073 | 931.202 | 907.073 | -0.000 | -0.000 | -0.000 |
| 907.073 | 907.073 | 931.202 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 495.977 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 495.977 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 495.977 |
| 931.202 | 907.073 | 907.073 | 0.000 | 0.000 | 0.000 |
| 907.073 | 931.202 | 907.073 | 0.000 | 0.000 | 0.000 |
| 907.073 | 907.073 | 931.202 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 495.977 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 495.977 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 495.977 |
| 931.202 | 907.073 | 907.073 | -0.000 | 0.000 | -0.000 |
| 907.073 | 931.202 | 907.073 | 0.000 | -0.000 | -0.000 |
| 907.073 | 907.073 | 931.202 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 495.977 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 495.977 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 495.977 |
| 931.202 | 907.073 | 907.073 | 0.000 | 0.000 | 0.000 |
| 907.073 | 931.202 | 907.073 | 0.000 | 0.000 | 0.000 |
| 907.073 | 907.073 | 931.202 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 495.977 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 495.977 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 495.977 |
| 931.203 | 907.075 | 907.075 | -0.000 | -0.000 | -0.000 |
| 907.075 | 931.203 | 907.075 | -0.000 | -0.000 | -0.000 |
| 907.075 | 907.075 | 931.203 | -0.000 | -0.000 | -0.000 |
| 0.001 | -0.000 | 0.001 | 495.977 | -0.000 | -0.000 |
| -0.000 | 0.001 | 0.001 | -0.000 | 495.977 | -0.000 |
| -0.000 | 0.001 | 0.001 | -0.000 | -0.000 | 495.977 |
| 931.201 | 907.071 | 907.071 | 0.000 | -0.000 | 0.000 |
| 907.071 | 931.201 | 907.071 | -0.000 | 0.000 | 0.000 |
| 907.071 | 907.071 | 931.201 | 0.000 | 0.000 | 0.000 |
| -0.001 | 0.000 | -0.001 | 495.977 | 0.000 | 0.000 |
| 0.000 | -0.001 | -0.001 | 0.000 | 495.977 | 0.000 |
| 0.000 | -0.001 | -0.001 | -0.000 | 0.000 | 495.977 |
| 1773.939 | 1033.337 | 907.863 | -0.000 | -0.000 | -0.000 |
| 997.256 | 1810.020 | 907.863 | -0.000 | 0.000 | -0.000 |
| 907.863 | 907.863 | 1878.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 307.334 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 307.334 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 370.301 |
| 1810.020 | 997.256 | 907.863 | -0.000 | -0.000 | 0.000 |
| 1033.337 | 1773.939 | 907.863 | 0.000 | 0.000 | 0.000 |
| 907.863 | 907.863 | 1878.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 307.334 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 307.334 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 370.301 |
| 1773.940 | 1033.337 | 907.863 | -0.000 | -0.000 | -0.000 |
| 1033.337 | 1773.940 | 907.863 | -0.000 | -0.000 | -0.000 |
| 907.863 | 907.863 | 1878.001 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 307.334 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 307.334 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 370.301 |
| 1773.939 | 1033.337 | 907.862 | 0.000 | 0.000 | 0.000 |
| 1033.337 | 1773.939 | 907.862 | 0.000 | 0.000 | 0.000 |
| 907.862 | 907.862 | 1878.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 307.334 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 307.334 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 370.301 |
| 1773.945 | 1033.340 | 907.864 | -0.000 | -0.000 | -0.000 |
| 997.260 | 1810.026 | 907.865 | -0.000 | 0.000 | -0.000 |
| 907.866 | 907.866 | 1878.005 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.001 | 307.334 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.001 | 0.000 | 307.334 | 0.000 |
| -0.000 | 0.001 | -0.000 | -0.000 | -0.000 | 370.301 |
| 1773.933 | 1033.335 | 907.861 | 0.000 | 0.000 | 0.000 |
| 997.253 | 1810.015 | 907.860 | -0.000 | -0.000 | 0.000 |
| 907.859 | 907.859 | 1877.995 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.001 | 307.334 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.001 | -0.000 | 307.334 | 0.000 |
| 0.000 | -0.001 | 0.000 | 0.000 | 0.000 | 370.301 |
| 798.645 | 673.678 | 545.470 | -0.000 | -0.000 | -0.000 |
| 504.314 | 968.009 | 545.470 | 0.000 | -0.000 | -0.000 |
| 545.470 | 545.470 | 1091.265 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 225.293 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 225.293 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 231.847 |
| 798.645 | 673.678 | 545.470 | 0.000 | 0.000 | 0.000 |
| 673.678 | 798.645 | 545.470 | 0.000 | -0.000 | 0.000 |
| 545.470 | 545.470 | 1091.266 | -0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 225.293 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 225.293 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 231.847 |
| 968.009 | 504.314 | 545.470 | -0.000 | -0.000 | -0.000 |
| 504.314 | 968.009 | 545.470 | -0.000 | -0.000 | -0.000 |
| 545.471 | 545.471 | 1091.265 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 225.293 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 225.293 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 231.847 |
| 968.008 | 504.314 | 545.471 | -0.000 | 0.000 | 0.000 |
| 673.677 | 798.645 | 545.471 | 0.000 | 0.000 | 0.000 |
| 545.470 | 545.470 | 1091.266 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 225.293 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 225.293 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 0.000 | 231.847 |
| 798.654 | 673.674 | 545.469 | -0.000 | -0.000 | -0.000 |
| 673.661 | 798.667 | 545.469 | -0.000 | -0.000 | -0.000 |
| 545.477 | 545.477 | 1091.259 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 225.293 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 225.293 | -0.000 |
| -0.001 | 0.000 | -0.000 | -0.000 | -0.000 | 62.483 |
| 798.663 | 673.655 | 545.471 | 0.000 | 0.000 | 0.000 |
| 673.667 | 798.650 | 545.471 | 0.000 | 0.000 | 0.000 |
| 545.464 | 545.464 | 1091.272 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 225.293 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 225.293 | 0.000 |
| 0.001 | -0.000 | 0.000 | 0.000 | 0.000 | 62.483 |
| 1766.185 | 1040.170 | 920.559 | 0.000 | -0.000 | 0.000 |
| 1040.170 | 1766.185 | 920.559 | 0.000 | -0.000 | 0.000 |
| 920.559 | 920.559 | 1841.252 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 278.399 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 278.399 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 390.526 |
| 1766.185 | 1040.169 | 920.559 | 0.000 | 0.000 | -0.000 |
| 1012.651 | 1793.703 | 920.559 | -0.000 | 0.000 | 0.000 |
| 920.559 | 920.559 | 1841.252 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 278.399 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 278.399 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 390.526 |
| 1766.185 | 1040.170 | 920.559 | 0.000 | -0.000 | -0.000 |
| 1012.652 | 1793.703 | 920.560 | 0.000 | -0.000 | -0.000 |
| 920.560 | 920.560 | 1841.252 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 278.399 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 278.399 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 363.008 |
| 1766.184 | 1040.169 | 920.559 | -0.000 | 0.000 | -0.000 |
| 1040.169 | 1766.184 | 920.559 | -0.000 | 0.000 | -0.000 |
| 920.559 | 920.559 | 1841.251 | -0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 278.399 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 278.399 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 0.000 | 363.008 |
| 1766.191 | 1040.172 | 920.561 | 0.000 | -0.000 | 0.000 |
| 1040.173 | 1766.190 | 920.561 | 0.000 | -0.000 | 0.000 |
| 920.562 | 920.562 | 1841.257 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 278.399 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 278.399 | 0.000 |
| -0.000 | 0.001 | -0.000 | -0.000 | -0.000 | 363.008 |
| 1766.179 | 1040.167 | 920.557 | 0.000 | 0.000 | -0.000 |
| 1040.166 | 1766.180 | 920.557 | -0.000 | 0.000 | 0.000 |
| 920.556 | 920.556 | 1841.247 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 278.399 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 278.399 | 0.000 |
| 0.000 | -0.001 | 0.000 | 0.000 | 0.000 | 363.008 |
| 406.606 | 351.767 | 351.767 | 0.000 | -0.000 | -0.000 |
| 351.767 | 406.606 | 351.767 | -0.000 | 0.000 | -0.000 |
| 351.767 | 351.767 | 406.606 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 63.500 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 63.500 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 63.500 |
| 406.606 | 351.767 | 351.767 | -0.000 | 0.000 | 0.000 |
| 351.767 | 406.606 | 351.767 | 0.000 | -0.000 | 0.000 |
| 351.767 | 351.767 | 406.606 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 63.500 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 63.500 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 63.500 |
| 406.606 | 351.767 | 351.767 | 0.000 | -0.000 | -0.000 |
| 351.767 | 406.606 | 351.767 | -0.000 | 0.000 | -0.000 |
| 351.767 | 351.767 | 406.606 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 63.500 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 63.500 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 63.500 |
| 406.606 | 351.766 | 351.766 | -0.000 | 0.000 | 0.000 |
| 351.766 | 406.606 | 351.766 | 0.000 | -0.000 | 0.000 |
| 351.766 | 351.766 | 406.606 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 63.500 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 63.500 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 63.500 |
| 406.607 | 351.767 | 351.767 | 0.000 | -0.000 | -0.000 |
| 351.767 | 406.607 | 351.767 | -0.000 | 0.000 | -0.000 |
| 351.767 | 351.767 | 406.607 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 63.500 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 63.500 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 63.500 |
| 406.605 | 351.766 | 351.766 | 0.000 | 0.000 | 0.000 |
| 351.766 | 406.605 | 351.766 | 0.000 | 0.000 | 0.000 |
| 351.766 | 351.766 | 406.605 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 63.500 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 63.500 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 63.500 |
| 638.445 | 692.430 | 487.414 | 0.000 | -0.000 | 0.000 |
| 692.430 | 638.446 | 487.414 | 0.000 | -0.000 | 0.000 |
| 487.414 | 487.415 | 1437.712 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | -6.942 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -6.942 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 27.758 |
| 638.446 | 692.430 | 487.414 | -0.000 | -0.000 | 0.000 |
| 692.430 | 638.445 | 487.414 | -0.000 | 0.000 | 0.000 |
| 487.415 | 487.414 | 1437.712 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -6.942 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -6.942 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 27.758 |
| 638.445 | 692.430 | 487.414 | -0.000 | -0.000 | 0.000 |
| 692.430 | 638.445 | 487.414 | 0.000 | -0.000 | 0.000 |
| 487.415 | 487.415 | 1437.712 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -6.942 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -6.942 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 27.758 |
| 638.446 | 692.430 | 487.414 | -0.000 | 0.000 | 0.000 |
| 692.430 | 638.446 | 487.414 | -0.000 | 0.000 | 0.000 |
| 487.414 | 487.414 | 1437.711 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -6.942 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -6.942 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 27.758 |
| 638.444 | 692.433 | 487.415 | 0.000 | -0.000 | 0.000 |
| 692.433 | 638.444 | 487.415 | 0.000 | -0.000 | -0.000 |
| 487.416 | 487.416 | 1437.718 | 0.000 | -0.000 | 0.000 |
| -0.001 | -0.000 | -0.001 | -6.942 | -0.000 | 0.000 |
| -0.000 | -0.001 | -0.001 | -0.000 | -6.942 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 27.758 |
| 638.447 | 692.427 | 487.414 | -0.000 | 0.000 | 0.000 |
| 692.427 | 638.447 | 487.414 | -0.000 | 0.000 | -0.000 |
| 487.413 | 487.413 | 1437.705 | -0.000 | 0.000 | -0.000 |
| 0.001 | 0.000 | 0.001 | -6.942 | 0.000 | -0.000 |
| 0.000 | 0.001 | 0.001 | 0.000 | -6.942 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 27.758 |
| 870.869 | 269.020 | 0.000 | 0.000 | 0.000 | -0.000 |
| 269.020 | 870.869 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 5.124 |
| 870.869 | 269.020 | -0.000 | -0.000 | -0.000 | -0.000 |
| 269.020 | 870.869 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 5.124 |
| 870.870 | 269.020 | 0.000 | 0.000 | 0.000 | -0.000 |
| 269.020 | 870.870 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 5.124 |
| 870.869 | 269.020 | -0.000 | -0.000 | -0.000 | 0.000 |
| 269.020 | 870.869 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 5.124 |
| 870.873 | 269.020 | 0.000 | 0.000 | 0.000 | 0.000 |
| 269.020 | 870.873 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 5.124 |
| 870.865 | 269.019 | -0.000 | -0.000 | -0.000 | -0.000 |
| 269.019 | 870.865 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 5.124 |
| 889.994 | 274.927 | 0.000 | 0.000 | 0.000 | -0.000 |
| 274.927 | 889.994 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 5.237 |
| 889.994 | 274.927 | -0.000 | -0.000 | -0.000 | 0.000 |
| 274.927 | 889.994 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 5.237 |
| 889.994 | 274.927 | 0.000 | 0.000 | 0.000 | -0.000 |
| 274.927 | 889.994 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 5.237 |
| 889.993 | 274.927 | -0.000 | -0.000 | -0.000 | -0.000 |
| 274.927 | 889.993 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 5.237 |
| 889.998 | 274.928 | 0.000 | 0.000 | 0.000 | 0.000 |
| 274.928 | 889.998 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 5.237 |
| 889.989 | 274.927 | -0.000 | -0.000 | -0.000 | -0.000 |
| 274.927 | 889.989 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 5.237 |
| 1280.751 | 272.125 | 272.125 | 0.000 | 0.000 | -0.000 |
| 272.125 | 1280.751 | 272.125 | 0.000 | 0.000 | 0.000 |
| 272.125 | 272.125 | 1280.751 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 7.207 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 7.207 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 7.207 |
| 1280.751 | 272.125 | 272.125 | 0.000 | -0.000 | 0.000 |
| 272.125 | 1280.751 | 272.125 | -0.000 | 0.000 | -0.000 |
| 272.125 | 272.125 | 1280.751 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 7.207 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 7.207 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 7.207 |
| 1280.752 | 272.125 | 272.125 | 0.000 | 0.000 | -0.000 |
| 272.125 | 1280.752 | 272.125 | 0.000 | -0.000 | 0.000 |
| 272.125 | 272.125 | 1280.752 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 7.207 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 7.207 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 7.207 |
| 1280.751 | 272.125 | 272.125 | -0.000 | -0.000 | -0.000 |
| 272.125 | 1280.751 | 272.125 | -0.000 | 0.000 | -0.000 |
| 272.125 | 272.125 | 1280.751 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 7.207 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 7.207 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 7.207 |
| 1280.757 | 272.126 | 272.126 | 0.000 | 0.000 | 0.000 |
| 272.126 | 1280.757 | 272.126 | -0.000 | -0.000 | -0.000 |
| 272.126 | 272.126 | 1280.757 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.001 | 7.207 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.001 | -0.000 | 7.207 | -0.000 |
| -0.000 | -0.001 | -0.000 | -0.000 | -0.000 | 7.207 |
| 1280.745 | 272.124 | 272.124 | 0.000 | 0.000 | -0.000 |
| 272.124 | 1280.745 | 272.124 | -0.000 | -0.000 | -0.000 |
| 272.124 | 272.124 | 1280.745 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.001 | 7.207 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.001 | -0.000 | 7.207 | -0.000 |
| 0.000 | 0.001 | 0.000 | -0.000 | -0.000 | 7.207 |
| 506.347 | 200.589 | 223.128 | -0.000 | 0.000 | -0.000 |
| 306.900 | 651.763 | 294.830 | -0.000 | 0.000 | 0.000 |
| 168.746 | 262.143 | 688.429 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 231.444 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 9.167 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 116.189 |
| 621.377 | 313.111 | 207.644 | 0.000 | -0.000 | -0.000 |
| 203.169 | 499.942 | 387.576 | -0.000 | -0.000 | 0.000 |
| 221.535 | 326.247 | 661.818 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 231.444 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 9.167 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 116.189 |
| 621.377 | 313.111 | 207.644 | -0.000 | 0.000 | -0.000 |
| 306.901 | 651.763 | 294.830 | -0.000 | 0.000 | -0.000 |
| 207.842 | 330.397 | 636.210 | 0.000 | -0.000 | -0.000 |
| -1.257 | 0.488 | -2.519 | 117.471 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 9.167 | -0.000 |
| -2.059 | 0.736 | -4.025 | 0.000 | 0.000 | 52.566 |
| 522.274 | 193.969 | 255.719 | 0.000 | -0.000 | 0.000 |
| 191.413 | 503.787 | 365.149 | 0.000 | -0.000 | 0.000 |
| 210.455 | 329.543 | 641.194 | 0.000 | -0.000 | 0.000 |
| 1.257 | -0.488 | 2.519 | 117.471 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 9.167 | 0.000 |
| 2.059 | -0.736 | 4.025 | 0.000 | -0.000 | 52.566 |
| 519.278 | 196.356 | 247.799 | -0.000 | 0.000 | -0.000 |
| 189.973 | 504.254 | 362.411 | -0.000 | 0.000 | -0.000 |
| 209.018 | 330.013 | 638.455 | 0.000 | 0.000 | -0.000 |
| -0.124 | 0.051 | -0.253 | 117.471 | 0.000 | -0.000 |
| -0.128 | 0.046 | -0.251 | 0.000 | 9.167 | 0.000 |
| -2.654 | -2.382 | 0.022 | 0.000 | 0.000 | 42.201 |
| 519.540 | 196.277 | 248.294 | 0.000 | -0.000 | -0.000 |
| 190.240 | 504.174 | 362.904 | 0.000 | -0.000 | 0.000 |
| 209.279 | 329.927 | 638.949 | 0.000 | 0.000 | 0.000 |
| 0.124 | -0.051 | 0.253 | 117.471 | 0.000 | 0.000 |
| 0.128 | -0.046 | 0.251 | -0.000 | 9.167 | 0.000 |
| 2.654 | 2.382 | -0.022 | -0.000 | -0.000 | 42.201 |
| 715.751 | 230.419 | 174.844 | -0.000 | 0.000 | 0.000 |
| 230.419 | 715.751 | 174.844 | -0.000 | 0.000 | 0.000 |
| 174.625 | 174.625 | 578.310 | 0.000 | 0.000 | 0.000 |
| 0.027 | 0.027 | -0.018 | 74.949 | 0.000 | 0.000 |
| 0.027 | 0.027 | -0.018 | 0.000 | 74.949 | 0.000 |
| 0.039 | 0.039 | -0.026 | -0.000 | 0.000 | 45.949 |
| 382.723 | 256.628 | 276.704 | 0.000 | -0.000 | 0.000 |
| 256.628 | 382.723 | 276.704 | -0.000 | -0.000 | 0.000 |
| 174.625 | 174.625 | 578.310 | 0.000 | -0.000 | -0.000 |
| -0.027 | -0.027 | 0.018 | 74.949 | -0.000 | -0.000 |
| -0.027 | -0.027 | 0.018 | 0.000 | 74.949 | -0.000 |
| -0.039 | -0.039 | 0.026 | 0.000 | 0.000 | 45.949 |
| 422.078 | 146.519 | 300.193 | 0.000 | 0.000 | 0.000 |
| 146.519 | 422.078 | 300.193 | 0.000 | 0.000 | 0.000 |
| 276.473 | 276.473 | 510.686 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 145.259 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 145.259 | 0.000 |
| 0.004 | 0.004 | -0.003 | -0.000 | -0.000 | 45.949 |
| 422.007 | 146.495 | 300.225 | -0.000 | -0.000 | -0.000 |
| 146.495 | 422.007 | 300.225 | 0.000 | -0.000 | 0.000 |
| 276.470 | 276.470 | 510.688 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 145.259 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 145.259 | -0.000 |
| -0.004 | -0.004 | 0.003 | 0.000 | -0.000 | 45.949 |
| 382.711 | 256.636 | 276.707 | 0.000 | 0.000 | -0.000 |
| 256.636 | 382.711 | 276.707 | -0.000 | 0.000 | -0.000 |
| 174.626 | 174.626 | 578.312 | 0.000 | 0.000 | 0.000 |
| 0.001 | 0.000 | -0.000 | 74.949 | 0.000 | 0.000 |
| 0.000 | 0.001 | -0.000 | 0.000 | 74.949 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 45.949 |
| 382.775 | 256.660 | 276.675 | -0.000 | -0.000 | -0.000 |
| 256.660 | 382.775 | 276.675 | 0.000 | -0.000 | -0.000 |
| 276.471 | 276.471 | 510.685 | -0.000 | -0.000 | 0.000 |
| -0.001 | -0.000 | 0.000 | 74.949 | -0.000 | -0.000 |
| -0.000 | -0.001 | 0.000 | 0.000 | 74.949 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 45.949 |
| 124.931 | 188.913 | 188.913 | -0.000 | -0.000 | 0.000 |
| 188.913 | 124.931 | 188.913 | 0.000 | -0.000 | -0.000 |
| 188.913 | 188.913 | 124.931 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -124.670 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -124.670 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -124.670 |
| 124.931 | 188.913 | 188.913 | -0.000 | -0.000 | 0.000 |
| 188.913 | 124.931 | 188.913 | -0.000 | -0.000 | -0.000 |
| 188.913 | 188.913 | 124.931 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -124.670 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -124.670 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -124.670 |
| 124.931 | 188.913 | 188.913 | 0.000 | 0.000 | 0.000 |
| 188.913 | 124.931 | 188.913 | 0.000 | 0.000 | 0.000 |
| 188.913 | 188.913 | 124.931 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -124.670 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -124.670 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -124.670 |
| 124.930 | 188.914 | 188.914 | 0.000 | -0.000 | -0.000 |
| 188.914 | 124.930 | 188.914 | 0.000 | 0.000 | -0.000 |
| 188.914 | 188.914 | 124.930 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -124.670 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -124.670 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -124.670 |
| 124.934 | 188.913 | 188.913 | 0.000 | 0.000 | 0.000 |
| 188.913 | 124.934 | 188.913 | 0.000 | -0.000 | 0.000 |
| 188.913 | 188.913 | 124.934 | 0.000 | 0.000 | 0.000 |
| 0.001 | 0.000 | 0.000 | -124.670 | -0.000 | -0.000 |
| 0.000 | 0.001 | 0.000 | 0.000 | -124.670 | 0.000 |
| 0.000 | 0.000 | 0.001 | 0.000 | -0.000 | -124.670 |
| 124.928 | 188.914 | 188.914 | -0.000 | -0.000 | -0.000 |
| 188.914 | 124.928 | 188.914 | -0.000 | -0.000 | -0.000 |
| 188.914 | 188.914 | 124.928 | -0.000 | -0.000 | -0.000 |
| -0.001 | -0.000 | -0.000 | -124.670 | -0.000 | -0.000 |
| -0.000 | -0.001 | -0.000 | -0.000 | -124.670 | -0.000 |
| -0.000 | -0.000 | -0.001 | -0.000 | -0.000 | -124.670 |
| 573.387 | 563.169 | 0.000 | 0.000 | 0.000 | 0.000 |
| 563.169 | 573.387 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 300.045 |
| 573.387 | 563.169 | -0.000 | -0.000 | -0.000 | -0.000 |
| 563.169 | 573.387 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 300.045 |
| 573.387 | 563.169 | 0.000 | 0.000 | 0.000 | -0.000 |
| 563.169 | 573.387 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 300.045 |
| 573.387 | 563.169 | -0.000 | -0.000 | -0.000 | -0.000 |
| 563.169 | 573.387 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 300.045 |
| 573.388 | 563.171 | 0.000 | 0.000 | 0.000 | 0.000 |
| 563.171 | 573.388 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.001 | 0.000 | 0.000 | 0.000 | 300.045 |
| 573.387 | 563.167 | -0.000 | -0.000 | -0.000 | 0.000 |
| 563.167 | 573.387 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.001 | -0.000 | -0.000 | -0.000 | 300.045 |
| 252.978 | 246.801 | 331.862 | -0.000 | 0.000 | -0.000 |
| 103.540 | 436.420 | 383.836 | 0.000 | 0.000 | -0.000 |
| 175.454 | 235.225 | 611.222 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 162.981 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 122.072 | 0.000 |
| -120.418 | -58.300 | 69.318 | 0.000 | -0.000 | 52.501 |
| 503.230 | 304.797 | 229.438 | -0.000 | -0.000 | -0.000 |
| 208.683 | 672.873 | 237.175 | -0.000 | -0.000 | -0.000 |
| 384.285 | 367.370 | 452.703 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 162.981 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 122.072 | -0.000 |
| 120.418 | 58.300 | -69.318 | -0.000 | -0.000 | 52.501 |
| 694.847 | 207.801 | 176.521 | -0.000 | 0.000 | 0.000 |
| 260.706 | 440.772 | 328.265 | 0.000 | 0.000 | -0.000 |
| 175.454 | 235.225 | 611.222 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 162.981 | 0.000 | 0.000 |
| -13.663 | -6.615 | 7.865 | 0.000 | 56.396 | 0.000 |
| -15.606 | -7.556 | 8.984 | 0.000 | 0.000 | 43.425 |
| 694.847 | 207.801 | 176.521 | -0.000 | -0.000 | -0.000 |
| 226.364 | 458.253 | 330.602 | 0.000 | -0.000 | -0.000 |
| 175.453 | 235.225 | 611.222 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 162.981 | -0.000 | -0.000 |
| 13.663 | 6.615 | -7.865 | -0.000 | 56.396 | -0.000 |
| 15.606 | 7.556 | -8.984 | -0.000 | -0.000 | 43.425 |
| 375.463 | 269.461 | 281.880 | 0.000 | 0.000 | 0.000 |
| 211.848 | 458.171 | 335.736 | 0.000 | 0.000 | 0.000 |
| 288.198 | 323.968 | 504.170 | 0.000 | 0.000 | 0.000 |
| 3.950 | 7.305 | -6.148 | 53.572 | 0.000 | 0.000 |
| -1.920 | 0.221 | 0.228 | -0.000 | 54.282 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 83.158 |
| 377.924 | 270.260 | 280.664 | -0.000 | -0.000 | -0.000 |
| 274.740 | 444.110 | 322.570 | -0.000 | -0.000 | -0.000 |
| 289.905 | 324.612 | 503.406 | -0.000 | -0.000 | -0.000 |
| -3.950 | -7.305 | 6.148 | 53.572 | -0.000 | -0.000 |
| 1.920 | -0.221 | -0.228 | -0.000 | 54.282 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 83.158 |
| 192.921 | 361.836 | 139.536 | -0.000 | -64.776 | 0.000 |
| 334.630 | 271.738 | 227.057 | -0.000 | -41.890 | 0.000 |
| 248.385 | 253.964 | 543.670 | 0.000 | 50.358 | 0.000 |
| -0.000 | 0.000 | 0.000 | 192.249 | -0.000 | -66.305 |
| -86.624 | 62.451 | -192.431 | -0.000 | -70.358 | 0.000 |
| -73.633 | 38.037 | -177.876 | 100.759 | -81.339 | -66.328 |
| 298.206 | 332.984 | 346.814 | -0.000 | 143.731 | -0.000 |
| 175.344 | 540.962 | 381.972 | -0.000 | 74.207 | 0.000 |
| 379.026 | 421.622 | 380.059 | 0.000 | 29.866 | -0.000 |
| 0.000 | -0.000 | -0.000 | 192.249 | -0.000 | -66.305 |
| 48.596 | 9.009 | 139.811 | 0.000 | 54.189 | -0.000 |
| 73.633 | -38.037 | 177.876 | 100.759 | 81.339 | -66.328 |
| 394.028 | 276.827 | 249.088 | 0.000 | 5.972 | 0.000 |
| 286.431 | 478.557 | 289.881 | 0.000 | -29.304 | -0.000 |
| 248.385 | 253.964 | 543.670 | 0.000 | 50.358 | 0.000 |
| 11.412 | -7.749 | -6.987 | 74.647 | -9.840 | -8.777 |
| 23.853 | -65.384 | 50.358 | -0.000 | 158.741 | -0.000 |
| -2.838 | 1.466 | -6.855 | -3.460 | -3.135 | 34.571 |
| 363.935 | 278.293 | 305.808 | -0.000 | 22.969 | 0.000 |
| 270.828 | 487.267 | 312.864 | 0.000 | -25.756 | -0.000 |
| 293.213 | 313.007 | 431.927 | 0.000 | 26.419 | 0.000 |
| -11.412 | 7.749 | 6.987 | 74.647 | 9.840 | -8.777 |
| 23.853 | -65.384 | 50.358 | 0.000 | 158.741 | -0.000 |
| 2.838 | -1.466 | 6.855 | -3.460 | 3.135 | 34.571 |
| 375.270 | 273.043 | 280.132 | 0.000 | 12.290 | 0.000 |
| 273.275 | 485.945 | 298.520 | 0.000 | -26.999 | -0.000 |
| 255.210 | 311.011 | 436.717 | 0.000 | 15.444 | 0.000 |
| 0.117 | -0.103 | -0.532 | 74.426 | -0.333 | -7.548 |
| 6.824 | -30.304 | 18.170 | 0.000 | 100.167 | 0.000 |
| 0.000 | 0.000 | 0.000 | -68.814 | 0.000 | 97.843 |
| 377.591 | 275.084 | 288.271 | -0.000 | 16.697 | -0.000 |
| 267.826 | 489.525 | 306.541 | -0.000 | -28.737 | -0.000 |
| 281.743 | 310.186 | 432.098 | -0.000 | 20.735 | -0.000 |
| -0.117 | 0.103 | 0.532 | 74.426 | 0.333 | -7.548 |
| 4.978 | -23.735 | 17.589 | 0.000 | 98.411 | -0.000 |
| -0.000 | -0.000 | -0.000 | -68.814 | -0.000 | 97.843 |
| 27.421 | 31.707 | -351.970 | -0.000 | 0.000 | 0.000 |
| 31.707 | 27.421 | -351.970 | 0.000 | -0.000 | 0.000 |
| 38.009 | 38.009 | 128.920 | -0.000 | 0.000 | -0.000 |
| 107.400 | 107.400 | -355.700 | 18.779 | 0.000 | 0.000 |
| 107.400 | 107.400 | -355.700 | 0.000 | 18.779 | 0.000 |
| 805.875 | 805.875 | 365.350 | -0.000 | -0.000 | 8.885 |
| -261.820 | -254.434 | 305.208 | -0.000 | -0.000 | -0.000 |
| -254.434 | -261.820 | 305.208 | -0.000 | -0.000 | -0.000 |
| -810.962 | -810.962 | -381.463 | 0.000 | -0.000 | -0.000 |
| -107.400 | -107.400 | 355.700 | 18.779 | -0.000 | -0.000 |
| -107.400 | -107.400 | 355.700 | -0.000 | 18.779 | -0.000 |
| -805.875 | -805.875 | -365.350 | 0.000 | -0.000 | 8.885 |
| 12.443 | 23.114 | 24.552 | -0.000 | 0.000 | 0.000 |
| 23.114 | 12.443 | 24.552 | 0.000 | 0.000 | -0.000 |
| 38.009 | 38.009 | 128.921 | 0.000 | 0.000 | -0.000 |
| 82.180 | 82.180 | 37.736 | 13.996 | -0.000 | 0.000 |
| 82.180 | 82.180 | 37.736 | -0.000 | 13.996 | 0.000 |
| 71.507 | 71.507 | 29.296 | -0.000 | -0.000 | 8.179 |
| -115.315 | -108.841 | -91.543 | -0.000 | -0.000 | -0.000 |
| -108.841 | -115.315 | -91.543 | -0.000 | -0.000 | -0.000 |
| -75.068 | -75.068 | -31.507 | 0.000 | -0.000 | -0.000 |
| -82.180 | -82.180 | -37.736 | 13.996 | -0.000 | -0.000 |
| -82.180 | -82.180 | -37.736 | -0.000 | 13.996 | -0.000 |
| -71.507 | -71.507 | -29.296 | 0.000 | -0.000 | 8.179 |
| -7590289.472 | -2826932.217 | -1298675.041 | 0.000 | 0.000 | 0.000 |
| -2826932.267 | -7590289.488 | -1298675.043 | 0.000 | -0.000 | 0.000 |
| -13.188 | -13.188 | 10.978 | 0.000 | 0.000 | 0.000 |
| 8.452 | 8.454 | 3.261 | 9.973 | -0.000 | -0.000 |
| 8.454 | 8.452 | 3.261 | 0.000 | 9.973 | -0.000 |
| 0.353 | 0.353 | -1.169 | 0.000 | 0.000 | 30.439 |
| 2826964.096 | 7590281.568 | 1298671.802 | 0.000 | -0.000 | 0.000 |
| 7590281.548 | 2826964.088 | 1298671.796 | 0.000 | -0.000 | -0.000 |
| -30.174 | -30.174 | 4.289 | -0.000 | -0.000 | 0.000 |
| -8.452 | -8.454 | -3.261 | 9.973 | -0.000 | -0.000 |
| -8.454 | -8.452 | -3.261 | 0.000 | 9.973 | -0.000 |
| -0.353 | -0.353 | 1.169 | 0.000 | -0.000 | 30.439 |
| 371.374 | 364.755 | 0.000 | 0.000 | 0.000 | 0.000 |
| 364.755 | 371.374 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 194.334 |
| 371.374 | 364.755 | -0.000 | -0.000 | -0.000 | -0.000 |
| 364.755 | 371.374 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 194.334 |
| 371.374 | 364.755 | 0.000 | 0.000 | 0.000 | -0.000 |
| 364.755 | 371.374 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 194.334 |
| 371.374 | 364.755 | -0.000 | -0.000 | -0.000 | -0.000 |
| 364.755 | 371.374 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 194.334 |
| 371.374 | 364.757 | 0.000 | 0.000 | 0.000 | 0.000 |
| 364.757 | 371.374 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.001 | 0.000 | 0.000 | 0.000 | 194.334 |
| 371.373 | 364.754 | -0.000 | -0.000 | -0.000 | 0.000 |
| 364.754 | 371.373 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.001 | -0.000 | -0.000 | -0.000 | 194.334 |
| -332.580 | 10.937 | 200.297 | 1.499 | 0.294 | 3.546 |
| -362.690 | 24.877 | 205.030 | 1.526 | 0.299 | 3.609 |
| 509.143 | 43.889 | 610.302 | 3.490 | 1.757 | 1.706 |
| -449.697 | -79.521 | 115.095 | 7.202 | 0.297 | 3.589 |
| -449.758 | -79.531 | 115.110 | 1.517 | 13.973 | 3.589 |
| -449.891 | -79.555 | 115.145 | 1.518 | 0.298 | 26.045 |
| 565.520 | 170.225 | -30.459 | -1.538 | -0.302 | -3.638 |
| 537.050 | 184.944 | -25.131 | -1.510 | -0.296 | -3.571 |
| 534.606 | 168.663 | 10.098 | -1.512 | -0.296 | -3.576 |
| 449.708 | 79.523 | -115.098 | 4.166 | -0.297 | -3.589 |
| 449.768 | 79.533 | -115.113 | -1.517 | 13.377 | -3.589 |
| 449.770 | 79.534 | -115.114 | -1.517 | -0.298 | 18.875 |
| -15.887 | 3.962 | 37.311 | 0.348 | 0.110 | 0.051 |
| 86.181 | 196.107 | 101.259 | -0.000 | 0.000 | -0.000 |
| 7.294 | -8.737 | 42.657 | 0.359 | 0.116 | 0.004 |
| 8.616 | -29.842 | 23.061 | 5.334 | 0.205 | 0.171 |
| -43.969 | -7.775 | 11.253 | 0.148 | 14.833 | 0.351 |
| 7.654 | -34.064 | 14.916 | 0.340 | 0.179 | -17.828 |
| 148.459 | 99.543 | 70.490 | -0.179 | -0.035 | -0.423 |
| 132.453 | 116.866 | 78.840 | -0.145 | -0.028 | -0.343 |
| 129.940 | 97.515 | 118.667 | -0.145 | -0.028 | -0.342 |
| -0.000 | -0.000 | -0.000 | 61.136 | 0.000 | 0.000 |
| 43.977 | 7.777 | -11.256 | -0.148 | 14.765 | -0.351 |
| 44.510 | 7.871 | -11.392 | -0.150 | -0.029 | 22.478 |
| 12.142 | 8.844 | 18.192 | 0.029 | 0.011 | 0.026 |
| 2.000 | 4.477 | 3.781 | 0.027 | 0.008 | 0.013 |
| 37.282 | 40.940 | 68.327 | 0.031 | 0.010 | 0.001 |
| -2.107 | -2.502 | 0.421 | 12.300 | 0.007 | 0.008 |
| 1.266 | 0.011 | 3.214 | 0.027 | 1.856 | -0.006 |
| -2.804 | -2.613 | 0.600 | 0.025 | 0.007 | 13.088 |
| 91.710 | 91.562 | 81.137 | -0.020 | -0.004 | -0.046 |
| 0.118 | 6.310 | -7.746 | -0.026 | -0.008 | 0.003 |
| 9.722 | 20.342 | 30.393 | -0.031 | -0.012 | -0.027 |
| 2.107 | 2.502 | -0.421 | 12.256 | -0.007 | -0.008 |
| -1.263 | -0.008 | -3.212 | -0.027 | 1.842 | 0.006 |
| 2.809 | 2.612 | -0.601 | -0.025 | -0.007 | 13.049 |
| 4605.420 | 6193.654 | 7198.853 | 0.109 | -0.003 | 0.036 |
| 3752.978 | 4975.248 | 5118.651 | -2.751 | -0.281 | 0.014 |
| 3895.873 | 5236.676 | 4887.805 | -0.469 | -0.031 | -0.021 |
| 1469.407 | 2403.808 | 854.704 | 10.389 | -0.003 | 0.003 |
| 1469.389 | 2403.778 | 854.693 | 0.012 | -3.046 | 0.003 |
| 1469.420 | 2403.828 | 854.711 | 0.012 | -0.003 | 15.107 |
| -4945.777 | -779.852 | -5352.348 | -1.131 | -0.333 | -0.000 |
| -7547.359 | -3445.927 | -11322.690 | -1.134 | 0.217 | 0.000 |
| -1395.321 | -2329.270 | -752.383 | -0.012 | 0.003 | -0.003 |
| -1469.407 | -2403.808 | -854.704 | 10.366 | 0.003 | -0.003 |
| -1469.389 | -2403.778 | -854.693 | -0.012 | -3.040 | -0.003 |
| -1469.420 | -2403.828 | -854.711 | -0.012 | 0.003 | 15.101 |
| 395.516 | 466.475 | 533.929 | -0.259 | -0.093 | 0.001 |
| 76.692 | 196.167 | 96.111 | -0.000 | 0.000 | -0.000 |
| 206.615 | 495.327 | 642.621 | -0.001 | 0.000 | 0.000 |
| 427.203 | 277.018 | 333.153 | -75.991 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 57.597 | 0.000 |
| 555.507 | 623.592 | 679.913 | -0.206 | -0.026 | -5.441 |
| -410.437 | -345.594 | -255.212 | -0.076 | -0.014 | -0.000 |
| -404.270 | -495.584 | -494.333 | 0.275 | 0.035 | 0.001 |
| -412.190 | -513.229 | -487.385 | -0.299 | -0.103 | -0.000 |
| -427.202 | -277.017 | -333.154 | -75.990 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 57.597 | -0.000 |
| -448.236 | -368.165 | -207.684 | -0.083 | -0.021 | 8.720 |
| 41.815 | 37.396 | 30.712 | -0.008 | -0.002 | -0.000 |
| 76.692 | 196.168 | 96.111 | -0.000 | 0.000 | -0.000 |
| 95.995 | 106.876 | 144.375 | 0.000 | -0.000 | 0.000 |
| 71.607 | 32.212 | 11.761 | -12.806 | 0.000 | 0.000 |
| 52.750 | 64.529 | 70.770 | 0.000 | 7.744 | 0.000 |
| 34.316 | 41.955 | 90.547 | 0.027 | 0.003 | 4.718 |
| -37.096 | -47.700 | -51.197 | 0.052 | 0.005 | -0.000 |
| -32.687 | -45.090 | -33.854 | -0.004 | 0.002 | -0.000 |
| -61.657 | -48.430 | -51.976 | 0.051 | 0.005 | 0.000 |
| -71.607 | -32.212 | -11.761 | -12.806 | -0.000 | -0.000 |
| -52.750 | -64.529 | -70.770 | -0.000 | 7.744 | -0.000 |
| -34.317 | -41.959 | -90.543 | -0.027 | -0.003 | 4.719 |
| 665.569 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 665.569 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 665.569 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 665.569 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 665.571 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 665.571 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 665.566 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 665.566 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 131.148 | -293.642 | -87.512 | -0.001 | 186.088 | 0.001 |
| 32.317 | 148.244 | 49.246 | 0.000 | -11.611 | -0.000 |
| -10.572 | -319.915 | 73.970 | -0.002 | 169.989 | 0.001 |
| 199.970 | -271.387 | 45.514 | 18.664 | 181.777 | -3.313 |
| 160.159 | -273.236 | -67.670 | -0.001 | 184.002 | 0.001 |
| 3.854 | -310.102 | 54.248 | 4.972 | 172.114 | -6.555 |
| 83.386 | 32.300 | 40.498 | -0.000 | 10.520 | -0.000 |
| 32.317 | 148.244 | 49.246 | 0.000 | -11.611 | 0.000 |
| -177.062 | 302.344 | 38.656 | 0.004 | -188.603 | -0.001 |
| -199.970 | 271.387 | -45.514 | 18.672 | -181.777 | -3.315 |
| -168.224 | 279.502 | 59.741 | 0.001 | -198.418 | -0.001 |
| -3.854 | 310.102 | -54.248 | 4.975 | -172.114 | -6.558 |
| 11.563 | -32.310 | 1.773 | -0.000 | 16.351 | 0.000 |
| 6.290 | -32.899 | -14.180 | -0.000 | 17.290 | 0.000 |
| 8.561 | -31.721 | 33.554 | -0.000 | 19.549 | 0.000 |
| 17.752 | -24.092 | 4.040 | 25.187 | 16.137 | -2.402 |
| 10.455 | -11.611 | 18.423 | -0.000 | 44.378 | -0.000 |
| -0.000 | 0.000 | 0.000 | 4.803 | -0.000 | 36.323 |
| -28.502 | 10.361 | -13.273 | 0.000 | -19.352 | -0.000 |
| -17.461 | 33.104 | 0.194 | 0.000 | -20.814 | -0.000 |
| 3.771 | 59.282 | 85.203 | 0.000 | -23.703 | -0.000 |
| -2.417 | 28.783 | -5.015 | 8.445 | -16.301 | 4.095 |
| -19.504 | 28.500 | 2.358 | 0.000 | -22.644 | -0.000 |
| 0.000 | 0.000 | -0.000 | 4.803 | 0.000 | 36.323 |
| 1.866 | -3.828 | 1.059 | -0.000 | 1.805 | 0.000 |
| -1.369 | 8.499 | -1.676 | -0.000 | 1.011 | 0.000 |
| 3.511 | -1.800 | 33.921 | -0.000 | 3.650 | 0.000 |
| 1.432 | -2.547 | -0.301 | 11.837 | 1.718 | 2.145 |
| 1.055 | -2.050 | -3.946 | -0.000 | 8.351 | 0.000 |
| 1.118 | -2.237 | -0.184 | 2.724 | 1.477 | 2.879 |
| 5.338 | 7.412 | 4.041 | 0.000 | -3.828 | -0.000 |
| -2.527 | 5.463 | -0.093 | 0.000 | -2.359 | -0.000 |
| -7.845 | -1.171 | 37.282 | 0.000 | -2.423 | -0.000 |
| -1.432 | 2.547 | 0.301 | 11.837 | -1.718 | 2.145 |
| -0.408 | 3.353 | -4.904 | 0.000 | 4.900 | -0.000 |
| -1.118 | 2.237 | 0.184 | 2.724 | -1.477 | 2.879 |
| 42.015 | 26.007 | 36.993 | -0.000 | -0.013 | 0.000 |
| 159.118 | 536.364 | 399.949 | -0.000 | 103.431 | 0.000 |
| 293.254 | 498.950 | 337.115 | 0.000 | 120.637 | 0.000 |
| 142.815 | 506.055 | 377.581 | 7.316 | 102.971 | 0.007 |
| 235.898 | 442.622 | 259.240 | -0.000 | 119.271 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.014 | 0.000 | 16.549 |
| 42.015 | 26.007 | 36.993 | 0.000 | -0.013 | 0.000 |
| -231.232 | -433.680 | -253.480 | 0.000 | -116.202 | -0.000 |
| -232.937 | -436.369 | -239.059 | 0.000 | -116.193 | -0.000 |
| -142.815 | -506.055 | -377.581 | 7.316 | -102.971 | 0.007 |
| -306.958 | -514.468 | -340.448 | -0.000 | -118.855 | -0.000 |
| -62.464 | 411.103 | -174.565 | -0.018 | -93.429 | 8.620 |
| 35.478 | 66.324 | 58.745 | -0.000 | 10.289 | 0.000 |
| 27.580 | 51.856 | 30.444 | -0.000 | 11.542 | 0.000 |
| 25.923 | 49.431 | 45.372 | -0.000 | 11.452 | 0.000 |
| 23.564 | 44.189 | 25.873 | 3.574 | 11.600 | -0.001 |
| 23.641 | 44.181 | 25.818 | -0.000 | 14.708 | 0.000 |
| 14.313 | 50.717 | 37.841 | 0.007 | 10.320 | 6.625 |
| 6.123 | -36.254 | -17.981 | 0.000 | -10.536 | -0.000 |
| -18.981 | -34.874 | -19.574 | 0.000 | -11.682 | -0.000 |
| -20.983 | -38.277 | -6.181 | 0.000 | -11.745 | -0.000 |
| -23.564 | -44.189 | -25.873 | 3.574 | -11.600 | -0.001 |
| -30.432 | -51.254 | -33.899 | -0.000 | -10.228 | -0.000 |
| -14.313 | -50.717 | -37.841 | 0.007 | -10.320 | 6.625 |
| 5.897 | 6.729 | 2.662 | -0.000 | 1.070 | 0.000 |
| 2.972 | 9.674 | 4.147 | -0.000 | 0.971 | 0.000 |
| 2.596 | 9.099 | 21.487 | -0.000 | 1.114 | 0.000 |
| 2.307 | 3.966 | 2.200 | 4.005 | 1.086 | -0.001 |
| 2.260 | 4.245 | 2.489 | -0.000 | 5.029 | 0.000 |
| 2.363 | 4.370 | 2.538 | -0.001 | 1.155 | 2.717 |
| 11.176 | 4.738 | 9.575 | 0.000 | -1.432 | -0.000 |
| 0.216 | 1.988 | 2.068 | 0.000 | -1.238 | -0.000 |
| -3.717 | -6.809 | 11.821 | -0.000 | -1.125 | -0.000 |
| -2.307 | -3.966 | -2.200 | 4.005 | -1.086 | -0.001 |
| -2.702 | -4.915 | -3.223 | -0.000 | 1.773 | -0.000 |
| -2.363 | -4.370 | -2.538 | -0.001 | -1.155 | 2.717 |
| 212.210 | 262.242 | 262.242 | -0.000 | -0.000 | -0.000 |
| 262.242 | 212.210 | 262.242 | -0.000 | 0.000 | -0.000 |
| 262.242 | 262.242 | 212.210 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 92.258 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 92.258 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 92.258 |
| 444.833 | 172.900 | 172.900 | 0.000 | -0.000 | -0.000 |
| 172.900 | 444.833 | 172.900 | 0.000 | -0.000 | -0.000 |
| 172.900 | 172.900 | 444.833 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 92.258 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 92.258 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 92.258 |
| 444.833 | 172.900 | 172.900 | -0.000 | 0.000 | 0.000 |
| 172.900 | 444.833 | 172.900 | -0.000 | 0.000 | 0.000 |
| 172.900 | 172.900 | 444.833 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 92.258 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 92.258 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 92.258 |
| 444.833 | 172.900 | 172.900 | -0.000 | 0.000 | -0.000 |
| 172.900 | 444.833 | 172.900 | 0.000 | 0.000 | 0.000 |
| 172.900 | 172.900 | 444.833 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 92.258 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 92.258 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 92.258 |
| 26.239 | 363.198 | 363.198 | -0.000 | 0.000 | -0.000 |
| 363.198 | 26.239 | 363.198 | -0.000 | 0.000 | -0.000 |
| 363.198 | 363.198 | 26.239 | 0.000 | 0.000 | -0.000 |
| 3367924.608 | 1392128.748 | 1392122.879 | -2934543.211 | 0.000 | 0.000 |
| 1392128.728 | 3367924.611 | 1392122.859 | -0.000 | -2934543.221 | -0.000 |
| 1392128.718 | 1392122.849 | 3367924.658 | -0.000 | -0.000 | -2934543.226 |
| 26.145 | 363.231 | 363.231 | 0.000 | -0.000 | -0.000 |
| 363.231 | 26.145 | 363.231 | -0.000 | -0.000 | 0.000 |
| 363.231 | 363.231 | 26.145 | 0.000 | 0.000 | -0.000 |
| -3367924.595 | -1392128.729 | -1392122.861 | -2934543.207 | 0.000 | -0.000 |
| -1392128.739 | -3367924.601 | -1392122.870 | -0.000 | -2934543.238 | 0.000 |
| -1392128.746 | -1392122.878 | -3367924.595 | -0.000 | 0.000 | -2934543.219 |
| 408.476 | 261.423 | 281.497 | -0.000 | 0.000 | -0.000 |
| 167.000 | 617.775 | 256.659 | -0.000 | -0.000 | -0.000 |
| 277.996 | 256.659 | 541.819 | -0.000 | -0.000 | -0.000 |
| -0.001 | 0.000 | 0.000 | 76.092 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 164.050 | -0.000 |
| -0.001 | 0.000 | 0.000 | -0.000 | 0.000 | 41.345 |
| 391.610 | 262.921 | 309.031 | 0.000 | -0.000 | 0.000 |
| 264.742 | 467.565 | 307.102 | 0.000 | -0.000 | 0.000 |
| 277.996 | 256.659 | 541.819 | 0.000 | -0.000 | 0.000 |
| 0.001 | -0.000 | -0.000 | 76.092 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 164.050 | 0.000 |
| 0.001 | -0.000 | -0.000 | 0.000 | -0.000 | 41.345 |
| 408.475 | 261.423 | 281.497 | -0.000 | 0.000 | -0.000 |
| 265.104 | 467.394 | 307.678 | -0.000 | 0.000 | -0.000 |
| 277.996 | 256.659 | 541.819 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 202.487 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 88.489 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 41.345 |
| 408.452 | 261.432 | 281.497 | 0.000 | 0.000 | 0.000 |
| 167.000 | 617.775 | 256.659 | 0.000 | 0.000 | 0.000 |
| 291.088 | 310.983 | 440.955 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 202.487 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 88.489 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 41.345 |
| 385.161 | 260.962 | 300.316 | 0.000 | 0.000 | -0.000 |
| 264.742 | 467.566 | 307.103 | -0.000 | 0.000 | -0.000 |
| 281.153 | 309.964 | 449.184 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 76.092 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 90.119 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 83.628 |
| 408.489 | 261.417 | 281.496 | 0.000 | -0.000 | 0.000 |
| 263.879 | 467.445 | 306.973 | 0.000 | -0.000 | 0.000 |
| 305.078 | 307.532 | 444.875 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 76.092 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 90.119 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 83.628 |
| -259.114 | 236.697 | 292.991 | -0.000 | 0.000 | 0.000 |
| 173.676 | 627.879 | 264.752 | -0.000 | -0.000 | 0.000 |
| -223.627 | 254.103 | 355.311 | -0.000 | 0.000 | 0.000 |
| -477.550 | -124.364 | 36.553 | -89.021 | -0.000 | 0.000 |
| 940.452 | -680.673 | -57.156 | -0.000 | 19.778 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 80.799 |
| 623.294 | 178.208 | 255.381 | 0.000 | -0.000 | -0.000 |
| 173.676 | 627.879 | 264.752 | -0.000 | 0.000 | -0.000 |
| 255.204 | 269.107 | 521.575 | -0.000 | 0.000 | -0.000 |
| 477.550 | 124.364 | -36.553 | -89.021 | -0.000 | 0.000 |
| -940.452 | 680.673 | 57.156 | 0.000 | 19.778 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 80.799 |
| 623.294 | 178.208 | 255.381 | 0.000 | 0.000 | 0.000 |
| 288.433 | 447.708 | 293.936 | -0.000 | 0.000 | 0.000 |
| 255.204 | 269.107 | 521.575 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 206.855 | -0.000 | 0.000 |
| 108.197 | -68.776 | -7.975 | -0.000 | 93.316 | 0.000 |
| -37.397 | -9.739 | 2.862 | -0.000 | 0.000 | 6.020 |
| 339.986 | 308.512 | 316.777 | -0.000 | 0.000 | -0.000 |
| 230.195 | 532.406 | 293.193 | 0.000 | -0.000 | -0.000 |
| 125.151 | 375.568 | 428.635 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 206.855 | -0.000 | -0.000 |
| -108.197 | 68.776 | 7.975 | 0.000 | 93.316 | -0.000 |
| 37.397 | 9.739 | -2.862 | 0.000 | 0.000 | 6.020 |
| 392.462 | 268.411 | 257.443 | -0.000 | 0.000 | 0.000 |
| 271.633 | 475.311 | 310.849 | -0.000 | -0.000 | 0.000 |
| 277.273 | 309.634 | 428.043 | 0.000 | 0.000 | 0.000 |
| -2.108 | -0.549 | 0.161 | 76.252 | 0.000 | 0.000 |
| 5.251 | -4.027 | 1.443 | 0.000 | 94.771 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 80.799 |
| 368.155 | 273.315 | 276.498 | -0.000 | -0.000 | -0.000 |
| 265.110 | 482.435 | 307.414 | 0.000 | 0.000 | -0.000 |
| 256.499 | 325.930 | 419.817 | 0.000 | 0.000 | -0.000 |
| 2.108 | 0.549 | -0.161 | 76.252 | -0.000 | 0.000 |
| -5.251 | 4.027 | -1.443 | -0.000 | 94.771 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 80.799 |
| -36.888 | -45.854 | -45.836 | 1.791 | 1.627 | 1.487 |
| 74.125 | 56.823 | 74.283 | -0.368 | -0.353 | -0.398 |
| 110.673 | 110.762 | 230.047 | 0.058 | 0.079 | 0.071 |
| 43.722 | 44.025 | 44.061 | -5.896 | -0.020 | 0.032 |
| 10.326 | 10.627 | 10.671 | 0.359 | 0.159 | 0.393 |
| 8.057 | 7.925 | 7.911 | 0.101 | 0.127 | -12.883 |
| -34.921 | -78.248 | -78.279 | 0.083 | 0.010 | -0.018 |
| -20.327 | -30.896 | -20.528 | -0.094 | -0.113 | -0.121 |
| 52.707 | 52.564 | 35.830 | -0.912 | -1.035 | -1.036 |
| 0.035 | -0.013 | -0.019 | 180.438 | -0.042 | -0.038 |
| 0.035 | -0.013 | -0.019 | -0.031 | 180.407 | -0.038 |
| -12.898 | -13.030 | -13.046 | 0.103 | 0.059 | -22.815 |
| 2.849 | 2.458 | 2.462 | 0.106 | 0.066 | 0.099 |
| -0.998 | 4.829 | -0.979 | 0.031 | 0.037 | 0.036 |
| -3.721 | -3.695 | 3.195 | 0.127 | 0.157 | 0.135 |
| 0.015 | 0.032 | 0.034 | 62.341 | 0.029 | 0.027 |
| 0.015 | 0.032 | 0.034 | 0.023 | 62.347 | 0.027 |
| 0.015 | 0.032 | 0.034 | 0.023 | 0.029 | 62.345 |
| 0.468 | 0.308 | 0.306 | -0.034 | -0.040 | -0.040 |
| -1.099 | 0.784 | -1.117 | 0.504 | 0.564 | 0.576 |
| -1.099 | -1.116 | 0.782 | 0.504 | 0.564 | 0.576 |
| -0.605 | -0.572 | -0.568 | 6.545 | -0.020 | -0.019 |
| -2.152 | -2.210 | -2.218 | -0.038 | -0.077 | -0.043 |
| -1.087 | -1.116 | -1.122 | -0.037 | -0.049 | -0.094 |
| 1.077 | -0.491 | -0.490 | -0.008 | -0.009 | -0.009 |
| -0.379 | 1.806 | -0.377 | -0.019 | -0.020 | -0.021 |
| 0.085 | 0.088 | 0.182 | 0.003 | 0.004 | 0.003 |
| 0.098 | 0.101 | 0.101 | 0.846 | 0.001 | 0.001 |
| 0.076 | 0.120 | 0.084 | 0.001 | 0.666 | 0.001 |
| 0.069 | 0.072 | 0.073 | 0.003 | 0.005 | 0.000 |
| 1.210 | -0.311 | -0.311 | -0.019 | -0.021 | -0.022 |
| 0.069 | -0.080 | 0.065 | 0.007 | 0.006 | 0.007 |
| -0.310 | -0.311 | 1.208 | -0.019 | -0.021 | -0.021 |
| -0.227 | -0.231 | -0.232 | -0.325 | -0.002 | -0.001 |
| -0.100 | -0.100 | -0.102 | -0.001 | 0.890 | -0.001 |
| -0.033 | -0.037 | -0.036 | -0.003 | -0.005 | -0.007 |
| 1019.646 | 397.663 | 397.663 | 0.000 | 0.000 | 0.000 |
| 397.663 | 1019.646 | 397.663 | 0.000 | 0.000 | -0.000 |
| 397.663 | 397.663 | 1019.646 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 306.882 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 306.882 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 306.882 |
| 648.488 | 280.135 | 280.135 | 0.000 | -0.000 | -0.000 |
| 280.135 | 648.488 | 280.135 | 0.000 | -0.000 | -0.000 |
| 280.135 | 280.135 | 648.488 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 306.882 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 306.882 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 306.882 |
| 1019.646 | 397.663 | 397.663 | 0.000 | 0.000 | 0.000 |
| 397.663 | 1019.646 | 397.663 | 0.000 | 0.000 | 0.000 |
| 397.663 | 397.663 | 1019.646 | 0.000 | 0.000 | 0.000 |
| 0.089 | 0.089 | 0.090 | 118.913 | 0.000 | 0.000 |
| 0.090 | 0.089 | 0.089 | -0.000 | 118.913 | 0.000 |
| 0.089 | 0.090 | 0.089 | 0.000 | 0.000 | 118.913 |
| 487.191 | 138.053 | 138.053 | -0.000 | -0.000 | -0.000 |
| 138.053 | 487.191 | 138.053 | -0.000 | 0.000 | -0.000 |
| 138.053 | 138.053 | 487.191 | -0.000 | -0.000 | -0.000 |
| -0.089 | -0.089 | -0.090 | 118.913 | -0.000 | -0.000 |
| -0.090 | -0.089 | -0.089 | -0.000 | 118.913 | -0.000 |
| -0.089 | -0.090 | -0.089 | -0.000 | -0.000 | 118.913 |
| 649.290 | 280.445 | 280.446 | 0.000 | 0.000 | 0.000 |
| 280.446 | 649.290 | 280.445 | 0.000 | 0.000 | 0.000 |
| 280.445 | 280.446 | 649.290 | 0.000 | -0.000 | 0.000 |
| 0.001 | 0.001 | 0.002 | 197.786 | 0.000 | 0.000 |
| 0.007 | 0.008 | 0.008 | 0.000 | 128.269 | 0.000 |
| 0.001 | 0.002 | 0.001 | 0.000 | 0.000 | 197.786 |
| 388.785 | 136.267 | 136.267 | -0.000 | -0.000 | -0.000 |
| 136.267 | 388.785 | 136.267 | -0.000 | -0.000 | -0.000 |
| 136.267 | 136.267 | 388.785 | -0.000 | -0.000 | -0.000 |
| -0.001 | -0.001 | -0.002 | 197.786 | -0.000 | -0.000 |
| -0.007 | -0.008 | -0.008 | -0.000 | 128.269 | -0.000 |
| -0.001 | -0.002 | -0.001 | -0.000 | -0.000 | 197.786 |
| 952.365 | 904.296 | 904.295 | -0.000 | 0.000 | -0.000 |
| 904.295 | 952.365 | 904.296 | -0.000 | 0.000 | -0.000 |
| 904.296 | 904.295 | 952.365 | 0.000 | 0.000 | -0.000 |
| 243.334 | 243.334 | 243.334 | -16.559 | 0.000 | -0.000 |
| 243.334 | 243.334 | 243.334 | -0.000 | -16.559 | 0.000 |
| 243.334 | 243.334 | 243.334 | 0.000 | 0.000 | -16.559 |
| -434.688 | -434.159 | -434.159 | 0.000 | -0.000 | -0.000 |
| -434.159 | -434.688 | -434.159 | 0.000 | -0.000 | 0.000 |
| -434.159 | -434.159 | -434.688 | 0.000 | -0.000 | -0.000 |
| -243.334 | -243.334 | -243.334 | -16.559 | -0.000 | 0.000 |
| -243.334 | -243.334 | -243.334 | -0.000 | -16.559 | 0.000 |
| -243.334 | -243.334 | -243.334 | 0.000 | -0.000 | -16.559 |
| 109.340 | 93.653 | 93.653 | -0.000 | 0.000 | -0.000 |
| 93.653 | 109.340 | 93.653 | 0.000 | 0.000 | -0.000 |
| 93.653 | 93.653 | 109.340 | -0.000 | 0.000 | -0.000 |
| 24.486 | 24.486 | 24.486 | -16.449 | 0.000 | 0.000 |
| 24.486 | 24.486 | 24.486 | -0.000 | -16.449 | 0.000 |
| 24.486 | 24.486 | 24.486 | -0.000 | 0.000 | -16.449 |
| 2.969 | -31.973 | -31.973 | -0.000 | -0.000 | 0.000 |
| -31.973 | 2.969 | -31.973 | 0.000 | -0.000 | 0.000 |
| -31.973 | -31.973 | 2.969 | -0.000 | -0.000 | 0.000 |
| -24.486 | -24.486 | -24.486 | -16.449 | -0.000 | 0.000 |
| -24.486 | -24.486 | -24.486 | 0.000 | -16.449 | -0.000 |
| -24.486 | -24.486 | -24.486 | 0.000 | -0.000 | -16.449 |
| -32.026 | -60.004 | -60.004 | 0.000 | 0.000 | -0.000 |
| 23.241 | 38.625 | 23.241 | 0.000 | 0.000 | 0.000 |
| -60.004 | -60.004 | -32.026 | -0.000 | -0.000 | -0.000 |
| -1.902 | -1.902 | -1.902 | 17.272 | 0.000 | -0.000 |
| -1.902 | -1.902 | -1.902 | -0.000 | 17.272 | 0.000 |
| -1.902 | -1.902 | -1.902 | 0.000 | 0.000 | 17.272 |
| 2.539 | -28.337 | -28.337 | 0.000 | 0.000 | 0.000 |
| -28.337 | 2.539 | -28.337 | 0.000 | -0.000 | 0.000 |
| -28.337 | -28.337 | 2.539 | 0.000 | 0.000 | 0.000 |
| 1.902 | 1.902 | 1.902 | 17.272 | -0.000 | 0.000 |
| 1.902 | 1.902 | 1.902 | -0.000 | 17.272 | 0.000 |
| 1.902 | 1.902 | 1.902 | -0.000 | -0.000 | 17.272 |
| -2130050.007 | 11385301.797 | -195954975.864 | -3438923.863 | 36851.744 | -2289555.472 |
| -2154299.080 | 11278338.244 | -195829738.648 | -3389711.607 | 24899.765 | -2429107.211 |
| -2131931.908 | 11385342.596 | -195953461.566 | -3439413.648 | 37668.569 | -2292693.248 |
| -2130878.168 | 11382019.744 | -195951467.256 | -3439684.022 | 36972.200 | -2294213.599 |
| -2130837.491 | 11383881.104 | -195953116.223 | -3438630.976 | 37273.427 | -2292810.137 |
| -2131030.938 | 11383758.811 | -195952743.598 | -3438709.032 | 37288.910 | -2293218.142 |
| 2160471.876 | -11293083.334 | 195835498.846 | 3395130.542 | -25628.450 | 2427247.532 |
| 2129895.217 | -11385694.282 | 195955495.851 | 3439078.745 | -36537.568 | 2288266.820 |
| 2154217.868 | -11393160.415 | 195937094.579 | 3439642.653 | -36374.918 | 2320780.360 |
| 2166822.365 | -11305154.102 | 195839916.052 | 3401303.533 | -27593.468 | 2432110.080 |
| 2130585.815 | -11385076.437 | 195954371.416 | 3438596.064 | -37141.170 | 2291053.452 |
| 2130416.382 | -11385072.719 | 195954587.582 | 3438535.068 | -37136.292 | 2290850.100 |
| -215425.212 | 1138908.191 | -19592742.442 | -344481.824 | 3494.750 | -231963.637 |
| -215985.102 | 1124602.384 | -19580457.108 | -337388.168 | 2785.445 | -245659.406 |
| -223252.928 | 1141616.747 | -19588632.182 | -344422.003 | 3811.975 | -241223.830 |
| -191341.729 | 1125315.690 | -19607095.093 | -336922.744 | 3789.551 | -211887.921 |
| -214402.287 | 1138560.511 | -19594166.144 | -344097.319 | 3748.488 | -231170.589 |
| -215833.554 | 1139160.970 | -19593129.295 | -344314.189 | 3698.205 | -232742.449 |
| 217331.944 | -1139701.188 | 19592343.956 | 344074.453 | -3878.602 | 235405.554 |
| 180265.948 | -1119023.096 | 19612932.749 | 331287.323 | -3779.056 | 203201.938 |
| 213622.419 | -1139644.034 | 19594736.734 | 344476.282 | -3012.946 | 230263.113 |
| 221146.616 | -1137567.417 | 19585529.057 | 342155.602 | -3119.313 | 243429.782 |
| 217316.883 | -1132084.633 | 19584514.728 | 340722.990 | -2744.322 | 242842.723 |
| 217076.821 | -1131834.077 | 19584551.649 | 340676.763 | -2760.639 | 242724.077 |
| -20317.279 | 113458.969 | -1960131.405 | -34316.244 | 295.015 | -21797.886 |
| -21068.776 | 111153.677 | -1958127.179 | -34137.820 | 168.276 | -24333.369 |
| -15627.633 | 108791.887 | -1960295.509 | -33090.755 | 133.383 | -21274.731 |
| -17868.517 | 110577.807 | -1959730.322 | -34040.513 | -343.899 | -22718.706 |
| -22200.933 | 113938.518 | -1958833.336 | -34301.598 | 454.877 | -24282.865 |
| -22181.542 | 114024.898 | -1958887.826 | -34278.162 | 473.481 | -24196.600 |
| 19257.723 | -108640.080 | 1957649.095 | 33583.681 | 94.236 | 22333.898 |
| 19822.616 | -112791.120 | 1960585.224 | 33857.378 | -243.480 | 20901.225 |
| 22228.185 | -114382.498 | 1959686.640 | 33842.815 | -210.163 | 24100.326 |
| 21132.332 | -112209.440 | 1958160.602 | 33885.311 | -221.883 | 24052.851 |
| 20424.352 | -113523.660 | 1960167.120 | 34278.621 | -332.929 | 21929.357 |
| 20598.385 | -113644.966 | 1960151.840 | 34335.451 | -285.788 | 22058.699 |
| 183343.622 | 1887852.156 | 4177744.815 | -1069025.947 | -3185534.800 | -2093944.305 |
| 195726.838 | 1919664.704 | 4183779.585 | -1060226.007 | -3169859.289 | -2106393.006 |
| 194153.459 | 1909098.328 | 4183843.551 | -1063573.073 | -3176150.215 | -2100325.800 |
| 198115.533 | 1912994.784 | 4191006.924 | -1061337.260 | -3177025.921 | -2102970.620 |
| 188621.209 | 1894419.692 | 4185413.337 | -1069077.663 | -3182574.813 | -2090806.745 |
| 197069.003 | 1918910.947 | 4183873.444 | -1060812.565 | -3169522.911 | -2104751.347 |
| -200156.271 | -1928663.964 | -4197703.233 | 1060093.640 | 3167810.432 | 2113577.579 |
| -184295.881 | -1894819.386 | -4188067.569 | 1068522.841 | 3187632.827 | 2098049.438 |
| -205939.209 | -1941072.128 | -4187624.596 | 1054335.250 | 3157842.145 | 2114837.951 |
| -204612.044 | -1935577.511 | -4189473.591 | 1053519.539 | 3160812.433 | 2112897.627 |
| -180994.026 | -1887335.673 | -4183450.037 | 1073246.185 | 3196186.163 | 2092552.197 |
| -201866.288 | -1932011.426 | -4184707.927 | 1056829.092 | 3157359.191 | 2108082.362 |
| 20294.208 | 193608.938 | 418730.504 | -105473.074 | -316143.992 | -211268.561 |
| 20663.585 | 194230.209 | 418884.348 | -105328.969 | -315782.369 | -211503.921 |
| 19044.942 | 189804.647 | 418418.556 | -106442.097 | -318108.263 | -209812.827 |
| 20682.996 | 194246.865 | 418852.751 | -105321.602 | -315743.686 | -211526.953 |
| 20099.138 | 192388.478 | 417953.073 | -105508.611 | -316669.730 | -210637.935 |
| 20153.725 | 192437.277 | 418332.725 | -105655.167 | -317138.942 | -210997.388 |
| -20077.019 | -192685.616 | -418989.869 | 105886.133 | 316936.588 | 210870.621 |
| -19946.258 | -192587.594 | -418956.754 | 105770.432 | 317076.242 | 210747.086 |
| -19911.357 | -192209.582 | -418296.781 | 105878.309 | 316733.634 | 210705.039 |
| -19878.452 | -192199.408 | -418469.105 | 105950.997 | 316944.672 | 210489.755 |
| -19350.815 | -190738.056 | -417643.673 | 105990.099 | 316876.549 | 209972.763 |
| -19560.817 | -191720.051 | -418244.308 | 106664.645 | 317023.357 | 210669.382 |
| 2044.459 | 19368.460 | 41854.837 | -10539.713 | -31619.133 | -21127.801 |
| 1838.843 | 18787.949 | 41752.708 | -10698.067 | -31909.730 | -20910.627 |
| 1967.661 | 19148.634 | 42137.519 | -10670.346 | -31918.844 | -21108.780 |
| 1973.985 | 19268.259 | 41820.533 | -10573.500 | -31652.786 | -21073.771 |
| 1955.424 | 18993.872 | 41967.674 | -10649.228 | -31834.676 | -20980.845 |
| 2060.540 | 19417.068 | 41835.709 | -10540.053 | -31573.582 | -21147.029 |
| -1839.390 | -18831.770 | -41825.382 | 10700.077 | 31865.253 | 20914.289 |
| -2024.332 | -19317.969 | -41848.947 | 10546.603 | 31594.560 | 21075.971 |
| -2053.189 | -19359.539 | -41877.528 | 10535.398 | 31607.524 | 21130.910 |
| -1975.430 | -19240.932 | -41862.198 | 10561.810 | 31677.992 | 21085.400 |
| -1778.758 | -18680.386 | -41728.348 | 10780.867 | 31945.990 | 20849.239 |
| -1934.839 | -19188.491 | -41830.539 | 10620.486 | 31634.104 | 21050.274 |
| 2911818.397 | -10963793.975 | 17836874.115 | 191582.338 | 632318.212 | -38216.757 |
| 1817981.149 | -10454090.690 | 18292359.268 | 111421.041 | 747604.528 | -434968.643 |
| 2646662.232 | -10766400.002 | 17908934.082 | 143532.411 | 347137.493 | 74844.830 |
| 3399563.794 | -10951383.583 | 18432832.008 | 333296.323 | 544130.447 | -169966.556 |
| 2040679.799 | -10603580.453 | 18225500.193 | 341898.933 | 245733.492 | -303225.044 |
| 2643985.556 | -10769377.952 | 17894127.057 | 134273.353 | 440686.440 | -79815.868 |
| -2561975.873 | 10717534.556 | -17723163.382 | -371124.086 | -444905.220 | -84164.231 |
| -2191170.331 | 10619003.662 | -18088061.911 | -183210.211 | -649060.012 | 135198.203 |
| -2978110.166 | 10990690.386 | -18164139.770 | -290842.261 | -179748.721 | 211804.660 |
| -2022285.060 | 10546854.808 | -17962828.212 | -347390.427 | -533428.450 | -31252.720 |
| -2630658.728 | 10852444.275 | -18186243.439 | -384308.877 | -167014.397 | 356650.343 |
| -2440538.019 | 10245356.222 | -17590787.560 | -144054.514 | -104323.589 | -215757.520 |
| 330423.410 | -1114411.781 | 1823693.510 | 37175.343 | 34731.159 | -6628.660 |
| 216539.244 | -1054409.683 | 1783657.209 | 30043.428 | 34459.967 | 23271.484 |
| 253914.614 | -1141124.927 | 1896332.391 | 16681.283 | 53647.515 | 6765.786 |
| 312603.330 | -1161557.637 | 1806203.931 | 13388.662 | 27671.427 | -21774.752 |
| 260958.443 | -1072041.672 | 1790358.565 | 14604.835 | 39460.265 | 1908.671 |
| 267138.860 | -1072231.967 | 1792103.923 | 14152.320 | 44711.702 | -7665.177 |
| -258033.140 | 1071327.245 | -1791383.623 | -15666.196 | -34918.748 | -11159.353 |
| -294103.994 | 1146676.305 | -1801625.358 | -14883.984 | -48989.922 | 39045.346 |
| -272379.323 | 1069440.082 | -1781600.812 | -33094.242 | -56188.566 | -9582.859 |
| -308577.035 | 1166197.556 | -1819765.613 | -21369.405 | -26943.448 | 52799.092 |
| -317958.957 | 1103532.366 | -1826157.192 | -40441.025 | -41184.953 | 18696.130 |
| -361544.385 | 1154201.404 | -1830602.578 | -28074.117 | -7414.028 | 18585.692 |
| 31153.440 | -107962.018 | 182773.261 | 2004.440 | 3404.184 | -2192.956 |
| 25274.819 | -103771.775 | 179679.992 | 2486.519 | 3790.805 | -181.968 |
| 32984.583 | -108663.042 | 186750.768 | 5584.811 | 2379.603 | -4308.278 |
| 26313.370 | -106526.743 | 180039.643 | 3991.256 | 6179.352 | 924.606 |
| 34167.536 | -113561.652 | 182704.331 | 2656.385 | 2671.268 | 403.987 |
| 28914.257 | -107705.709 | 181934.943 | 2243.731 | 2303.351 | -957.067 |
| -21976.848 | 107167.368 | -178691.338 | -3194.238 | -4021.770 | -388.257 |
| -19685.121 | 106444.009 | -183674.120 | -630.658 | -5296.288 | 3476.717 |
| -26390.293 | 108038.861 | -179003.527 | -1209.586 | -4514.327 | 1033.822 |
| -26207.383 | 107618.368 | -181920.186 | -2325.553 | -1979.904 | 1545.074 |
| -24489.664 | 105733.051 | -177137.146 | -2194.410 | -3935.343 | 810.585 |
| -27944.140 | 110252.345 | -181184.075 | -1376.961 | -2866.410 | 1221.304 |
| 772.715 | 737.203 | -632.220 | -0.000 | -0.000 | 0.000 |
| 737.203 | 772.715 | -632.220 | -0.000 | -0.000 | 0.000 |
| 1209.112 | 1209.112 | -426.263 | 0.000 | -0.000 | 0.000 |
| 722.043 | 722.043 | -663.681 | 7.059 | 0.000 | 0.000 |
| 722.043 | 722.043 | -663.681 | 0.000 | 7.059 | 0.000 |
| 721.998 | 721.998 | -663.639 | -0.000 | -0.000 | 5.099 |
| -1479.112 | -1276.505 | 453.660 | -0.000 | -0.000 | -0.000 |
| -1276.505 | -1479.112 | 453.660 | -0.000 | -0.000 | 0.000 |
| -1135.383 | -1135.383 | 569.378 | -0.000 | -0.000 | 0.000 |
| -722.043 | -722.043 | 663.681 | 7.059 | 0.000 | -0.000 |
| -722.043 | -722.043 | 663.681 | -0.000 | 7.059 | -0.000 |
| -721.998 | -721.998 | 663.639 | 0.000 | 0.000 | 5.099 |
| 122.931 | 88.920 | -35.561 | -0.000 | -0.000 | 0.000 |
| 88.920 | 122.931 | -35.561 | -0.000 | -0.000 | -0.000 |
| 164.113 | 164.113 | 6.588 | 0.000 | 0.000 | 0.000 |
| 136.963 | 136.963 | -37.378 | -17.497 | 0.000 | -0.000 |
| 136.963 | 136.963 | -37.378 | 0.000 | -17.497 | -0.000 |
| 139.193 | 139.193 | -42.376 | 0.000 | 0.000 | -4.879 |
| -22.282 | -58.041 | 98.618 | 0.000 | 0.000 | 0.000 |
| -58.041 | -22.282 | 98.618 | 0.000 | 0.000 | -0.000 |
| -89.274 | -89.274 | 183.235 | 0.000 | 0.000 | -0.000 |
| -136.963 | -136.963 | 37.378 | -17.497 | -0.000 | 0.000 |
| -136.963 | -136.963 | 37.378 | -0.000 | -17.497 | 0.000 |
| -139.193 | -139.193 | 42.376 | -0.000 | 0.000 | -4.879 |
| 46.802 | 39.973 | 20.839 | -0.000 | -0.000 | -0.000 |
| 39.973 | 46.802 | 20.839 | -0.000 | -0.000 | 0.000 |
| 39.770 | 39.770 | 38.064 | -0.000 | -0.000 | 0.000 |
| 8.398 | 8.398 | -7.719 | 5.120 | -0.000 | 0.000 |
| 8.398 | 8.398 | -7.719 | 0.000 | 5.120 | 0.000 |
| 13.733 | 13.733 | -4.333 | 0.000 | 0.000 | 3.020 |
| 31.248 | 18.060 | 41.376 | 0.000 | 0.000 | 0.000 |
| 18.060 | 31.248 | 41.376 | 0.000 | 0.000 | 0.000 |
| 17.882 | 17.882 | 49.225 | -0.000 | -0.000 | 0.000 |
| -8.398 | -8.398 | 7.719 | 5.120 | 0.000 | 0.000 |
| -8.398 | -8.398 | 7.719 | 0.000 | 5.120 | -0.000 |
| -13.733 | -13.733 | 4.333 | -0.000 | -0.000 | 3.020 |
| 248.933 | 247.088 | -35.509 | -85.005 | 84.550 | 100.637 |
| 543.674 | 461.666 | 347.297 | -121.225 | -98.040 | 120.999 |
| 497.962 | 391.317 | 332.957 | -161.260 | -60.726 | 136.060 |
| 482.148 | 465.975 | 266.057 | -114.143 | -113.394 | 138.231 |
| 317.661 | 328.381 | 170.282 | -182.536 | -24.658 | 187.328 |
| 536.657 | 432.828 | 341.653 | -119.829 | -95.542 | 130.961 |
| -493.622 | -454.011 | -241.042 | 132.067 | 100.491 | -148.887 |
| -527.365 | -419.787 | -318.176 | 132.549 | 133.681 | -143.277 |
| -138.344 | -76.741 | 111.073 | 78.295 | -101.646 | -58.360 |
| -567.091 | -449.500 | -316.341 | 161.815 | 127.400 | -103.996 |
| -539.369 | -444.239 | -325.938 | 135.628 | 144.871 | -140.582 |
| -375.557 | -271.109 | -172.772 | 136.522 | 92.846 | -94.234 |
| 87.230 | 55.934 | 29.070 | -7.745 | 9.119 | 12.888 |
| 66.257 | 73.133 | 39.447 | -17.648 | -12.849 | 11.749 |
| 45.313 | 38.725 | 60.228 | -16.851 | -5.881 | 12.409 |
| 45.046 | 31.832 | 27.101 | 0.459 | -1.527 | 12.638 |
| 50.784 | 36.510 | 33.653 | -11.295 | 4.008 | 9.993 |
| 55.708 | 42.738 | 33.979 | -12.897 | -11.992 | 24.444 |
| -7.328 | -19.094 | -12.804 | 13.132 | 13.320 | -13.896 |
| -42.084 | -12.449 | -22.961 | 11.932 | 6.033 | -11.565 |
| -32.792 | -27.538 | 24.404 | 13.313 | 10.889 | -11.849 |
| -53.958 | -50.536 | -28.758 | 27.345 | 10.258 | -14.519 |
| -48.732 | -39.821 | -22.040 | 13.170 | 25.091 | -13.004 |
| -16.715 | -16.495 | 13.249 | 17.910 | -3.410 | 6.485 |
| 48.239 | 17.491 | 17.866 | -3.308 | 3.337 | 2.749 |
| 19.723 | 37.826 | 16.312 | -3.977 | -2.504 | 1.256 |
| 19.923 | 17.946 | 47.510 | -3.678 | -0.517 | 2.725 |
| 1.731 | -1.943 | 1.638 | 12.420 | 0.723 | 1.824 |
| 6.853 | 1.323 | 3.942 | -0.927 | 11.318 | -0.804 |
| 4.357 | 4.317 | 3.536 | -0.821 | -1.845 | 13.198 |
| 41.125 | 12.366 | 14.841 | 1.091 | 4.249 | -0.457 |
| 9.567 | 29.041 | 11.392 | -1.771 | -0.618 | -0.904 |
| 12.377 | 12.041 | 44.685 | 0.575 | 1.456 | -0.133 |
| -6.714 | -9.408 | -2.366 | 14.669 | 2.074 | -1.112 |
| -5.296 | -6.488 | -2.181 | 1.719 | 13.017 | -2.805 |
| -3.511 | -2.920 | 0.120 | 2.293 | -0.631 | 9.712 |
| 543.980 | 427.311 | 427.307 | 0.000 | -0.001 | -0.001 |
| 427.292 | 543.999 | 427.307 | 0.000 | -0.001 | -0.001 |
| 427.292 | 427.311 | 543.995 | 0.000 | -0.001 | -0.001 |
| -0.013 | 0.009 | 0.004 | 192.377 | -0.001 | -0.001 |
| 0.000 | 0.000 | 0.000 | -0.000 | 237.627 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 237.627 |
| 614.409 | 392.094 | 392.094 | 0.000 | -0.000 | 0.000 |
| 392.094 | 614.409 | 392.094 | 0.000 | -0.000 | 0.000 |
| 392.094 | 392.094 | 614.409 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 237.627 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 237.627 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 237.627 |
| 614.409 | 392.094 | 392.094 | -0.000 | -0.000 | -0.000 |
| 392.094 | 614.409 | 392.094 | -0.000 | 0.000 | -0.000 |
| 392.094 | 392.094 | 614.409 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 237.627 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 237.627 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 237.627 |
| 614.408 | 392.094 | 392.094 | 0.000 | -0.000 | 0.000 |
| 392.094 | 614.408 | 392.094 | 0.000 | 0.000 | 0.000 |
| 392.094 | 392.094 | 614.408 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 237.627 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 237.627 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 237.627 |
| 543.991 | 427.305 | 427.304 | 0.000 | -0.000 | -0.000 |
| 427.304 | 543.991 | 427.304 | 0.000 | -0.000 | -0.000 |
| 427.304 | 427.305 | 543.991 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 192.370 | -0.000 | -0.000 |
| -0.000 | 0.001 | 0.000 | 0.000 | 192.370 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 192.370 |
| 543.987 | 427.304 | 427.304 | -0.000 | 0.000 | 0.000 |
| 427.304 | 543.987 | 427.304 | -0.000 | 0.000 | 0.000 |
| 427.304 | 427.304 | 543.987 | -0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 192.370 | 0.000 | 0.000 |
| 0.000 | -0.001 | -0.000 | -0.000 | 192.370 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 192.370 |
| 789.342 | 251.880 | 235.056 | 0.000 | -0.000 | -0.000 |
| 251.880 | 789.342 | 235.056 | 0.000 | -0.000 | -0.000 |
| 299.569 | 299.569 | 1082.757 | 0.000 | 0.000 | -0.000 |
| -3.376 | -3.376 | 2.582 | 114.120 | 0.000 | -0.000 |
| -3.376 | -3.376 | 2.582 | 0.000 | 114.120 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 268.731 |
| 789.342 | 251.880 | 235.056 | -0.000 | -0.000 | 0.000 |
| 286.947 | 534.001 | 319.300 | -0.000 | 0.000 | 0.000 |
| 436.027 | 436.027 | 978.381 | -0.000 | 0.000 | 0.000 |
| 3.376 | 3.376 | -2.582 | 114.120 | 0.000 | 0.000 |
| 3.376 | 3.376 | -2.582 | -0.000 | 114.120 | 0.000 |
| 3.341 | 3.341 | -2.555 | -0.000 | -0.000 | 151.627 |
| 789.342 | 251.880 | 235.057 | 0.000 | -0.000 | -0.000 |
| 282.740 | 526.463 | 323.792 | 0.000 | -0.000 | 0.000 |
| 429.236 | 429.236 | 983.576 | 0.000 | -0.000 | -0.000 |
| -0.337 | -0.337 | 0.258 | 114.175 | 0.000 | -0.000 |
| -0.337 | -0.337 | 0.258 | 0.000 | 114.175 | -0.000 |
| -0.334 | -0.334 | 0.255 | 0.000 | -0.000 | 151.682 |
| 527.842 | 283.512 | 322.969 | -0.000 | 0.000 | 0.000 |
| 251.880 | 789.342 | 235.056 | -0.000 | 0.000 | 0.000 |
| 299.569 | 299.569 | 1082.757 | -0.000 | 0.000 | -0.000 |
| 0.337 | 0.337 | -0.258 | 114.175 | -0.000 | -0.000 |
| 0.337 | 0.337 | -0.258 | -0.000 | 114.175 | -0.000 |
| 0.334 | 0.334 | -0.255 | -0.000 | 0.000 | 151.682 |
| 527.077 | 283.090 | 323.425 | 0.000 | -0.000 | -0.000 |
| 198.528 | 501.881 | 365.402 | 0.000 | -0.000 | -0.000 |
| 429.799 | 429.799 | 983.151 | 0.000 | -0.000 | -0.000 |
| -0.034 | -0.034 | 0.026 | 114.175 | -0.000 | -0.000 |
| -0.034 | -0.034 | 0.026 | 0.000 | 114.175 | -0.000 |
| -0.033 | -0.033 | 0.026 | 0.000 | 0.000 | 151.683 |
| 502.035 | 198.660 | 365.289 | -0.000 | 0.000 | 0.000 |
| 198.660 | 502.045 | 365.285 | -0.000 | 0.000 | 0.000 |
| 429.906 | 429.906 | 983.057 | -0.000 | 0.000 | 0.000 |
| 0.034 | 0.034 | -0.026 | 114.175 | 0.000 | 0.000 |
| 0.034 | 0.034 | -0.026 | -0.000 | 114.175 | 0.000 |
| 0.033 | 0.033 | -0.026 | -0.000 | 0.000 | 151.683 |
| 405.327 | 178.701 | 136.059 | 0.000 | -0.000 | -0.000 |
| -45.412 | 40.167 | -246.414 | 0.025 | -0.018 | -0.000 |
| 137.946 | 117.827 | 380.563 | 0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 79.955 | -0.000 | -0.000 |
| -192.700 | -5.086 | -240.014 | -0.014 | 49.190 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 127.879 |
| 419.282 | 143.236 | 387.290 | 0.000 | 0.000 | -0.000 |
| 195.394 | 254.418 | 414.267 | -0.005 | 0.005 | -0.000 |
| 280.815 | 88.502 | 424.882 | 0.014 | -0.008 | 0.000 |
| 231.449 | 210.558 | 505.152 | 28.484 | -0.002 | -0.000 |
| 315.912 | 250.214 | 488.217 | 0.006 | 28.736 | 0.000 |
| 192.718 | 5.086 | 240.035 | 0.014 | -0.008 | 56.267 |
| 246.419 | 110.652 | 71.544 | -0.001 | 0.001 | -0.000 |
| 70.221 | 170.184 | 49.665 | -0.002 | 0.001 | -0.000 |
| 79.943 | 88.600 | 196.491 | -0.001 | 0.001 | -0.000 |
| -17.779 | -17.236 | -38.808 | 31.753 | 0.000 | 0.000 |
| -14.328 | -16.213 | -34.346 | -0.001 | 28.490 | -0.000 |
| -18.381 | -0.485 | -22.894 | -0.001 | 0.001 | 59.576 |
| 238.221 | 89.593 | 118.739 | 0.000 | -0.000 | -0.000 |
| 176.591 | 307.651 | 113.831 | -0.000 | -0.000 | 0.000 |
| 113.267 | 90.598 | 242.514 | 0.001 | -0.001 | 0.000 |
| 15.092 | 0.398 | 18.797 | 43.021 | -0.001 | 0.000 |
| 17.671 | 0.466 | 22.010 | 0.001 | 53.406 | 0.000 |
| 18.581 | 4.795 | 26.961 | 0.000 | -0.000 | 45.247 |
| 154.986 | 46.400 | 37.148 | -0.000 | 0.000 | 0.000 |
| 45.769 | 91.724 | 29.302 | -0.001 | 0.000 | 0.000 |
| 43.143 | 33.339 | 117.720 | -0.000 | 0.000 | -0.000 |
| -1.528 | -0.757 | -2.412 | 39.186 | 0.000 | 0.000 |
| -2.237 | -1.329 | -3.365 | -0.000 | 37.033 | -0.000 |
| -1.735 | -0.984 | -3.037 | -0.000 | 0.000 | 42.834 |
| 148.432 | 51.556 | 49.597 | 0.000 | -0.000 | 0.000 |
| 43.059 | 93.938 | 33.910 | -0.000 | 0.000 | 0.000 |
| 45.574 | 34.748 | 122.890 | 0.000 | -0.000 | 0.000 |
| 1.539 | 1.242 | 3.033 | 36.985 | -0.000 | 0.000 |
| 2.237 | 1.329 | 3.365 | 0.000 | 37.033 | 0.000 |
| 2.105 | 1.297 | 3.401 | 0.000 | -0.000 | 41.232 |
| 490.177 | 178.583 | 0.000 | 0.000 | 0.000 | -0.000 |
| 178.583 | 253.959 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 490.177 | 178.583 | -0.000 | -0.000 | -0.000 | -0.000 |
| 178.583 | 253.959 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 490.177 | 178.583 | 0.000 | 0.000 | 0.000 | 0.000 |
| 178.583 | 253.959 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 490.177 | 178.583 | -0.000 | -0.000 | -0.000 | -0.000 |
| 178.583 | 253.959 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 490.179 | 178.584 | 0.000 | 0.000 | 0.000 | -0.000 |
| 178.583 | 253.956 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.001 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 490.175 | 178.583 | -0.000 | -0.000 | -0.000 | -0.000 |
| 178.583 | 253.962 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.001 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 246.861 | 196.446 | 167.878 | 0.000 | -0.000 | -0.000 |
| 106.122 | 609.321 | 234.699 | 0.000 | 0.000 | -0.000 |
| 86.802 | 243.948 | 592.887 | 0.000 | 0.000 | -0.000 |
| -13.633 | 71.790 | -92.436 | 53.578 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 20.424 | -0.000 |
| -15.695 | 82.646 | -106.414 | 0.000 | 0.000 | 14.853 |
| -194.480 | 230.336 | 230.383 | 0.000 | -0.000 | 0.000 |
| 106.122 | 609.321 | 234.699 | -0.000 | -0.000 | 0.000 |
| 86.802 | 243.948 | 592.887 | -0.000 | -0.000 | 0.000 |
| 13.633 | -71.790 | 92.436 | 53.578 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 20.424 | 0.000 |
| 15.695 | -82.646 | 106.414 | 0.000 | -0.000 | 14.853 |
| -25.748 | 293.869 | 105.172 | 0.000 | 0.000 | 0.000 |
| 143.145 | 544.634 | 317.155 | 0.000 | 0.000 | -0.000 |
| 65.096 | 338.122 | 465.050 | 0.000 | -0.000 | -0.000 |
| -197961897.227 | -29698727.137 | -16598508.657 | 4149138.890 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 20.424 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 36.571 |
| 246.860 | 196.446 | 167.877 | -0.000 | 0.000 | 0.000 |
| 144.202 | 535.489 | 328.952 | -0.000 | -0.000 | -0.000 |
| 67.019 | 327.806 | 478.270 | -0.000 | -0.000 | 0.000 |
| 197961896.936 | 29698727.502 | 16598508.662 | 4149139.030 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 20.424 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 36.571 |
| -121.424 | 248.843 | 193.679 | -0.000 | -0.000 | -0.000 |
| 161.286 | 540.455 | 317.973 | -0.000 | 0.000 | -0.000 |
| 105.799 | 328.420 | 467.137 | -0.000 | 0.000 | 0.000 |
| -0.102 | 0.537 | -0.691 | 88.327 | 0.000 | -0.000 |
| -0.141 | 0.742 | -0.956 | 0.000 | 12.910 | -0.000 |
| 0.905 | 0.420 | -0.767 | 0.000 | 0.000 | 12.791 |
| -124.160 | 248.492 | 194.707 | -0.000 | -0.000 | 0.000 |
| 143.157 | 540.608 | 322.360 | -0.000 | -0.000 | 0.000 |
| 101.889 | 327.670 | 469.082 | 0.000 | -0.000 | 0.000 |
| 0.102 | -0.537 | 0.691 | 88.327 | 0.000 | 0.000 |
| 0.141 | -0.742 | 0.956 | -0.000 | 12.910 | 0.000 |
| -0.905 | -0.420 | 0.767 | 0.000 | -0.000 | 12.791 |
| 340.726 | 282.780 | 486.282 | 0.453 | 0.000 | 0.000 |
| 274.680 | 638.011 | 197.256 | -21.831 | -0.000 | -0.000 |
| 260.297 | 260.297 | 588.106 | -0.000 | 0.000 | 0.000 |
| 21.831 | -21.831 | -0.000 | 104.636 | -0.000 | -0.000 |
| -53.304 | -53.304 | 106.257 | -0.000 | 46.341 | 5.349 |
| -69.097 | -69.097 | 137.945 | -0.000 | 0.456 | 28.994 |
| 510.297 | 392.747 | 207.324 | 5.353 | -0.000 | -0.000 |
| 420.975 | 478.962 | 210.352 | -0.456 | -0.000 | 0.000 |
| 417.458 | 417.458 | 274.342 | 0.000 | -0.000 | -0.000 |
| 69.556 | 68.644 | -137.950 | 29.037 | -0.000 | 0.000 |
| 53.304 | 53.304 | -106.257 | 0.000 | 46.341 | 5.349 |
| 69.097 | 69.097 | -137.945 | 0.000 | 0.456 | 28.994 |
| 638.011 | 274.680 | 197.256 | 21.831 | -0.000 | 0.000 |
| 274.680 | 638.011 | 197.256 | -21.831 | -0.000 | 0.000 |
| 308.309 | 308.309 | 492.399 | -0.000 | 0.000 | 0.000 |
| 21.831 | -21.831 | 0.000 | 104.636 | -0.000 | -0.000 |
| -6.909 | -6.909 | 13.794 | -0.000 | 29.043 | 0.457 |
| -5.331 | -5.331 | 10.626 | 0.000 | 5.349 | 58.744 |
| 416.878 | 358.747 | 334.438 | 0.466 | -0.000 | 0.000 |
| 358.744 | 416.885 | 334.434 | -0.467 | -0.000 | 0.000 |
| 319.017 | 319.017 | 471.050 | 0.000 | -0.000 | -0.000 |
| 10.680 | -0.019 | -10.626 | 46.342 | -0.000 | -0.000 |
| 6.909 | 6.909 | -13.794 | 0.000 | 29.043 | 0.457 |
| 5.331 | 5.331 | -10.626 | -0.000 | 5.349 | 58.744 |
| 410.062 | 344.771 | 355.209 | -0.283 | 0.000 | -0.000 |
| 348.324 | 417.737 | 343.975 | -1.256 | 0.000 | -0.000 |
| 340.565 | 340.565 | 427.889 | -0.000 | 0.000 | -0.000 |
| 4.539 | -5.622 | 1.080 | 45.390 | 0.000 | -0.000 |
| -0.680 | -0.680 | 1.358 | -0.000 | 30.247 | 0.798 |
| -0.544 | -0.544 | 1.085 | 0.000 | 5.008 | 56.369 |
| 418.960 | 349.688 | 341.386 | 1.246 | -0.000 | 0.000 |
| 349.687 | 418.963 | 341.384 | -1.247 | -0.000 | 0.000 |
| 341.753 | 341.753 | 425.510 | 0.000 | -0.000 | -0.000 |
| 5.622 | -4.539 | -1.080 | 45.389 | -0.000 | -0.000 |
| 0.680 | 0.680 | -1.358 | 0.000 | 30.247 | 0.798 |
| 0.544 | 0.544 | -1.085 | -0.000 | 5.008 | 56.369 |
| 637.537 | 273.829 | 197.535 | 15.572 | 0.000 | -0.000 |
| -141.377 | -48.479 | 1296.572 | -2.009 | -0.000 | 0.000 |
| 197.433 | 197.433 | 716.524 | 0.000 | -0.000 | -0.000 |
| -402.197 | -410.644 | 811.228 | 48.157 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 104.264 | 15.572 |
| -0.000 | -0.000 | 0.000 | 0.000 | 15.572 | 181.854 |
| 637.537 | 273.829 | 197.535 | 15.572 | -0.000 | 0.000 |
| 273.829 | 637.537 | 197.535 | -15.572 | 0.000 | 0.000 |
| 934.996 | 934.996 | -755.679 | 0.000 | 0.000 | -0.000 |
| 15.572 | -15.572 | -0.000 | 104.264 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 104.264 | 15.572 |
| 0.000 | 0.000 | -0.000 | -0.000 | 15.572 | 181.854 |
| 637.537 | 273.829 | 197.535 | 15.572 | -0.000 | -0.000 |
| 294.619 | 372.423 | 441.373 | -1.257 | -0.000 | 0.000 |
| 197.433 | 197.433 | 716.524 | 0.000 | -0.000 | -0.000 |
| 15.572 | -15.572 | 0.000 | 104.264 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.264 | 15.572 |
| 0.000 | 0.000 | 0.000 | 0.000 | 15.572 | 181.854 |
| 464.189 | 404.377 | 240.248 | 0.356 | 0.000 | 0.000 |
| 273.829 | 637.536 | 197.535 | -15.572 | 0.000 | 0.000 |
| 386.011 | 386.011 | 340.115 | 0.000 | 0.000 | -0.000 |
| 15.572 | -15.572 | -0.000 | 104.264 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.264 | 15.572 |
| -0.000 | -0.000 | -0.000 | -0.000 | 15.572 | 181.854 |
| 637.539 | 273.829 | 197.535 | 15.572 | -0.000 | -0.000 |
| 273.829 | 637.539 | 197.535 | -15.572 | -0.000 | -0.000 |
| 323.329 | 323.329 | 465.235 | -0.000 | -0.000 | -0.000 |
| -2.361 | -7.062 | 9.405 | 38.898 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.264 | 15.572 |
| 0.000 | 0.000 | 0.000 | 0.000 | 15.572 | 181.854 |
| 637.534 | 273.828 | 197.534 | 15.572 | -0.000 | -0.000 |
| 347.049 | 444.617 | 316.993 | -2.655 | -0.000 | -0.000 |
| 332.706 | 332.706 | 446.510 | 0.000 | 0.000 | 0.000 |
| 7.147 | 2.240 | -9.369 | 39.407 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.264 | 15.572 |
| -0.000 | -0.000 | -0.000 | -0.000 | 15.572 | 181.854 |
| 324.247 | 162.846 | 162.846 | -0.000 | 0.000 | 0.000 |
| 162.846 | 324.247 | 162.846 | -0.000 | 0.000 | -0.000 |
| 162.846 | 162.846 | 324.247 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 55.754 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 55.754 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 55.754 |
| 289.299 | 221.873 | 221.873 | -0.000 | 0.000 | -0.000 |
| 221.873 | 289.299 | 221.873 | -0.000 | -0.000 | -0.000 |
| 221.873 | 221.873 | 289.299 | 0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 55.754 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 55.754 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 55.754 |
| 324.226 | 163.931 | 163.931 | 0.000 | -0.000 | -0.000 |
| 163.931 | 324.226 | 163.931 | -0.000 | -0.000 | -0.000 |
| 163.931 | 163.931 | 324.226 | 0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 55.754 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 55.754 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 55.754 |
| 324.221 | 164.174 | 164.174 | -0.000 | 0.000 | 0.000 |
| 164.174 | 324.221 | 164.174 | 0.000 | 0.000 | 0.000 |
| 164.174 | 164.174 | 324.221 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 55.754 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 55.754 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 55.754 |
| 288.357 | 220.941 | 220.941 | 0.000 | -0.000 | 0.000 |
| 220.941 | 288.357 | 220.941 | 0.000 | 0.000 | 0.000 |
| 220.941 | 220.941 | 288.357 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 55.754 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 55.754 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 55.754 |
| 288.403 | 220.962 | 220.962 | -0.000 | -0.000 | -0.000 |
| 220.962 | 288.403 | 220.962 | -0.000 | 0.000 | 0.000 |
| 220.962 | 220.962 | 288.403 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 55.754 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 55.754 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 55.754 |
| 314.270 | 152.941 | 152.941 | -0.000 | -0.000 | 0.000 |
| 152.941 | 314.270 | 152.941 | -0.000 | 0.000 | 0.000 |
| 152.941 | 152.941 | 314.270 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 35.208 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 35.208 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 35.208 |
| 279.261 | 208.790 | 208.790 | 0.000 | -0.000 | 0.000 |
| 208.790 | 279.261 | 208.790 | 0.000 | -0.000 | -0.000 |
| 208.790 | 208.790 | 279.261 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 35.208 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 35.208 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 35.208 |
| 494.684 | 179.188 | 179.188 | 0.000 | 0.000 | 0.000 |
| 179.188 | 494.684 | 179.188 | 0.000 | 0.000 | 0.000 |
| 179.188 | 179.188 | 494.684 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 39.906 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 39.906 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 39.906 |
| 314.272 | 152.941 | 152.941 | 0.000 | -0.000 | -0.000 |
| 152.941 | 314.272 | 152.941 | 0.000 | -0.000 | -0.000 |
| 152.941 | 152.941 | 314.272 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 39.906 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 39.906 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 39.906 |
| 279.247 | 208.790 | 208.790 | -0.000 | 0.000 | 0.000 |
| 208.790 | 279.247 | 208.790 | -0.000 | 0.000 | 0.000 |
| 208.790 | 208.790 | 279.247 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 55.405 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 55.405 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 55.405 |
| 279.274 | 208.792 | 208.792 | 0.000 | -0.000 | -0.000 |
| 208.792 | 279.274 | 208.792 | 0.000 | -0.000 | -0.000 |
| 208.792 | 208.792 | 279.274 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 55.405 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 55.405 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 55.405 |
| 202.099 | -50.618 | 182.561 | -0.000 | 0.000 | 0.000 |
| 41.026 | -61.185 | 174.189 | 0.000 | -0.000 | 0.000 |
| 10.380 | -89.700 | 245.961 | -0.000 | -0.000 | -0.000 |
| -9.328 | -109.169 | 186.487 | 0.764 | 0.000 | -0.000 |
| -10.004 | -110.757 | 184.215 | 0.000 | -2.494 | -0.000 |
| -10.971 | -111.105 | 182.789 | -0.000 | 0.000 | -14.385 |
| 336.659 | 120.495 | 90.869 | -0.000 | -0.000 | -0.000 |
| 119.455 | 166.093 | 38.576 | -0.000 | 0.000 | -0.000 |
| 100.497 | 220.240 | -33.892 | 0.000 | 0.000 | -0.000 |
| 9.328 | 109.169 | -186.487 | 0.764 | -0.000 | -0.000 |
| 10.004 | 110.757 | -184.215 | -0.000 | -2.494 | 0.000 |
| 10.971 | 111.105 | -182.789 | -0.000 | -0.000 | -14.385 |
| 240.431 | 91.695 | 74.706 | 0.000 | -0.000 | 0.000 |
| 59.672 | 48.345 | 44.244 | 0.000 | 0.000 | -0.000 |
| 91.021 | 39.768 | 221.916 | 0.000 | 0.000 | 0.000 |
| -4.626 | -17.837 | 21.618 | 1.875 | -0.000 | -0.000 |
| 0.474 | -9.932 | 20.643 | -0.000 | -3.150 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 79.157 |
| 336.659 | 120.495 | 90.869 | -0.000 | -0.000 | -0.000 |
| 57.439 | 65.859 | 6.870 | 0.000 | -0.000 | 0.000 |
| 1.350 | 12.169 | -18.396 | 0.000 | -0.000 | 0.000 |
| 4.626 | 17.837 | -21.618 | 1.875 | 0.000 | 0.000 |
| -0.474 | 9.932 | -20.643 | -0.000 | -3.150 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 79.157 |
| 244.589 | 101.034 | 67.197 | -0.000 | 0.000 | 0.000 |
| 58.582 | 51.606 | 1.839 | 0.000 | -0.000 | -0.000 |
| 0.042 | 0.129 | 1.881 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 18.802 | 0.000 | 0.000 |
| -152237.177 | -1192072.783 | 1.855 | 0.000 | 0.012 | 0.000 |
| -0.412 | -1.346 | 1.170 | -0.000 | -0.000 | 11.288 |
| 214.889 | 60.837 | -1.842 | 0.000 | -0.000 | -0.000 |
| 58.819 | 53.797 | -1.839 | -0.000 | -0.000 | 0.000 |
| 0.274 | 2.292 | -1.833 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 18.802 | 0.000 | -0.000 |
| 152237.175 | 1192072.802 | -1.870 | -0.000 | 0.007 | -0.000 |
| 0.412 | 1.346 | -1.170 | 0.000 | 0.000 | 11.288 |
| 638.236 | 275.084 | 197.121 | 0.000 | 0.000 | 0.000 |
| 340.463 | 437.469 | 332.757 | 0.000 | -0.000 | -0.000 |
| 197.215 | 197.215 | 713.711 | 0.000 | 0.000 | -0.000 |
| -2.176 | -2.176 | 4.360 | 39.526 | -0.000 | -0.000 |
| -2.176 | -2.176 | 4.360 | -0.000 | 39.526 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 181.576 |
| 441.776 | 344.880 | 324.018 | 0.000 | -0.000 | 0.000 |
| 275.084 | 638.236 | 197.121 | -0.000 | -0.000 | 0.000 |
| 329.442 | 329.442 | 448.771 | -0.000 | 0.000 | 0.000 |
| 2.176 | 2.176 | -4.360 | 39.526 | 0.000 | 0.000 |
| 2.176 | 2.176 | -4.360 | -0.000 | 39.526 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 181.576 |
| 638.236 | 275.084 | 197.121 | 0.000 | 0.000 | 0.000 |
| 342.450 | 439.409 | 328.824 | 0.000 | 0.000 | -0.000 |
| 327.050 | 327.050 | 453.563 | 0.000 | -0.000 | -0.000 |
| -0.218 | -0.218 | 0.436 | 39.529 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.813 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 181.576 |
| 638.236 | 275.084 | 197.121 | -0.000 | -0.000 | -0.000 |
| 275.084 | 638.235 | 197.121 | -0.000 | -0.000 | 0.000 |
| 197.215 | 197.215 | 713.711 | -0.000 | -0.000 | 0.000 |
| 0.218 | 0.218 | -0.436 | 39.529 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.813 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 181.576 |
| 439.142 | 351.614 | 319.913 | 0.000 | -0.000 | 0.000 |
| 342.588 | 439.784 | 328.313 | 0.000 | -0.000 | -0.000 |
| 327.255 | 327.255 | 453.156 | 0.000 | -0.000 | -0.000 |
| -0.021 | -0.022 | 0.044 | 39.529 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.813 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 181.576 |
| 439.184 | 351.657 | 319.821 | 0.000 | -0.000 | -0.000 |
| 351.654 | 439.193 | 319.815 | 0.000 | 0.000 | -0.000 |
| 327.472 | 327.472 | 452.714 | -0.000 | 0.000 | 0.000 |
| 0.021 | 0.022 | -0.044 | 39.529 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.813 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 181.576 |
| 638.117 | 274.790 | 197.239 | 0.000 | -0.000 | -0.000 |
| -1188.345 | -1266.495 | 3567.946 | -0.000 | -0.000 | 0.000 |
| 197.284 | 197.284 | 714.474 | -0.000 | 0.000 | -0.000 |
| -1613.365 | -1613.365 | 3229.566 | -7.534 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 104.688 | -0.000 |
| -1612.065 | -1612.065 | 3226.965 | -0.000 | -0.000 | -44.326 |
| 1943.267 | 2042.896 | -2878.720 | 0.000 | 0.000 | 0.000 |
| 274.790 | 638.117 | 197.239 | -0.000 | -0.000 | 0.000 |
| 1952.339 | 1952.339 | -2798.723 | -0.000 | 0.000 | -0.000 |
| 1613.365 | 1613.365 | -3229.566 | -7.534 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 104.688 | -0.000 |
| 1612.065 | 1612.065 | -3226.965 | 0.000 | 0.000 | -44.326 |
| 638.117 | 274.790 | 197.239 | 0.000 | 0.000 | -0.000 |
| 238.227 | 250.578 | 621.713 | 0.000 | -0.000 | 0.000 |
| 197.284 | 197.284 | 714.474 | -0.000 | 0.000 | -0.000 |
| -150.269 | -150.269 | 300.802 | 0.164 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.688 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 181.663 |
| 462.531 | 548.455 | 99.073 | 0.000 | 0.000 | 0.000 |
| 548.454 | 462.532 | 99.073 | 0.000 | 0.000 | -0.000 |
| 476.352 | 476.352 | 155.847 | 0.000 | 0.000 | -0.000 |
| 150.269 | 150.269 | -300.802 | 0.164 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.688 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 181.663 |
| 638.119 | 274.791 | 197.239 | -0.000 | -0.000 | -0.000 |
| 335.018 | 431.034 | 344.226 | 0.000 | -0.000 | -0.000 |
| 319.377 | 319.377 | 470.076 | -0.000 | -0.000 | 0.000 |
| -9.701 | -9.701 | 19.420 | 37.208 | 0.000 | 0.000 |
| -9.701 | -9.701 | 19.420 | -0.000 | 37.208 | -0.000 |
| -9.621 | -9.621 | 19.259 | 0.000 | -0.000 | 46.794 |
| 447.360 | 352.008 | 310.874 | 0.000 | 0.000 | 0.000 |
| 352.004 | 447.370 | 310.868 | 0.000 | -0.000 | 0.000 |
| 338.157 | 338.157 | 432.477 | 0.000 | -0.000 | 0.000 |
| 9.701 | 9.701 | -19.420 | 37.208 | -0.000 | -0.000 |
| 9.701 | 9.701 | -19.420 | 0.000 | 37.208 | 0.000 |
| 9.621 | 9.621 | -19.259 | -0.000 | 0.000 | 46.794 |
| 638.014 | 274.517 | 197.350 | 0.000 | 0.000 | 0.000 |
| 5337.695 | 5422.744 | -9650.463 | 0.000 | -0.000 | -0.000 |
| 5320.590 | 5320.590 | -9531.184 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 104.573 | 0.000 | 0.000 |
| 4989.811 | 4989.811 | -9979.526 | 0.000 | 35.463 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 181.748 |
| 638.014 | 274.517 | 197.351 | -0.000 | -0.000 | -0.000 |
| -4643.844 | -4558.530 | 10312.157 | -0.000 | 0.000 | 0.000 |
| -4662.532 | -4662.532 | 10434.870 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 104.573 | -0.000 | -0.000 |
| -4989.811 | -4989.811 | 9979.526 | -0.000 | 35.463 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 181.748 |
| 638.014 | 274.517 | 197.351 | 0.000 | 0.000 | -0.000 |
| 847.070 | 930.566 | -667.747 | -0.000 | -0.000 | -0.000 |
| 827.191 | 827.191 | -544.470 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 104.573 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.573 | 0.000 |
| 498.765 | 498.765 | -997.521 | 0.000 | -0.000 | 42.677 |
| 638.013 | 274.517 | 197.350 | -0.000 | -0.000 | -0.000 |
| -127.797 | -27.971 | 1265.639 | -0.000 | -0.000 | 0.000 |
| 197.350 | 197.350 | 715.198 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 104.573 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.573 | 0.000 |
| -498.765 | -498.765 | 997.521 | -0.000 | 0.000 | 42.677 |
| 638.016 | 274.518 | 197.351 | 0.000 | -0.000 | 0.000 |
| 391.181 | 483.969 | 234.735 | -0.000 | -0.000 | -0.000 |
| 376.398 | 376.398 | 357.109 | 0.000 | -0.000 | -0.000 |
| 48.642 | 48.642 | -97.283 | 37.203 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.573 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 181.748 |
| 638.011 | 274.516 | 197.350 | -0.000 | -0.000 | -0.000 |
| 274.517 | 638.011 | 197.350 | 0.000 | -0.000 | -0.000 |
| 197.349 | 197.349 | 715.195 | -0.000 | -0.000 | -0.000 |
| -48.642 | -48.642 | 97.283 | 37.203 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.573 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 181.748 |
| 371.389 | 152.746 | 144.826 | -0.000 | 0.000 | -0.000 |
| 153.711 | 214.380 | 29.732 | -0.000 | 0.000 | -0.000 |
| -521.648 | 21.776 | -1603.231 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 12.815 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 65.491 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 40.251 |
| 371.389 | 152.746 | 144.826 | -0.000 | -0.000 | 0.000 |
| 153.711 | 214.380 | 29.732 | -0.000 | -0.000 | 0.000 |
| -141.628 | -30.133 | 1078.523 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 12.815 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 65.491 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 40.251 |
| 371.389 | 152.746 | 144.826 | 0.000 | 0.000 | -0.000 |
| 153.711 | 214.380 | 29.732 | -0.000 | 0.000 | -0.000 |
| 88.102 | 86.505 | -77.044 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 12.815 | 0.000 | -0.000 |
| 14.549 | 5.931 | -62.217 | 0.000 | -26.892 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 40.251 |
| 371.389 | 152.746 | 144.826 | 0.000 | -0.000 | 0.000 |
| 153.711 | 214.380 | 29.732 | 0.000 | -0.000 | 0.000 |
| 144.937 | 28.877 | 288.577 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 12.815 | -0.000 | 0.000 |
| -14.549 | -5.931 | 62.217 | -0.000 | -26.892 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 40.251 |
| 117.341 | 112.559 | -12.540 | 0.000 | 0.000 | -0.000 |
| 116.597 | 21.714 | -16.254 | 0.000 | 0.000 | -0.000 |
| -12.716 | 2.048 | -15.901 | -0.000 | 0.000 | -0.000 |
| -16702612.349 | -11695140.272 | -25431142.058 | 5156239.392 | 129986.080 | -77944.216 |
| -26909088.856 | -32894219.471 | -44368648.775 | -46796.006 | 2856903.293 | -4402.340 |
| -12.762 | -11.541 | -40.888 | 0.000 | -0.000 | -179.582 |
| 128.594 | 113.459 | 23.539 | 0.000 | 0.000 | -0.000 |
| 126.739 | 25.616 | 16.266 | 0.000 | 0.000 | 0.000 |
| 5.170 | 1.099 | 16.260 | 0.000 | 0.000 | 0.000 |
| 16690088.762 | 11969094.019 | 25327404.502 | 4953983.994 | 54285.435 | -20078.739 |
| 27206903.273 | 32534967.968 | 44330701.156 | -30131.238 | 2412071.542 | -1666.690 |
| 12.762 | 11.541 | 40.888 | -0.000 | -0.000 | -179.582 |
| -66991.831 | -13593.031 | 3549.117 | -8610.195 | 2505.036 | 30700.156 |
| -69897.789 | -14361.650 | 2887.583 | -7470.104 | 855.025 | 30024.876 |
| -71198.548 | -15587.246 | 2793.079 | -6833.493 | -2363.069 | 29684.242 |
| -71214.018 | -15586.424 | 2756.374 | -6811.792 | -2347.546 | 29675.995 |
| -69835.132 | -14442.059 | 2884.599 | -7504.504 | 887.415 | 30050.396 |
| -69830.234 | -14370.019 | 2854.028 | -7493.336 | 890.671 | 30046.641 |
| 69975.886 | 14424.143 | -2845.104 | 7462.500 | -833.720 | -30023.868 |
| 71220.614 | 15580.785 | -2731.132 | 6853.609 | 2364.608 | -29622.140 |
| 705348.065 | 574573.796 | 324378.873 | 214632.747 | -75303.133 | -26205.047 |
| 69876.711 | 14357.388 | -2904.157 | 7456.491 | -838.615 | -30101.402 |
| 71206.254 | 15587.867 | -2762.130 | 6836.296 | 2372.461 | -29663.734 |
| 71238.743 | 15628.324 | -2764.287 | 6905.055 | 2301.768 | -29667.245 |
| -6784.372 | -1452.626 | 365.504 | -796.406 | -67.726 | 3033.311 |
| -72683.657 | -56894.354 | -38059.088 | -17531.234 | 4439.996 | 2023.480 |
| -6972.308 | -1424.228 | 319.844 | -747.449 | 84.471 | 3004.679 |
| -72435.171 | -56860.620 | -38010.794 | -17628.612 | 4605.739 | 2092.734 |
| -6976.683 | -1433.490 | 290.011 | -746.105 | 95.526 | 3006.069 |
| -6984.406 | -1433.267 | 285.683 | -748.255 | 85.393 | 3035.493 |
| 76953.277 | 61233.729 | 36122.014 | 20633.754 | -6473.890 | -281.224 |
| 72694.121 | 56968.481 | 38528.757 | 17899.027 | -4663.377 | -1762.658 |
| 7151.116 | 1592.198 | -245.315 | 683.887 | 233.461 | -2953.190 |
| 7125.034 | 1562.747 | -276.326 | 697.321 | 235.201 | -2961.337 |
| 6994.453 | 1441.650 | -287.394 | 747.171 | -70.470 | -3009.660 |
| 6985.097 | 1431.857 | -286.063 | 744.874 | -84.069 | -2987.185 |
| -615.950 | -105.472 | 55.010 | -86.172 | 25.125 | 307.556 |
| -7744.238 | -6051.700 | -3688.958 | -2123.504 | 671.258 | 109.292 |
| -7255.921 | -5681.267 | -3815.929 | -1794.019 | 496.669 | 181.269 |
| -7579.271 | -6088.467 | -3611.583 | -2020.002 | 650.905 | -69.263 |
| -739.127 | -88.973 | -2.099 | 64.689 | 34.386 | 276.468 |
| -7242.793 | -5686.189 | -3800.697 | -1763.946 | 460.185 | 230.794 |
| 7491.354 | 5776.993 | 3927.963 | 1894.184 | -534.976 | -314.932 |
| 727.443 | 191.880 | -7.873 | 74.968 | -8.387 | -300.238 |
| 7255.522 | 5702.587 | 3876.771 | 1801.395 | -482.937 | -182.359 |
| 670.920 | 136.192 | -34.937 | 100.989 | -25.259 | -306.848 |
| 7254.126 | 5679.681 | 3849.640 | 1796.658 | -487.218 | -180.606 |
| 818.493 | 148.328 | -16.327 | 21.013 | 13.524 | -301.479 |
| 528.389 | 212.529 | 0.000 | 0.000 | -0.000 | 0.000 |
| 212.535 | 335.641 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| 528.389 | 212.529 | 0.000 | -0.000 | -0.000 | -0.000 |
| 212.536 | 335.642 | 0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 528.390 | 212.529 | 0.000 | 0.000 | -0.000 | 0.000 |
| 212.535 | 335.639 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 528.389 | 212.529 | 0.000 | -0.000 | -0.000 | -0.000 |
| 212.536 | 335.645 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| 528.391 | 212.530 | 0.000 | 0.000 | -0.000 | 0.000 |
| 212.534 | 335.612 | 0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 528.387 | 212.528 | 0.000 | 0.000 | -0.000 | 0.000 |
| 212.493 | 334.992 | 0.000 | 0.000 | -0.001 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| 328.142 | 230.748 | 0.000 | -0.000 | -0.000 | 0.000 |
| 230.748 | 633.361 | 0.000 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 0.000 |
| 328.142 | 230.748 | 0.000 | 0.000 | 0.000 | -0.000 |
| 230.748 | 633.361 | 0.000 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| 328.142 | 230.748 | 0.000 | -0.000 | -0.000 | 0.000 |
| 230.748 | 633.361 | 0.000 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| 328.143 | 230.748 | 0.000 | 0.000 | 0.000 | -0.000 |
| 230.748 | 633.361 | 0.000 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| 328.138 | 230.748 | 0.000 | -0.000 | -0.000 | 0.000 |
| 230.749 | 633.364 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.001 | 0.000 | -0.000 | 0.000 | 0.000 |
| 328.146 | 230.748 | 0.000 | 0.000 | 0.000 | -0.000 |
| 230.747 | 633.359 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.001 | -0.000 | -0.000 | -0.000 | 0.000 |
| 153.195 | 48.466 | 48.465 | 0.000 | -0.000 | 0.000 |
| 48.464 | 153.196 | 48.465 | 0.000 | -0.000 | 0.000 |
| 48.464 | 48.466 | 153.196 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 34.519 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 34.519 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 34.519 |
| 152.647 | 47.573 | 47.574 | -0.000 | 0.000 | -0.000 |
| 47.574 | 152.646 | 47.574 | 0.000 | 0.000 | -0.000 |
| 47.574 | 47.573 | 152.646 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 34.519 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 34.519 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 34.519 |
| 133.582 | 44.641 | 44.641 | -0.000 | -0.000 | 0.000 |
| 44.641 | 133.592 | 44.641 | -0.000 | -0.000 | 0.000 |
| 44.641 | 44.641 | 133.582 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 34.519 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 34.519 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 34.519 |
| 152.911 | 47.974 | 47.974 | -0.000 | 0.000 | -0.000 |
| 47.974 | 152.911 | 47.974 | -0.000 | 0.000 | -0.000 |
| 47.974 | 47.974 | 152.911 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 34.519 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 34.519 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 34.519 |
| 152.936 | 48.023 | 48.023 | 0.000 | -0.000 | 0.000 |
| 48.023 | 152.936 | 48.023 | -0.000 | -0.000 | 0.000 |
| 48.023 | 48.023 | 152.936 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 34.519 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 34.519 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 34.519 |
| 152.941 | 48.014 | 48.014 | 0.000 | 0.000 | -0.000 |
| 48.014 | 152.941 | 48.014 | 0.000 | 0.000 | -0.000 |
| 48.014 | 48.014 | 152.941 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 34.519 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 34.519 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 34.519 |
| 26052264.734 | 26025090.989 | 23134954.367 | 297488.076 | 497235.607 | 279442.804 |
| 26052204.044 | 26025180.378 | 23134981.245 | 297386.027 | 497368.973 | 279465.290 |
| 26052241.965 | 26025141.638 | 23135124.719 | 297446.587 | 497270.147 | 279449.013 |
| 26052193.299 | 26025099.089 | 23134943.280 | 297540.205 | 497222.104 | 279442.407 |
| 26052195.562 | 26025090.382 | 23134957.404 | 297433.264 | 497378.811 | 279449.354 |
| 26052221.414 | 26025100.166 | 23134961.662 | 297459.675 | 497379.659 | 279496.319 |
| -26052149.359 | -26025118.705 | -23134944.895 | -297395.670 | -497382.286 | -279465.036 |
| -26052211.342 | -26025032.506 | -23134918.809 | -297493.430 | -497254.409 | -279443.335 |
| -26052175.560 | -26025072.758 | -23134774.722 | -297429.897 | -497366.528 | -279461.835 |
| -26052222.413 | -26025115.027 | -23134959.796 | -297337.550 | -497409.430 | -279467.392 |
| -26052219.844 | -26025118.055 | -23134941.055 | -297458.277 | -497236.252 | -279457.797 |
| -26052195.077 | -26025111.467 | -23134936.529 | -297423.401 | -497251.329 | -279413.579 |
| 2605278.484 | 2602495.835 | 2313495.382 | 29795.055 | 49644.048 | 27932.412 |
| 2605545.818 | 2603278.175 | 2313302.688 | 28699.760 | 49721.921 | 28067.203 |
| 2605407.503 | 2602957.000 | 2313714.137 | 28969.492 | 50231.145 | 28100.691 |
| 2605217.808 | 2602596.333 | 2313547.473 | 29562.872 | 50039.764 | 28004.997 |
| 2605208.388 | 2602509.209 | 2313506.177 | 29697.739 | 49839.683 | 27948.015 |
| 2605236.395 | 2602514.212 | 2313509.102 | 29743.781 | 49832.786 | 27992.923 |
| -2605317.097 | -2602955.203 | -2313490.656 | -28915.072 | -50244.265 | -28103.689 |
| -2605444.908 | -2602951.920 | -2313441.178 | -28889.598 | -50125.186 | -28099.783 |
| -2605258.419 | -2602698.405 | -2313387.440 | -29247.457 | -50345.248 | -28071.642 |
| -2605275.284 | -2602644.375 | -2313551.521 | -29374.699 | -50197.626 | -28027.848 |
| -2605458.847 | -2603044.872 | -2313422.236 | -28877.653 | -49976.104 | -28094.331 |
| -2605305.147 | -2602809.013 | -2313587.911 | -29100.429 | -50321.603 | -28060.166 |
| 260601.867 | 260274.231 | 231371.498 | 2941.069 | 4996.904 | 2806.545 |
| 260522.946 | 260342.581 | 231369.425 | 2904.126 | 5007.816 | 2804.921 |
| 260629.136 | 260399.027 | 231475.999 | 2868.936 | 4802.937 | 2790.214 |
| 260577.724 | 260325.448 | 231321.457 | 2999.043 | 4876.384 | 2798.360 |
| 260550.693 | 260317.242 | 231337.514 | 2869.633 | 5034.163 | 2802.890 |
| 260534.299 | 260237.183 | 231332.901 | 3039.232 | 4920.288 | 2818.538 |
| -260477.281 | -260318.236 | -231325.236 | -2854.057 | -5028.199 | -2811.018 |
| -260534.723 | -260169.365 | -231292.092 | -3083.332 | -4759.793 | -2766.774 |
| -260536.145 | -260308.295 | -231115.027 | -2857.053 | -4931.887 | -2803.533 |
| -260541.267 | -260172.584 | -231294.219 | -3239.135 | -4410.615 | -2715.510 |
| -260619.607 | -260399.561 | -231241.231 | -2846.898 | -4686.650 | -2788.158 |
| -260509.169 | -260250.938 | -231335.435 | -2970.982 | -4843.309 | -2743.256 |
| 233.612 | 90.271 | 114.190 | -0.002 | -0.052 | 0.001 |
| 35.069 | 84.539 | 62.625 | 3.399 | 5.099 | -0.973 |
| 43.592 | 44.449 | 54.346 | 8.060 | 1.001 | -2.215 |
| 13.140 | -0.308 | 15.568 | 5.269 | 3.485 | -3.689 |
| 6.828 | -1.491 | 17.173 | 5.805 | 5.973 | -0.833 |
| 5.657 | 3.810 | 17.370 | 2.597 | 2.816 | -6.599 |
| 7.561 | -26.844 | -49.320 | -9.156 | -0.965 | 2.505 |
| 87.801 | 176.000 | 84.503 | -0.004 | -0.022 | 0.008 |
| -72.819 | -106.365 | -96.367 | 0.281 | -3.617 | -2.332 |
| 0.499 | -9.224 | -20.068 | 2.039 | -1.564 | 2.116 |
| -1.491 | 1.712 | -2.440 | -2.905 | 17.755 | 1.544 |
| -5.410 | 4.269 | -7.076 | -8.079 | -1.012 | -2.951 |
| -10.704 | -65.492 | -54.770 | 0.018 | 0.311 | 0.126 |
| 15.191 | 44.827 | 25.423 | 0.334 | 0.589 | -0.071 |
| 5.946 | 27.002 | 25.006 | -0.380 | 0.089 | -0.362 |
| -0.002 | -0.004 | 0.022 | 54.398 | -0.003 | -0.009 |
| 0.071 | 0.768 | 1.237 | -0.042 | 8.598 | 0.192 |
| 0.022 | 0.684 | 1.186 | 0.009 | 0.320 | 10.377 |
| 233.612 | 90.271 | 114.190 | -0.002 | -0.052 | 0.001 |
| -22.291 | 1.899 | -24.787 | -0.055 | -0.420 | -0.372 |
| -20.686 | -20.197 | -10.678 | -0.839 | -0.117 | 0.221 |
| -0.520 | 0.398 | -0.691 | -10.977 | -0.061 | 0.212 |
| -0.050 | -0.021 | 0.027 | -0.003 | 70.307 | 0.003 |
| -0.519 | 0.426 | -0.656 | -0.778 | -0.121 | -2.234 |
| 36.842 | 2.379 | 13.830 | 0.009 | 0.032 | -0.023 |
| -8.129 | 5.213 | -8.785 | 0.047 | -0.005 | 0.003 |
| 5.715 | -0.033 | 14.845 | 0.014 | 0.047 | -0.023 |
| -0.003 | 0.052 | 0.134 | 0.527 | 0.003 | -0.017 |
| 0.164 | 0.129 | 0.347 | -0.163 | 11.076 | -0.059 |
| -0.014 | -0.013 | 0.054 | 0.034 | 0.012 | 7.864 |
| 42.173 | -6.037 | 2.369 | -0.025 | -0.051 | -0.017 |
| -0.244 | 3.283 | -5.748 | -0.040 | -0.020 | 0.013 |
| 2.759 | 1.706 | 20.440 | -0.053 | -0.015 | 0.024 |
| -0.039 | 0.024 | -0.049 | 9.403 | 0.013 | 0.016 |
| -0.002 | 0.001 | -0.060 | -0.055 | 11.141 | 0.005 |
| -0.082 | -0.092 | -0.217 | -0.007 | -0.030 | 7.904 |
| 172.649 | 130.229 | 130.472 | 0.484 | 0.474 | 1.062 |
| 30.978 | 47.516 | 28.889 | -1.214 | 0.244 | -0.053 |
| 41.292 | 38.039 | 36.667 | -1.282 | 0.313 | -0.030 |
| 37.881 | 36.370 | 36.551 | -2.294 | 0.263 | -0.045 |
| 22.074 | 19.114 | 19.467 | -1.049 | 71.953 | -0.006 |
| 45.318 | 42.674 | 42.990 | -1.225 | 0.251 | 13.437 |
| -29.562 | -44.616 | -44.900 | 1.225 | -0.248 | 0.048 |
| -41.464 | -35.884 | -38.621 | 1.276 | -0.277 | 0.080 |
| -42.371 | -39.818 | -36.475 | 1.238 | -0.257 | 0.056 |
| -0.000 | 0.000 | 0.000 | 543.364 | 0.000 | 0.000 |
| -45.315 | -42.673 | -42.988 | 1.224 | 13.267 | 0.035 |
| -3.190 | -2.578 | -2.651 | 0.824 | 0.042 | 204.325 |
| 6.947 | 2.098 | 2.129 | -0.126 | 0.027 | -0.003 |
| 3.096 | 7.355 | 2.879 | -0.038 | 0.062 | 0.088 |
| 9.819 | 9.579 | 30.162 | -0.397 | 0.082 | 0.157 |
| 3.962 | 3.566 | 3.613 | 0.218 | 0.027 | -0.006 |
| 4.085 | 3.870 | 3.896 | -0.122 | 0.944 | -0.005 |
| 4.628 | 4.242 | 4.289 | -0.123 | 0.025 | 19.472 |
| -1.520 | -3.564 | -3.594 | 0.150 | -0.020 | 0.025 |
| -4.951 | -2.492 | -4.725 | 0.134 | -0.027 | -0.001 |
| -12.474 | -12.234 | 3.498 | 0.114 | -0.023 | 0.008 |
| -0.865 | -0.720 | -0.738 | 144.269 | -0.009 | -0.012 |
| -4.052 | -3.744 | -3.786 | 0.126 | -0.035 | 0.005 |
| -0.865 | -0.720 | -0.738 | 0.096 | -0.009 | 144.195 |
| 7.964 | -0.745 | -0.742 | -0.013 | 0.003 | 0.000 |
| -0.757 | 2.155 | -0.780 | -0.013 | 0.003 | -0.000 |
| -0.116 | -0.141 | 2.491 | 0.005 | 0.013 | 0.025 |
| 0.634 | 0.598 | 0.602 | 0.042 | 0.003 | -0.000 |
| 0.409 | 0.377 | 0.381 | -0.012 | 1.169 | -0.000 |
| 0.523 | 0.325 | 0.327 | -0.012 | 0.003 | 0.939 |
| 3.293 | 0.294 | 0.291 | 0.015 | -0.004 | -0.004 |
| -2.495 | 1.145 | -2.471 | 0.004 | -0.013 | -0.023 |
| -0.423 | -0.390 | -0.359 | 0.013 | -0.003 | 0.001 |
| -0.395 | -0.359 | -0.364 | 0.009 | -0.003 | 0.001 |
| -0.374 | -0.398 | -0.371 | 0.012 | 8.529 | 0.000 |
| -0.358 | -0.372 | -0.377 | 0.012 | -0.003 | 1.207 |
| 151.847 | 76.103 | 76.103 | 0.000 | -0.000 | -0.000 |
| 76.103 | 151.847 | 76.103 | 0.000 | -0.000 | -0.000 |
| 76.103 | 76.103 | 151.847 | 0.000 | -0.000 | -0.000 |
| 5.037 | 5.037 | 5.037 | 4.284 | -0.000 | -0.000 |
| 5.037 | 5.037 | 5.037 | -0.000 | 4.284 | -0.000 |
| 5.037 | 5.037 | 5.037 | 0.000 | -0.000 | 4.284 |
| 151.847 | 76.103 | 76.103 | 0.000 | -0.000 | 0.000 |
| 76.103 | 151.847 | 76.103 | -0.000 | 0.000 | 0.000 |
| 76.103 | 76.103 | 151.847 | -0.000 | 0.000 | 0.000 |
| -5.037 | -5.037 | -5.037 | 4.284 | 0.000 | 0.000 |
| -5.037 | -5.037 | -5.037 | -0.000 | 4.284 | 0.000 |
| -5.037 | -5.037 | -5.037 | 0.000 | 0.000 | 4.284 |
| 13.034 | -0.805 | -0.805 | 0.000 | -0.000 | -0.000 |
| -0.805 | 13.034 | -0.805 | 0.000 | -0.000 | -0.000 |
| -0.805 | -0.805 | 13.034 | 0.000 | -0.000 | -0.000 |
| 0.503 | 0.503 | 0.503 | 4.317 | -0.000 | -0.000 |
| 0.503 | 0.503 | 0.503 | -0.000 | 4.317 | 0.000 |
| 0.503 | 0.503 | 0.503 | -0.000 | -0.000 | 4.317 |
| 40.800 | 3.737 | 3.737 | 0.000 | 0.000 | -0.000 |
| 3.737 | 40.800 | 3.737 | 0.000 | -0.000 | -0.000 |
| 3.737 | 3.737 | 40.800 | 0.000 | -0.000 | -0.000 |
| -0.503 | -0.503 | -0.503 | 4.317 | 0.000 | 0.000 |
| -0.503 | -0.503 | -0.503 | 0.000 | 4.317 | 0.000 |
| -0.503 | -0.503 | -0.503 | -0.000 | 0.000 | 4.317 |
| 9.139 | -13.157 | -13.157 | -0.000 | -0.000 | -0.000 |
| -13.157 | 9.139 | -13.157 | 0.000 | -0.000 | -0.000 |
| -13.157 | -13.157 | 9.139 | -0.000 | -0.000 | 0.000 |
| 0.049 | 0.049 | 0.049 | 4.317 | -0.000 | -0.000 |
| 0.049 | 0.049 | 0.049 | -0.000 | 4.317 | -0.000 |
| 0.049 | 0.049 | 0.049 | 0.000 | -0.000 | 4.317 |
| 2.013 | 0.248 | 0.248 | 0.000 | 0.000 | 0.000 |
| 0.248 | 2.013 | 0.248 | -0.000 | 0.000 | 0.000 |
| 0.248 | 0.248 | 2.013 | -0.000 | -0.000 | -0.000 |
| -0.049 | -0.049 | -0.049 | 4.317 | 0.000 | 0.000 |
| -0.049 | -0.049 | -0.049 | -0.000 | 4.317 | 0.000 |
| -0.049 | -0.049 | -0.049 | -0.000 | 0.000 | 4.317 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 753.132 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 753.132 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 753.132 | -0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 753.131 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 753.135 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 753.129 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 601.401 | 590.683 | 0.000 | 0.000 | 0.000 | 0.000 |
| 590.683 | 601.401 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 314.704 |
| 601.401 | 590.683 | -0.000 | -0.000 | -0.000 | -0.000 |
| 590.683 | 601.401 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 314.704 |
| 601.401 | 590.683 | 0.000 | 0.000 | 0.000 | 0.000 |
| 590.683 | 601.401 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 314.704 |
| 601.401 | 590.683 | -0.000 | -0.000 | -0.000 | -0.000 |
| 590.683 | 601.401 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 314.704 |
| 601.402 | 590.685 | 0.000 | 0.000 | 0.000 | 0.000 |
| 590.685 | 601.402 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.001 | 0.000 | 0.000 | 0.000 | 314.704 |
| 601.400 | 590.681 | -0.000 | -0.000 | -0.000 | -0.000 |
| 590.681 | 601.400 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.001 | -0.000 | -0.000 | -0.000 | 314.704 |
| 324.536 | 164.437 | 164.437 | -0.000 | 0.000 | 0.000 |
| 164.437 | 324.536 | 164.437 | -0.000 | 0.000 | -0.000 |
| 164.437 | 164.437 | 324.536 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 55.765 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 55.765 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 55.765 |
| 324.538 | 164.372 | 164.372 | -0.000 | 0.000 | 0.000 |
| 164.372 | 324.538 | 164.372 | 0.000 | -0.000 | 0.000 |
| 164.372 | 164.372 | 324.538 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 55.765 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 55.765 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 55.765 |
| 324.536 | 164.407 | 164.407 | 0.000 | -0.000 | -0.000 |
| 164.407 | 324.536 | 164.407 | 0.000 | 0.000 | -0.000 |
| 164.407 | 164.407 | 324.536 | -0.000 | 0.000 | -0.000 |
| 0.004 | 0.004 | 0.004 | 38.814 | 0.000 | -0.000 |
| 0.004 | 0.004 | 0.004 | -0.000 | 38.814 | -0.000 |
| 0.004 | 0.004 | 0.004 | 0.000 | 0.000 | 38.814 |
| 324.538 | 164.401 | 164.401 | -0.000 | -0.000 | -0.000 |
| 164.401 | 324.538 | 164.401 | -0.000 | -0.000 | 0.000 |
| 164.401 | 164.401 | 324.538 | -0.000 | 0.000 | -0.000 |
| -0.004 | -0.004 | -0.004 | 38.814 | -0.000 | 0.000 |
| -0.004 | -0.004 | -0.004 | -0.000 | 38.814 | 0.000 |
| -0.004 | -0.004 | -0.004 | -0.000 | 0.000 | 38.814 |
| 288.654 | 221.335 | 221.335 | 0.000 | 0.000 | -0.000 |
| 221.335 | 288.654 | 221.335 | 0.000 | 0.000 | -0.000 |
| 221.335 | 221.335 | 288.654 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 33.319 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 33.319 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 33.319 |
| 288.680 | 221.339 | 221.339 | -0.000 | -0.000 | 0.000 |
| 221.339 | 288.680 | 221.339 | -0.000 | -0.000 | 0.000 |
| 221.339 | 221.339 | 288.680 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 33.319 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 33.319 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 33.319 |
| 638.176 | 274.577 | 197.386 | 0.000 | 0.000 | 0.000 |
| 274.577 | 638.176 | 197.386 | 0.000 | 0.000 | -0.000 |
| 197.386 | 197.386 | 715.366 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 104.610 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 39.447 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 181.800 |
| 435.828 | 344.358 | 329.954 | -0.000 | -0.000 | -0.000 |
| 344.358 | 435.828 | 329.953 | 0.000 | -0.000 | 0.000 |
| 197.386 | 197.386 | 715.366 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 104.610 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 39.446 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 181.800 |
| 638.176 | 274.577 | 197.387 | 0.000 | 0.000 | -0.000 |
| 346.621 | 435.817 | 327.702 | 0.000 | 0.000 | -0.000 |
| 327.714 | 327.714 | 454.711 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 39.447 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 39.447 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 181.800 |
| 435.828 | 344.358 | 329.954 | 0.000 | -0.000 | 0.000 |
| 344.357 | 435.829 | 329.953 | -0.000 | -0.000 | 0.000 |
| 327.712 | 327.712 | 454.715 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 39.447 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 39.447 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 181.800 |
| 435.801 | 346.628 | 327.714 | -0.000 | 0.000 | -0.000 |
| 346.631 | 435.792 | 327.720 | -0.000 | 0.000 | -0.000 |
| 327.723 | 327.723 | 454.696 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 39.447 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 39.447 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 44.586 |
| 638.174 | 274.576 | 197.386 | -0.000 | -0.000 | 0.000 |
| 346.623 | 435.806 | 327.707 | -0.000 | -0.000 | -0.000 |
| 327.703 | 327.703 | 454.730 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 39.447 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 39.447 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 44.586 |
| 789.379 | 251.973 | 234.944 | -0.000 | -0.000 | -0.000 |
| 251.973 | 789.379 | 234.944 | -0.000 | 0.000 | -0.000 |
| -181.919 | -181.919 | 1450.411 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 174.207 | 0.000 | 0.000 |
| -588.557 | -588.557 | 450.421 | -0.000 | 68.793 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 268.703 |
| 789.379 | 251.973 | 234.944 | 0.000 | -0.000 | -0.000 |
| 251.973 | 789.379 | 234.944 | 0.000 | 0.000 | 0.000 |
| 299.529 | 299.530 | 1081.960 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 174.207 | -0.000 | 0.000 |
| 588.557 | 588.557 | -450.421 | 0.000 | 68.793 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 268.703 |
| 789.379 | 251.973 | 234.944 | -0.000 | 0.000 | -0.000 |
| 240.768 | 456.543 | 366.591 | -0.000 | 0.000 | -0.000 |
| 299.530 | 299.530 | 1081.961 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 174.207 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 174.207 | -0.000 |
| -34.583 | -34.583 | 26.466 | -0.000 | 0.000 | 146.410 |
| 789.379 | 251.973 | 234.944 | 0.000 | -0.000 | 0.000 |
| 251.973 | 789.379 | 234.944 | 0.000 | -0.000 | 0.000 |
| 490.990 | 490.990 | 935.435 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 174.207 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 174.207 | -0.000 |
| 34.583 | 34.583 | -26.466 | 0.000 | -0.000 | 146.410 |
| 495.914 | 192.750 | 369.902 | 0.000 | 0.000 | -0.000 |
| 192.758 | 495.927 | 369.894 | 0.000 | 0.000 | -0.000 |
| 299.530 | 299.530 | 1081.966 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 174.207 | 0.000 | -0.000 |
| -3.348 | -3.348 | 2.562 | -0.000 | 114.245 | -0.000 |
| -3.314 | -3.313 | 2.536 | -0.000 | 0.000 | 151.529 |
| 789.376 | 251.972 | 234.944 | 0.000 | -0.000 | -0.000 |
| 287.089 | 533.536 | 319.403 | -0.000 | 0.000 | 0.000 |
| 435.906 | 435.906 | 977.586 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 174.207 | -0.000 | 0.000 |
| 3.348 | 3.348 | -2.562 | 0.000 | 114.245 | 0.000 |
| 3.314 | 3.313 | -2.536 | 0.000 | -0.000 | 151.529 |
| 638.107 | 274.924 | 197.157 | 0.000 | -0.000 | -0.000 |
| 339.846 | 436.868 | 333.665 | -0.000 | -0.000 | -0.000 |
| 324.750 | 324.750 | 458.577 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 104.740 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 104.740 | 0.000 |
| -2.626 | -2.626 | 5.260 | -0.000 | -0.000 | 47.875 |
| 442.001 | 345.107 | 323.256 | 0.000 | -0.000 | 0.000 |
| 345.107 | 442.001 | 323.256 | 0.000 | 0.000 | 0.000 |
| 329.931 | 329.931 | 448.201 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 104.740 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 104.740 | 0.000 |
| 2.626 | 2.626 | -5.260 | 0.000 | 0.000 | 47.875 |
| 439.184 | 342.210 | 328.978 | -0.000 | 0.000 | -0.000 |
| 342.204 | 439.202 | 328.966 | -0.000 | 0.000 | -0.000 |
| 197.229 | 197.229 | 713.976 | 0.000 | -0.000 | -0.000 |
| -0.259 | -0.259 | 0.519 | 39.489 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.740 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 181.591 |
| 638.106 | 274.924 | 197.157 | 0.000 | 0.000 | -0.000 |
| 351.798 | 439.242 | 319.318 | -0.000 | -0.000 | -0.000 |
| 327.595 | 327.595 | 452.878 | 0.000 | 0.000 | 0.000 |
| 0.259 | 0.259 | -0.519 | 39.489 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.740 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 181.591 |
| 638.109 | 274.925 | 197.158 | 0.000 | -0.000 | 0.000 |
| 347.289 | 435.811 | 327.273 | 0.000 | 0.000 | -0.000 |
| 327.321 | 327.321 | 453.431 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 104.740 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.740 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 181.591 |
| 638.104 | 274.924 | 197.157 | 0.000 | 0.000 | 0.000 |
| 351.557 | 439.003 | 319.796 | 0.000 | -0.000 | -0.000 |
| 197.229 | 197.229 | 713.972 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 104.740 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.740 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 181.591 |
| 638.176 | 274.577 | 197.386 | 0.000 | 0.000 | -0.000 |
| 342.037 | 439.264 | 328.839 | -0.000 | -0.000 | -0.000 |
| 327.713 | 327.713 | 454.713 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 39.446 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 104.610 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 48.043 |
| 638.176 | 274.576 | 197.386 | -0.000 | -0.000 | -0.000 |
| 342.037 | 439.264 | 328.839 | 0.000 | 0.000 | 0.000 |
| 197.386 | 197.386 | 715.366 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 39.447 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 104.610 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 48.043 |
| 439.264 | 342.037 | 328.839 | -0.000 | 0.000 | -0.000 |
| 274.577 | 638.177 | 197.387 | -0.000 | 0.000 | 0.000 |
| 327.714 | 327.714 | 454.711 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 104.610 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 39.447 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 181.800 |
| 638.176 | 274.576 | 197.386 | -0.000 | 0.000 | -0.000 |
| 342.036 | 439.264 | 328.838 | 0.000 | 0.000 | 0.000 |
| 327.712 | 327.712 | 454.715 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 104.610 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 39.447 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 181.800 |
| 638.179 | 274.577 | 197.387 | 0.000 | -0.000 | 0.000 |
| 342.041 | 439.256 | 328.846 | -0.000 | -0.000 | -0.000 |
| 327.723 | 327.723 | 454.696 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 39.447 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 39.447 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 181.800 |
| 638.174 | 274.576 | 197.386 | -0.000 | -0.000 | -0.000 |
| 342.097 | 439.083 | 328.957 | 0.000 | 0.000 | 0.000 |
| 327.703 | 327.703 | 454.730 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 39.447 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 39.447 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 181.800 |
| 638.292 | 275.100 | 197.136 | 0.000 | 0.000 | 0.000 |
| 1226.707 | 1333.224 | -1452.407 | 0.000 | -0.000 | -0.000 |
| 197.229 | 197.229 | 713.786 | -0.000 | 0.000 | -0.000 |
| 1042.652 | 1042.652 | -2089.106 | 32.062 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 104.824 | -0.000 |
| 934.713 | 934.713 | -1872.836 | 0.000 | -0.000 | 49.167 |
| 638.291 | 275.100 | 197.136 | -0.000 | -0.000 | -0.000 |
| 275.100 | 638.291 | 197.136 | -0.000 | -0.000 | 0.000 |
| -721.999 | -721.999 | 2555.595 | -0.000 | 0.000 | -0.000 |
| -1042.652 | -1042.652 | 2089.106 | 32.062 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 104.824 | -0.000 |
| -934.713 | -934.713 | 1872.836 | -0.000 | 0.000 | 49.167 |
| 638.292 | 275.100 | 197.136 | -0.000 | 0.000 | 0.000 |
| 453.758 | 523.462 | 133.190 | 0.000 | -0.000 | 0.000 |
| 426.657 | 426.657 | 254.095 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 104.824 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 104.824 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 181.596 |
| 638.291 | 275.100 | 197.136 | -0.000 | -0.000 | 0.000 |
| 275.100 | 638.291 | 197.136 | -0.000 | -0.000 | 0.000 |
| 197.229 | 197.229 | 713.786 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 104.824 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 104.824 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 181.596 |
| 638.294 | 275.101 | 197.136 | 0.000 | 0.000 | -0.000 |
| 351.474 | 449.870 | 309.391 | 0.000 | -0.000 | 0.000 |
| 336.610 | 336.610 | 434.520 | -0.000 | -0.000 | -0.000 |
| 9.385 | 9.385 | -18.804 | 39.329 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.824 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 181.596 |
| 638.289 | 275.099 | 197.135 | 0.000 | -0.000 | 0.000 |
| 335.710 | 431.269 | 343.811 | -0.000 | 0.000 | 0.000 |
| 318.049 | 318.049 | 471.703 | -0.000 | 0.000 | -0.000 |
| -9.385 | -9.385 | 18.804 | 39.329 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 104.824 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 181.596 |
| 638.366 | 275.175 | 197.124 | 0.000 | 0.000 | 0.000 |
| 274.994 | 333.566 | 502.720 | 0.000 | 0.000 | -0.000 |
| 283.317 | 283.317 | 541.168 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 104.860 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 104.860 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 181.595 |
| 638.366 | 275.175 | 197.124 | -0.000 | -0.000 | -0.000 |
| 445.417 | 471.719 | 193.523 | 0.000 | -0.000 | 0.000 |
| 417.425 | 417.425 | 272.410 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 104.860 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 104.860 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 0.000 | 181.595 |
| 433.943 | 335.231 | 341.782 | -0.000 | 0.000 | 0.000 |
| 275.176 | 638.367 | 197.124 | 0.000 | 0.000 | 0.000 |
| 197.227 | 197.227 | 713.693 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 104.860 | -0.000 | 0.000 |
| -6.544 | -6.544 | 13.115 | -0.000 | 40.215 | 0.000 |
| -6.727 | -6.727 | 13.481 | -0.000 | -0.000 | 48.637 |
| 638.366 | 275.175 | 197.124 | -0.000 | 0.000 | 0.000 |
| 350.334 | 448.776 | 311.787 | -0.000 | 0.000 | -0.000 |
| 197.227 | 197.227 | 713.692 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 104.860 | 0.000 | -0.000 |
| 6.544 | 6.544 | -13.115 | 0.000 | 40.215 | 0.000 |
| 6.727 | 6.727 | -13.481 | 0.000 | -0.000 | 48.637 |
| 440.810 | 340.973 | 329.152 | -0.000 | 0.000 | 0.000 |
| 340.976 | 440.800 | 329.159 | -0.000 | 0.000 | 0.000 |
| 326.616 | 326.616 | 454.399 | -0.000 | 0.000 | 0.000 |
| -0.663 | -0.663 | 1.330 | 39.549 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.860 | 0.000 |
| -0.668 | -0.668 | 1.339 | -0.000 | 0.000 | 49.581 |
| 441.088 | 343.203 | 326.632 | -0.000 | -0.000 | -0.000 |
| 345.274 | 441.820 | 323.823 | -0.000 | 0.000 | 0.000 |
| 327.938 | 327.938 | 451.742 | 0.000 | 0.000 | -0.000 |
| 0.663 | 0.663 | -1.330 | 39.549 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.860 | 0.000 |
| 0.668 | 0.668 | -1.339 | 0.000 | -0.000 | 49.581 |
| 638.118 | 274.854 | 197.203 | 21.838 | -0.000 | 0.000 |
| 246.712 | 91.202 | 772.905 | 14.489 | -0.000 | -0.000 |
| 197.260 | 197.260 | 714.251 | -0.000 | -0.000 | 0.000 |
| 21.838 | -21.838 | -0.000 | 104.713 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 104.713 | 21.838 |
| -177.765 | -177.765 | 355.927 | 0.000 | -18.676 | -107.651 |
| 638.118 | 274.854 | 197.203 | 21.838 | -0.000 | -0.000 |
| 408.248 | 532.521 | 169.375 | -5.656 | 0.000 | -0.000 |
| 580.243 | 580.243 | -52.572 | -0.000 | 0.000 | 0.000 |
| 21.838 | -21.838 | 0.000 | 104.713 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 104.713 | 21.838 |
| 177.765 | 177.765 | -355.927 | -0.000 | -18.676 | -107.651 |
| 421.305 | 336.821 | 352.223 | 2.316 | -0.000 | -0.000 |
| 274.854 | 638.118 | 197.203 | -21.838 | 0.000 | 0.000 |
| 325.312 | 325.312 | 457.862 | 0.000 | -0.000 | -0.000 |
| -8.872 | -9.859 | 18.752 | 29.201 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.713 | 21.838 |
| -8.846 | -8.846 | 17.713 | -0.000 | 1.676 | 37.670 |
| 436.506 | 354.717 | 319.089 | 2.127 | 0.000 | 0.000 |
| 354.716 | 436.507 | 319.088 | -2.128 | 0.000 | 0.000 |
| 197.260 | 197.260 | 714.251 | 0.000 | -0.000 | -0.000 |
| 9.859 | 8.872 | -18.752 | 29.201 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.713 | 21.838 |
| 8.846 | 8.846 | -17.713 | 0.000 | 1.676 | 37.670 |
| 638.121 | 274.855 | 197.203 | 21.838 | -0.000 | -0.000 |
| 274.855 | 638.121 | 197.203 | -21.838 | 0.000 | 0.000 |
| 326.814 | 326.814 | 454.857 | -0.000 | 0.000 | 0.000 |
| 21.838 | -21.837 | 0.000 | 104.713 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 104.713 | 21.838 |
| 0.000 | 0.000 | 0.000 | 0.000 | 21.838 | 181.632 |
| 638.116 | 274.853 | 197.202 | 21.838 | 0.000 | 0.000 |
| 342.787 | 440.764 | 326.766 | -3.261 | 0.000 | 0.000 |
| 328.209 | 328.209 | 452.056 | 0.000 | -0.000 | 0.000 |
| 21.837 | -21.838 | -0.000 | 104.713 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 104.713 | 21.838 |
| -0.000 | -0.000 | -0.000 | -0.000 | 21.838 | 181.632 |
| 1426.713 | 1419.761 | 1419.770 | -0.000 | -0.001 | -0.000 |
| 1419.759 | 1426.715 | 1419.770 | -0.000 | -0.001 | -0.000 |
| 1419.759 | 1419.762 | 1426.724 | -0.000 | -0.001 | -0.000 |
| 893.432 | 893.432 | 893.434 | -6.979 | -0.009 | -0.000 |
| 893.432 | 893.433 | 893.434 | 0.001 | -6.989 | -0.000 |
| 893.432 | 893.433 | 893.434 | 0.001 | -0.009 | -6.980 |
| -868.564 | -866.453 | -866.454 | -0.001 | 0.009 | 0.000 |
| -866.453 | -868.564 | -866.454 | -0.001 | 0.009 | 0.000 |
| -866.453 | -866.453 | -868.566 | -0.001 | 0.009 | 0.000 |
| -893.432 | -893.433 | -893.434 | -6.980 | 0.009 | 0.000 |
| -893.432 | -893.433 | -893.434 | -0.001 | -6.970 | 0.000 |
| -893.432 | -893.433 | -893.434 | -0.001 | 0.009 | -6.979 |
| 351.862 | 285.040 | 285.039 | 0.000 | -0.002 | -0.000 |
| 113.033 | 135.520 | 113.033 | 0.000 | -0.001 | -0.000 |
| 113.032 | 113.032 | 135.520 | 0.000 | -0.001 | -0.000 |
| 129.197 | 129.197 | 129.198 | -6.067 | -0.000 | -0.000 |
| 129.197 | 129.197 | 129.198 | -0.000 | -6.067 | -0.000 |
| 129.197 | 129.197 | 129.198 | -0.000 | -0.000 | -6.067 |
| -1.523 | -48.158 | -48.158 | 0.000 | 0.000 | 0.000 |
| -48.158 | -1.523 | -48.159 | 0.000 | 0.000 | 0.000 |
| -48.158 | -48.158 | -1.523 | 0.000 | 0.000 | 0.000 |
| -129.197 | -129.197 | -129.198 | -6.067 | 0.000 | 0.000 |
| -129.197 | -129.197 | -129.198 | 0.000 | -6.067 | 0.000 |
| -129.197 | -129.197 | -129.198 | 0.000 | 0.000 | -6.067 |
| 139.460 | 55.241 | 55.241 | 0.000 | -0.000 | -0.000 |
| 55.240 | 139.460 | 55.241 | 0.000 | -0.000 | -0.000 |
| 55.241 | 55.240 | 139.460 | 0.000 | -0.000 | -0.000 |
| -159178.955 | -171209.784 | -149866.297 | -364757.524 | -93470.993 | 4452.423 |
| -148020.700 | -159076.780 | -171362.477 | 13775.069 | -364422.980 | 2516.345 |
| -168643.108 | -147431.674 | -161832.686 | 16762.680 | -93940.264 | -364885.480 |
| 110.322 | 40.699 | 40.699 | -0.000 | 0.000 | 0.000 |
| 40.699 | 110.322 | 40.699 | -0.000 | 0.000 | 0.000 |
| 40.699 | 40.699 | 110.322 | -0.000 | 0.000 | 0.000 |
| 159305.362 | 171296.086 | 149738.772 | -364319.488 | 93948.825 | -572.244 |
| 148027.047 | 159145.169 | 171223.391 | -14939.590 | -364434.735 | -2973.105 |
| 168610.779 | 147337.693 | 161925.761 | -8765.047 | 93846.181 | -364267.022 |
| 357.194 | 128.132 | 96.758 | -0.000 | -0.000 | -0.000 |
| 126.836 | 175.893 | 40.606 | -0.000 | -0.000 | 0.000 |
| -218.421 | 318.248 | -223.675 | 0.000 | 0.000 | -0.000 |
| -254.457 | 235.901 | -328.219 | 0.453 | 0.000 | -0.000 |
| -330.462 | 113.494 | -684.025 | 0.000 | 3.532 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 82.927 |
| 508.012 | -152.305 | 419.239 | -0.000 | -0.000 | 0.000 |
| 126.836 | 175.893 | 40.606 | 0.000 | -0.000 | -0.000 |
| 297.932 | -157.477 | 452.204 | -0.000 | -0.000 | 0.000 |
| 254.457 | -235.901 | 328.219 | 0.453 | -0.000 | 0.000 |
| 330.462 | -113.494 | 684.025 | -0.000 | 3.532 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 82.927 |
| 357.194 | 128.132 | 96.758 | -0.000 | 0.000 | 0.000 |
| 29.382 | 65.268 | -68.398 | -0.000 | 0.000 | -0.000 |
| 3.946 | 104.227 | 37.093 | 0.000 | 0.000 | -0.000 |
| -23.757 | 22.025 | -30.644 | 1.734 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 45.817 | -0.000 |
| -32.806 | 11.317 | -67.659 | 0.000 | 0.000 | -18.190 |
| 273.772 | 85.336 | 98.461 | -0.000 | 0.000 | -0.000 |
| 102.617 | 49.023 | 86.623 | -0.000 | -0.000 | 0.000 |
| 82.977 | 18.475 | 97.702 | -0.000 | -0.000 | 0.000 |
| 23.757 | -22.025 | 30.644 | 1.734 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 45.817 | -0.000 |
| 32.806 | -11.317 | 67.659 | -0.000 | -0.000 | -18.190 |
| 258.009 | 111.133 | 67.953 | 0.000 | 0.000 | -0.000 |
| 95.412 | 70.655 | 54.264 | 0.000 | 0.000 | -0.000 |
| -3.215 | 2.510 | -6.830 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 19.769 | 0.000 | 0.000 |
| -1.084 | 1.712 | -6.452 | 0.000 | -3.780 | -0.000 |
| -1.149 | 1.830 | -6.948 | 0.000 | 0.000 | -17.973 |
| 236.665 | 55.625 | 36.149 | -0.000 | -0.000 | 0.000 |
| 65.854 | 53.106 | 6.841 | -0.000 | -0.000 | 0.000 |
| 22.693 | 10.748 | 10.606 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 19.769 | -0.000 | -0.000 |
| 1.084 | -1.712 | 6.452 | 0.000 | -3.780 | -0.000 |
| 1.149 | -1.830 | 6.948 | -0.000 | -0.000 | -17.973 |
| 260.740 | 229.384 | 156.680 | 0.000 | 0.000 | 0.000 |
| 395.671 | 457.973 | 418.876 | 0.000 | -0.000 | 0.000 |
| 87.376 | 246.600 | 597.132 | 0.000 | 0.000 | 0.000 |
| 559.897 | -75.830 | -10.900 | 178.525 | -0.000 | 0.000 |
| 221.760 | -35.422 | 23.686 | 0.000 | 9.727 | 0.000 |
| 221.660 | -35.406 | 23.675 | 0.000 | 0.000 | 9.201 |
| -220.191 | 311.312 | 103.095 | -0.000 | 0.000 | -0.000 |
| -446.169 | 687.831 | 254.212 | -0.000 | 0.000 | -0.000 |
| 87.376 | 246.600 | 597.132 | -0.000 | 0.000 | -0.000 |
| -559.897 | 75.830 | 10.900 | 178.525 | 0.000 | -0.000 |
| -221.760 | 35.422 | -23.686 | -0.000 | 9.727 | -0.000 |
| -221.660 | 35.406 | -23.675 | -0.000 | 0.000 | 9.201 |
| -51.212 | 248.431 | 185.473 | 0.000 | -0.000 | 0.000 |
| 168.440 | 530.698 | 346.027 | 0.000 | -0.000 | 0.000 |
| 87.376 | 246.600 | 597.133 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 193.910 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 19.722 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 35.248 |
| -122.557 | 312.686 | 106.158 | -0.000 | 0.000 | -0.000 |
| 92.232 | 556.025 | 325.330 | 0.000 | 0.000 | -0.000 |
| 50.775 | 346.834 | 462.131 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 193.910 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 19.722 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 35.248 |
| -112.245 | 250.785 | 196.498 | 0.000 | 0.000 | 0.000 |
| 146.266 | 548.480 | 324.672 | 0.000 | -0.000 | 0.000 |
| 67.998 | 334.502 | 477.279 | -0.000 | -0.000 | -0.000 |
| -18861374.112 | -3314640.584 | -1978403.719 | 401157.842 | -0.000 | -0.000 |
| -1.826 | 3.034 | -3.488 | -0.000 | 8.750 | 0.000 |
| 4.622 | -0.646 | 0.012 | -0.000 | 0.000 | 11.623 |
| -130.637 | 255.038 | 194.587 | -0.000 | 0.000 | -0.000 |
| 107.973 | 617.276 | 236.990 | -0.000 | -0.000 | -0.000 |
| 93.069 | 331.238 | 474.283 | -0.000 | -0.000 | -0.000 |
| 18861374.067 | 3314640.599 | 1978403.720 | 401157.795 | -0.000 | 0.000 |
| 1.826 | -3.034 | 3.488 | 0.000 | 8.750 | -0.000 |
| -4.622 | 0.646 | -0.012 | 0.000 | 0.000 | 11.623 |
| -816.110 | 733.187 | -331.961 | -0.000 | 0.000 | 0.000 |
| 108.388 | 619.510 | 237.459 | 0.000 | 0.000 | 0.000 |
| 87.470 | 247.149 | 598.159 | 0.000 | 0.000 | 0.000 |
| -1199.330 | 230.205 | -38.996 | 137.211 | 0.000 | 0.000 |
| -780.616 | 429.561 | -432.180 | -0.000 | 8.843 | 0.000 |
| -780.214 | 429.340 | -431.957 | -0.000 | 0.000 | 6.846 |
| 254.738 | 200.382 | 169.774 | -0.000 | -0.000 | -0.000 |
| 108.388 | 619.510 | 237.459 | -0.000 | -0.000 | 0.000 |
| 1427.356 | 115.393 | 457.261 | -0.000 | -0.000 | 0.000 |
| 1199.330 | -230.205 | 38.996 | 137.211 | -0.000 | -0.000 |
| 780.616 | -429.561 | 432.180 | 0.000 | 8.843 | -0.000 |
| 780.214 | -429.340 | 431.957 | 0.000 | -0.000 | 6.846 |
| -155.359 | 354.653 | 57.955 | -0.000 | 0.000 | 0.000 |
| 108.388 | 619.510 | 237.459 | 0.000 | 0.000 | 0.000 |
| -41.221 | 361.721 | 466.809 | -0.000 | 0.000 | -0.000 |
| -117.484 | 29.847 | -14.431 | 93.773 | 0.000 | 0.000 |
| -151.298 | 32.447 | -8.416 | 0.000 | 11.713 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 34.552 |
| 45.480 | 212.023 | 212.991 | -0.000 | -0.000 | 0.000 |
| 279.400 | 523.435 | 331.626 | 0.000 | -0.000 | -0.000 |
| 113.265 | 333.946 | 469.923 | 0.000 | -0.000 | 0.000 |
| 117.484 | -29.847 | 14.431 | 93.773 | -0.000 | -0.000 |
| 151.298 | -32.447 | 8.416 | -0.000 | 11.713 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 34.552 |
| -138.748 | 260.534 | 190.001 | 0.000 | 0.000 | -0.000 |
| 140.710 | 554.025 | 321.557 | 0.000 | 0.000 | -0.000 |
| 85.391 | 333.971 | 474.680 | 0.000 | 0.000 | 0.000 |
| -3.877 | 2.134 | -2.146 | 91.164 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 19.454 | 0.000 |
| -5.840 | 3.214 | -3.233 | -0.000 | 0.000 | 13.814 |
| -104.380 | 249.275 | 197.991 | -0.000 | -0.000 | -0.000 |
| 149.213 | 548.595 | 327.299 | 0.000 | -0.000 | -0.000 |
| 122.685 | 327.147 | 474.962 | -0.000 | -0.000 | -0.000 |
| 3.877 | -2.134 | 2.146 | 91.164 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 19.454 | 0.000 |
| 5.840 | -3.214 | 3.233 | -0.000 | -0.000 | 13.814 |
| 22286666.138 | 21793012.411 | 19127805.844 | 157492.558 | 147136.276 | -249236.882 |
| 22286638.231 | 21793100.176 | 19127753.248 | 157599.125 | 147599.664 | -249180.020 |
| 22286654.165 | 21793057.787 | 19127943.394 | 157533.229 | 147230.541 | -249222.991 |
| 22286569.885 | 21792995.857 | 19127766.889 | 157580.316 | 146977.360 | -249256.743 |
| 22286570.178 | 21792994.024 | 19127772.173 | 157494.117 | 147290.664 | -249235.507 |
| 22286602.899 | 21793007.686 | 19127727.454 | 157547.725 | 147364.753 | -249169.784 |
| -22286518.426 | -21792989.688 | -19127676.337 | -157573.275 | -147292.688 | 249216.221 |
| -22286542.584 | -21792902.531 | -19127733.857 | -157454.713 | -146863.178 | 249269.830 |
| -22286536.241 | -21792945.693 | -19127532.491 | -157539.414 | -147272.718 | 249220.731 |
| -22286593.951 | -21793002.526 | -19127739.659 | -157442.203 | -147370.222 | 249208.998 |
| -22286623.717 | -21793013.416 | -19127699.084 | -157579.125 | -147309.976 | 249197.244 |
| -22286585.948 | -21792997.226 | -19127749.725 | -157520.746 | -147163.997 | 249271.810 |
| 2228558.207 | 2179224.504 | 1913154.367 | 15787.445 | 11648.085 | -25366.069 |
| 2228758.510 | 2179376.774 | 1912713.009 | 15777.250 | 15749.796 | -24773.452 |
| 2228724.001 | 2179359.952 | 1912972.119 | 15756.462 | 14795.128 | -24912.776 |
| 2228614.784 | 2179298.116 | 1912832.765 | 15843.748 | 13947.703 | -25030.646 |
| 2228644.698 | 2179293.617 | 1912796.030 | 15737.226 | 14758.459 | -24935.074 |
| 2228665.954 | 2179305.565 | 1912764.885 | 15762.743 | 14813.964 | -24873.099 |
| -2228588.935 | -2179289.909 | -1912704.792 | -15800.654 | -14822.053 | 24909.514 |
| -2228622.840 | -2179205.596 | -1912752.457 | -15736.152 | -14244.640 | 24982.803 |
| -2228697.533 | -2179225.764 | -1912442.340 | -15750.109 | -15830.027 | 24757.049 |
| -2228675.044 | -2179300.166 | -1912756.554 | -15703.498 | -14837.502 | 24907.246 |
| -2228482.801 | -2179210.788 | -1913096.080 | -15858.669 | -11408.828 | 25383.387 |
| -2228634.074 | -2179291.023 | -1912807.283 | -15748.201 | -14257.093 | 25021.719 |
| 222898.971 | 217868.629 | 191370.856 | 1546.898 | 1168.684 | -2569.362 |
| 222901.360 | 217969.873 | 191242.639 | 1686.994 | 1540.635 | -2501.800 |
| 222871.643 | 217854.066 | 191530.423 | 1561.380 | 2056.492 | -2436.834 |
| 222846.008 | 217895.562 | 191242.802 | 1625.581 | 1632.340 | -2475.043 |
| 222924.148 | 217896.376 | 191221.965 | 1714.075 | 1686.718 | -2474.096 |
| 222882.258 | 217865.594 | 191203.261 | 1680.922 | 1511.479 | -2466.296 |
| -222829.035 | -217905.991 | -191171.423 | -1680.269 | -1532.613 | 2475.362 |
| -222895.560 | -217832.635 | -191178.027 | -1624.806 | -1507.776 | 2475.599 |
| -222788.842 | -217864.531 | -191104.779 | -1580.046 | -1166.425 | 2537.535 |
| -222853.496 | -217917.114 | -191302.225 | -1507.949 | -1212.157 | 2526.808 |
| -222888.431 | -217913.493 | -191250.631 | -1553.569 | -1445.895 | 2464.589 |
| -222848.736 | -217901.443 | -191334.195 | -1601.948 | -1184.785 | 2579.781 |
| 2545.638 | 904.512 | -524.418 | 0.000 | 0.000 | 0.000 |
| 173.428 | 627.880 | 264.989 | -0.000 | -0.000 | -0.000 |
| 2956.152 | 583.885 | 885.749 | -0.000 | -0.000 | 0.000 |
| 2315.984 | 570.177 | -851.663 | 13.403 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 166.029 | -0.000 |
| 1973.963 | 331.897 | -300.192 | 0.000 | 0.000 | 46.822 |
| -2076.773 | -237.376 | 1175.587 | -0.000 | -0.000 | -0.000 |
| -1974.001 | -176.869 | 1180.754 | -0.000 | -0.000 | -0.000 |
| -2023.859 | -245.863 | 1230.595 | -0.000 | 0.000 | -0.000 |
| -2315.984 | -570.177 | 851.663 | 13.403 | -0.000 | 0.000 |
| -2318.637 | -570.359 | 851.004 | -0.000 | 75.882 | 0.000 |
| -1973.963 | -331.897 | 300.192 | -0.000 | 0.000 | 46.822 |
| 623.026 | 177.990 | 255.492 | -0.000 | 0.000 | -0.000 |
| 526.906 | 433.094 | 383.946 | -0.000 | -0.000 | -0.000 |
| 497.220 | 354.730 | 323.963 | 0.000 | -0.000 | -0.000 |
| 227.650 | 19.007 | 44.581 | 2.763 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 166.029 | -0.000 |
| 249.095 | 20.798 | 48.781 | -0.000 | -0.000 | 8.825 |
| 623.025 | 177.990 | 255.492 | 0.000 | -0.000 | 0.000 |
| 71.844 | 386.551 | 296.725 | 0.000 | -0.000 | -0.000 |
| 57.320 | 318.937 | 360.499 | 0.000 | 0.000 | -0.000 |
| -227.650 | -19.007 | -44.581 | 2.763 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 166.029 | 0.000 |
| -249.095 | -20.798 | -48.781 | 0.000 | 0.000 | 8.825 |
| 402.790 | 273.511 | 260.799 | -0.000 | -0.000 | 0.000 |
| 275.862 | 483.647 | 318.927 | -0.000 | 0.000 | -0.000 |
| 273.323 | 324.004 | 426.815 | -0.000 | 0.000 | 0.000 |
| 15.820 | 3.603 | -4.808 | 73.574 | -0.000 | -0.000 |
| 14.521 | 1.213 | 2.844 | 0.000 | 98.216 | -0.000 |
| 15.527 | 1.296 | 3.041 | -0.000 | -0.000 | 35.957 |
| 376.635 | 268.141 | 254.322 | -0.000 | -0.000 | 0.000 |
| 246.796 | 477.639 | 314.028 | -0.000 | 0.000 | 0.000 |
| 257.625 | 305.607 | 428.797 | 0.000 | 0.000 | 0.000 |
| -15.820 | -3.603 | 4.808 | 73.574 | 0.000 | -0.000 |
| -14.521 | -1.213 | -2.844 | 0.000 | 98.216 | 0.000 |
| -15.527 | -1.296 | -3.041 | -0.000 | 0.000 | 35.957 |
| 942.927 | 39.069 | 650.515 | -0.000 | 218.627 | 0.000 |
| 189.824 | 628.152 | 248.542 | 0.000 | -67.837 | -0.000 |
| 860.832 | 48.553 | 828.836 | -0.000 | 230.694 | 0.000 |
| -0.000 | 0.000 | 0.000 | 192.048 | -0.000 | -66.377 |
| 23.836 | -65.351 | 50.478 | 0.000 | 158.705 | -0.000 |
| -0.000 | 0.000 | 0.000 | -68.863 | -0.000 | 97.948 |
| 620.924 | 186.447 | 252.850 | -0.000 | 37.723 | 0.000 |
| 189.824 | 628.152 | 248.542 | -0.000 | -67.837 | 0.000 |
| -305.652 | 562.826 | 61.956 | 0.000 | -185.848 | 0.000 |
| 0.000 | -0.000 | -0.000 | 192.048 | 0.000 | -66.377 |
| -587.794 | 227.964 | -340.163 | 0.000 | -112.052 | -0.000 |
| 0.000 | -0.000 | -0.000 | -68.863 | -0.000 | 97.948 |
| 620.924 | 186.447 | 252.851 | 0.000 | 37.723 | -0.000 |
| 328.912 | 457.840 | 335.322 | 0.000 | -7.167 | -0.000 |
| 339.093 | 282.383 | 469.598 | -0.000 | 40.703 | -0.000 |
| 0.000 | 0.000 | 0.000 | 192.048 | -0.000 | -66.377 |
| 62.453 | -42.777 | 51.569 | -0.000 | 119.299 | 0.000 |
| 0.000 | 0.000 | 0.000 | -68.863 | 0.000 | 97.948 |
| 354.840 | 294.088 | 211.493 | -0.000 | -17.785 | 0.000 |
| 189.824 | 628.152 | 248.542 | 0.000 | -67.837 | 0.000 |
| 204.063 | 328.752 | 406.578 | -0.000 | -7.040 | 0.000 |
| -0.000 | -0.000 | -0.000 | 192.048 | 0.000 | -66.377 |
| -40.365 | -10.765 | -17.415 | -0.000 | 78.667 | 0.000 |
| -0.000 | -0.000 | -0.000 | -68.863 | -0.000 | 97.948 |
| 380.192 | 271.032 | 284.360 | -0.000 | 15.045 | 0.000 |
| 272.640 | 487.325 | 310.573 | -0.000 | -26.103 | 0.000 |
| 261.148 | 309.728 | 439.903 | 0.000 | 18.076 | -0.000 |
| 5.252 | -2.213 | 3.423 | 74.219 | 2.052 | -7.481 |
| 11.336 | -26.356 | 19.984 | 0.000 | 99.222 | -0.000 |
| 5.457 | -2.406 | 4.284 | -8.502 | 2.775 | 39.513 |
| 368.599 | 275.244 | 277.181 | 0.000 | 10.687 | -0.000 |
| 269.942 | 488.118 | 299.457 | -0.000 | -29.953 | 0.000 |
| 249.816 | 312.707 | 434.773 | -0.000 | 13.597 | 0.000 |
| -5.252 | 2.213 | -3.423 | 74.219 | -2.052 | -7.481 |
| 1.526 | -23.374 | 14.054 | -0.000 | 95.010 | 0.000 |
| -5.457 | 2.406 | -4.284 | -8.502 | -2.775 | 39.513 |
| 717.319 | 231.506 | 174.039 | -0.000 | -0.000 | 0.000 |
| 231.506 | 717.319 | 174.039 | -0.000 | -0.000 | 0.000 |
| 276.376 | 276.376 | 509.855 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 145.412 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 145.412 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 45.616 |
| 383.993 | 258.149 | 276.376 | 0.000 | -0.000 | 0.000 |
| 258.149 | 383.993 | 276.376 | 0.000 | 0.000 | 0.000 |
| 276.376 | 276.376 | 509.855 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 145.412 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 145.412 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 45.616 |
| 423.275 | 147.873 | 300.065 | 0.000 | -0.000 | -0.000 |
| 147.873 | 423.275 | 300.065 | -0.000 | 0.000 | -0.000 |
| 276.375 | 276.375 | 509.855 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 75.171 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 75.171 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 0.000 | 55.893 |
| 423.281 | 147.875 | 300.063 | -0.000 | -0.000 | 0.000 |
| 147.875 | 423.281 | 300.063 | -0.000 | -0.000 | 0.000 |
| 276.376 | 276.376 | 509.855 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 75.171 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 75.171 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 55.893 |
| 383.960 | 258.137 | 276.392 | -0.000 | -0.000 | -0.000 |
| 258.137 | 383.960 | 276.392 | -0.000 | -0.000 | -0.000 |
| 276.372 | 276.372 | 509.860 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 75.171 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 75.171 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 45.616 |
| 384.025 | 258.161 | 276.360 | -0.000 | 0.000 | 0.000 |
| 258.161 | 384.025 | 276.360 | -0.000 | -0.000 | 0.000 |
| 276.380 | 276.380 | 509.850 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 75.171 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 75.171 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 45.616 |
| 382.803 | 256.718 | 276.670 | -0.000 | -0.000 | 0.000 |
| 256.718 | 382.803 | 276.670 | -0.000 | -0.000 | -0.000 |
| 276.483 | 276.483 | 510.646 | -0.000 | -0.000 | 0.000 |
| 0.020 | 0.020 | -0.013 | 74.957 | -0.000 | -0.000 |
| 0.020 | 0.020 | -0.013 | -0.000 | 74.957 | -0.000 |
| 0.029 | 0.029 | -0.019 | -0.000 | -0.000 | 45.937 |
| 382.774 | 256.688 | 276.689 | 0.000 | -0.000 | 0.000 |
| 256.688 | 382.774 | 276.689 | 0.000 | -0.000 | 0.000 |
| 174.603 | 174.603 | 578.304 | 0.000 | 0.000 | 0.000 |
| -0.020 | -0.020 | 0.013 | 74.957 | 0.000 | 0.000 |
| -0.020 | -0.020 | 0.013 | 0.000 | 74.957 | 0.000 |
| -0.029 | -0.029 | 0.019 | 0.000 | 0.000 | 45.937 |
| 422.050 | 146.548 | 300.220 | -0.000 | -0.000 | -0.000 |
| 146.548 | 422.050 | 300.220 | -0.000 | -0.000 | -0.000 |
| 276.469 | 276.469 | 510.655 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 145.264 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 145.264 | -0.000 |
| 0.003 | 0.003 | -0.002 | -0.000 | -0.000 | 45.937 |
| 422.054 | 146.546 | 300.219 | 0.000 | 0.000 | 0.000 |
| 146.546 | 422.054 | 300.219 | 0.000 | 0.000 | -0.000 |
| 276.467 | 276.467 | 510.656 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 145.264 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 145.264 | 0.000 |
| -0.003 | -0.003 | 0.002 | 0.000 | 0.000 | 45.937 |
| 382.756 | 256.691 | 276.695 | -0.000 | -0.000 | -0.000 |
| 256.691 | 382.756 | 276.695 | -0.000 | 0.000 | -0.000 |
| 276.465 | 276.465 | 510.660 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 74.957 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 74.957 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 45.937 |
| 382.821 | 256.715 | 276.663 | 0.000 | 0.000 | 0.000 |
| 256.715 | 382.821 | 276.663 | 0.000 | -0.000 | -0.000 |
| 276.472 | 276.472 | 510.651 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 74.957 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 74.957 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 45.937 |
| 444.741 | 172.864 | 172.864 | 0.000 | 0.000 | -0.000 |
| 172.864 | 444.741 | 172.864 | 0.000 | 0.000 | 0.000 |
| 172.864 | 172.864 | 444.741 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 92.240 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 92.240 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 92.240 |
| 212.102 | 262.261 | 262.261 | 0.000 | -0.000 | 0.000 |
| 262.263 | 212.100 | 262.262 | 0.000 | -0.000 | 0.000 |
| 262.263 | 262.262 | 212.100 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 92.240 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 92.240 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 92.240 |
| 444.741 | 172.864 | 172.864 | -0.000 | 0.000 | -0.000 |
| 172.864 | 444.741 | 172.864 | -0.000 | 0.000 | 0.000 |
| 172.864 | 172.864 | 444.741 | -0.000 | 0.000 | -0.000 |
| 40367098.655 | 20117139.508 | 20133924.795 | -14167523.763 | 0.001 | -0.002 |
| -5801252.677 | 13586695.427 | -4986868.445 | 443912.012 | -13827777.392 | -97483.984 |
| -5398329.445 | -4924756.967 | 12619245.928 | -21175.836 | -98014.746 | -13978269.429 |
| 212.092 | 262.430 | 262.430 | 0.000 | 0.000 | 0.000 |
| 262.430 | 212.092 | 262.430 | 0.000 | 0.000 | 0.000 |
| 262.430 | 262.430 | 212.092 | 0.000 | 0.000 | 0.000 |
| -8162360.099 | 9695841.800 | 8825830.305 | -14947438.992 | -155039.567 | -483890.507 |
| 6512642.374 | -11654920.778 | 6938087.626 | 227677.782 | -14420092.511 | 99098.623 |
| 9555501.829 | 8562682.713 | -9405186.253 | -203993.477 | -640013.500 | -12713613.116 |
| 26.031 | 363.218 | 363.218 | -0.000 | 0.000 | -0.000 |
| 363.218 | 26.031 | 363.218 | -0.000 | 0.000 | -0.000 |
| 363.218 | 363.218 | 26.031 | -0.000 | 0.000 | -0.000 |
| 3368351.134 | 1392927.039 | 1392921.169 | -2935005.547 | 0.000 | 0.000 |
| 1392927.029 | 3368351.142 | 1392921.159 | -0.000 | -2935005.518 | -0.000 |
| 1392927.067 | 1392921.198 | 3368351.160 | -0.000 | -0.000 | -2935005.528 |
| 212.108 | 262.439 | 262.439 | 0.000 | 0.000 | 0.000 |
| 262.439 | 212.108 | 262.439 | 0.000 | 0.000 | 0.000 |
| 262.439 | 262.439 | 212.108 | 0.000 | -0.000 | 0.000 |
| -3368351.120 | -1392927.021 | -1392921.152 | -2935005.550 | 0.000 | -0.000 |
| -1392927.044 | -3368351.125 | -1392921.174 | -0.000 | -2935005.567 | 0.000 |
| -1392927.028 | -1392921.159 | -3368351.094 | 0.000 | -0.000 | -2935005.546 |
| 171.011 | 118.684 | 118.684 | 0.000 | -0.000 | 0.000 |
| 118.684 | 171.011 | 118.684 | 0.000 | -0.000 | 0.000 |
| 118.684 | 118.684 | 171.011 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 114.577 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 114.577 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 114.577 |
| 171.014 | 118.686 | 118.686 | -0.000 | 0.000 | -0.000 |
| 118.686 | 171.014 | 118.686 | -0.000 | 0.000 | -0.000 |
| 169.204 | 169.204 | 360.881 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 114.577 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 114.577 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 0.000 | 114.577 |
| 230.289 | 140.653 | 140.653 | 0.000 | -0.000 | 0.000 |
| 140.653 | 230.289 | 140.653 | 0.000 | -0.000 | 0.000 |
| 140.653 | 140.653 | 230.289 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 41.637 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 41.637 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 41.637 |
| 107.095 | 41.881 | 41.881 | -0.000 | 0.000 | -0.000 |
| 169.204 | 360.881 | 169.204 | -0.000 | 0.000 | 0.000 |
| 169.204 | 169.204 | 360.881 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 41.637 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 41.637 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 41.637 |
| 230.337 | 140.664 | 140.664 | 0.000 | -0.000 | 0.000 |
| 140.664 | 230.337 | 140.664 | 0.000 | -0.000 | 0.000 |
| 140.664 | 140.664 | 230.337 | 0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 41.637 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 51.950 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 41.637 |
| 127.330 | 82.693 | 82.694 | -0.000 | 0.000 | -0.000 |
| 82.694 | 127.330 | 82.693 | -0.000 | 0.000 | -0.000 |
| 82.693 | 82.694 | 127.330 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 41.637 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 51.950 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 41.637 |
| -216.276 | -243.843 | -243.830 | 0.017 | 0.014 | 0.040 |
| 77.657 | 206.032 | 77.657 | -0.000 | -0.000 | -0.000 |
| 77.657 | 77.657 | 206.032 | 0.000 | -0.000 | -0.000 |
| -306.105 | -306.054 | -306.041 | 14.320 | 0.014 | 0.039 |
| -306.105 | -306.054 | -306.041 | 0.016 | 14.318 | 0.039 |
| -954.980 | -954.905 | -954.885 | 0.025 | 0.021 | 6.695 |
| 206.032 | 77.657 | 77.657 | 0.000 | -0.000 | -0.000 |
| 77.657 | 206.032 | 77.657 | 0.000 | -0.000 | 0.000 |
| 77.657 | 77.657 | 206.032 | -0.000 | 0.000 | 0.000 |
| 765.209 | 765.235 | 765.242 | 32.022 | 0.007 | 0.021 |
| 954.977 | 954.902 | 954.882 | -0.025 | 6.620 | -0.059 |
| 306.102 | 306.052 | 306.039 | -0.016 | -0.014 | 14.265 |
| 65.514 | 37.952 | 37.954 | 0.002 | 0.001 | 0.004 |
| 37.946 | 65.514 | 37.952 | 0.002 | 0.001 | 0.004 |
| 37.945 | 37.950 | 65.513 | 0.002 | 0.001 | 0.004 |
| -0.000 | 0.000 | 0.000 | 50.014 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 50.014 | 0.000 |
| -27.714 | -27.709 | -27.708 | 0.001 | 0.001 | 17.687 |
| 206.032 | 77.657 | 77.657 | -0.000 | 0.000 | 0.000 |
| 77.657 | 206.032 | 77.657 | 0.000 | 0.000 | -0.000 |
| 77.657 | 77.657 | 206.032 | 0.000 | 0.000 | 0.000 |
| 27.712 | 27.708 | 27.706 | 17.683 | -0.001 | -0.004 |
| -0.000 | -0.000 | -0.000 | -0.000 | 50.014 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 50.014 |
| 47.128 | 19.117 | 19.117 | 0.000 | 0.000 | 0.000 |
| 19.117 | 47.128 | 19.118 | 0.000 | 0.000 | 0.000 |
| 19.117 | 19.117 | 47.128 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 50.014 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 50.014 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 50.014 |
| 87.330 | 51.705 | 51.705 | -0.000 | -0.000 | -0.000 |
| 51.706 | 87.329 | 51.705 | -0.000 | -0.000 | -0.000 |
| 51.706 | 51.705 | 87.329 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 50.014 | 0.000 | -0.000 |
| 7.388 | 7.388 | 7.388 | -0.000 | 20.678 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 50.014 |
| -332534.727 | -3453996.329 | 25512035.298 | 95268.588 | -233650.177 | -559193.896 |
| -433159.668 | -3246292.608 | 25075486.450 | 72040.266 | -144096.966 | -443878.570 |
| -338400.052 | -2934664.203 | 24996943.467 | 59993.993 | -89901.177 | -404126.069 |
| -535259.415 | -3037875.402 | 24689679.720 | 39062.316 | -121392.893 | -356539.780 |
| -340176.778 | -3042208.121 | 24986618.044 | 23643.422 | 38403.047 | -643559.222 |
| -410656.551 | -3210869.478 | 25090298.028 | 52200.096 | -43613.785 | -484063.321 |
| 383261.143 | 2994631.259 | -24943794.430 | -66691.442 | 91212.129 | 449542.326 |
| 321535.856 | 3076827.836 | -25212913.774 | -49165.976 | 61818.177 | 439973.428 |
| 325493.400 | 2968657.363 | -25064560.468 | -54867.320 | 108828.449 | 422376.571 |
| 378382.144 | 2975448.180 | -24938123.592 | -71543.055 | 69237.405 | 426845.069 |
| 303414.895 | 3138861.689 | -25111013.999 | -66186.142 | -124545.706 | 511349.157 |
| 370304.582 | 3254154.255 | -25266740.877 | -87861.034 | -4187.296 | 318754.178 |
| -42766.118 | -312437.269 | 2500055.497 | 3096.537 | -21822.324 | -26808.843 |
| -37531.967 | -344947.534 | 2532929.688 | 5626.403 | 8757.918 | -35286.329 |
| -48263.141 | -319245.115 | 2501832.938 | 3751.411 | -19118.671 | -34673.892 |
| -43728.577 | -352167.283 | 2531866.996 | 5320.070 | -11839.495 | -63556.855 |
| -29461.842 | -331020.303 | 2509551.623 | 7584.865 | -34899.043 | -45824.156 |
| -57315.799 | -297012.716 | 2453550.459 | 5754.094 | -6957.222 | -40161.721 |
| 33612.298 | 302662.880 | -2498191.071 | -6431.188 | -1313.931 | 22336.726 |
| 37742.995 | 341359.503 | -2532257.512 | -8041.096 | 17618.825 | 56045.787 |
| 35599.485 | 323309.529 | -2519372.665 | -7628.447 | 17876.027 | 51566.998 |
| 62447.616 | 292248.581 | -2438591.283 | -6002.513 | 7818.368 | 40718.968 |
| 34792.250 | 312395.303 | -2507846.821 | -8012.527 | -4321.147 | 27004.860 |
| 53696.071 | 298913.679 | -2470749.643 | -4862.550 | 8029.521 | 43560.311 |
| -3642.301 | -32336.694 | 251784.468 | 218.493 | 156.882 | -5122.378 |
| -3520.175 | -31066.739 | 252337.694 | 357.592 | -365.710 | -4540.642 |
| -3482.961 | -29579.427 | 250206.981 | 656.509 | -762.210 | -4355.598 |
| -3582.275 | -31471.791 | 251507.282 | 839.564 | -927.490 | -4114.596 |
| -4933.579 | -32575.815 | 249728.798 | 568.415 | -1975.248 | -4994.510 |
| -3534.846 | -31095.283 | 250432.043 | 606.443 | -1875.476 | -2457.039 |
| 4689.784 | 36392.571 | -252666.132 | -778.679 | 3320.560 | 7844.757 |
| 5046.700 | 34463.005 | -250624.674 | -43.322 | 239.977 | 3507.604 |
| 3290.187 | 30999.351 | -251371.339 | -846.643 | 1305.637 | 4239.204 |
| 2753.450 | 33522.743 | -253844.088 | -804.389 | 2445.958 | 3649.293 |
| 3177.587 | 32260.782 | -253240.181 | -339.838 | 136.209 | 4859.469 |
| 4477.675 | 33320.497 | -251633.266 | -914.208 | 2045.068 | 5712.007 |
| 198.671 | 150.238 | -830.315 | 0.000 | -0.000 | 0.000 |
| 150.238 | 198.671 | -830.315 | 0.000 | -0.000 | 0.000 |
| 102.318 | 102.318 | 215.159 | 0.000 | -0.000 | -0.000 |
| 50.950 | 50.950 | -922.019 | 8.693 | -0.000 | 0.000 |
| 50.950 | 50.950 | -922.019 | 0.000 | 8.692 | 0.000 |
| 50.951 | 50.951 | -922.037 | 0.000 | -0.000 | 15.564 |
| 95.908 | 48.214 | 1013.739 | -0.000 | 0.000 | -0.000 |
| 48.214 | 95.908 | 1013.739 | -0.000 | 0.000 | -0.000 |
| 102.318 | 102.318 | 215.159 | 0.000 | 0.000 | 0.000 |
| -50.950 | -50.950 | 922.019 | 8.692 | 0.000 | -0.000 |
| -50.950 | -50.950 | 922.019 | -0.000 | 8.693 | -0.000 |
| -50.951 | -50.951 | 922.037 | -0.000 | 0.000 | 15.564 |
| 156.404 | 104.880 | -0.148 | 0.000 | -0.000 | 0.000 |
| 104.880 | 156.404 | -0.148 | 0.000 | -0.000 | 0.000 |
| 179.636 | 179.636 | -86.217 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 64.920 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 64.920 | -0.000 |
| 5.073 | 5.073 | -91.813 | 0.000 | -0.000 | 15.782 |
| 244.717 | 112.655 | 102.408 | -0.000 | 0.000 | 0.000 |
| 112.655 | 244.717 | 102.408 | -0.000 | 0.000 | 0.000 |
| -57.909 | -57.909 | 280.065 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 64.920 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 64.920 | 0.000 |
| -5.073 | -5.073 | 91.813 | -0.000 | 0.000 | 15.782 |
| 92.599 | 63.322 | 34.398 | 0.000 | -0.000 | 0.000 |
| 63.322 | 92.599 | 34.398 | 0.000 | -0.000 | 0.000 |
| 63.350 | 63.350 | 55.552 | 0.000 | -0.000 | 0.000 |
| 11.877 | 11.877 | -34.969 | 4.106 | -0.000 | 0.000 |
| 11.877 | 11.877 | -34.969 | 0.000 | 4.106 | 0.000 |
| 0.440 | 0.440 | -7.961 | 0.000 | -0.000 | 22.607 |
| 37.405 | 17.292 | 54.252 | -0.000 | 0.000 | -0.000 |
| 17.292 | 37.405 | 54.252 | -0.000 | 0.000 | -0.000 |
| 9.301 | 9.301 | 45.647 | -0.000 | 0.000 | -0.000 |
| -11.877 | -11.877 | 34.969 | 4.106 | 0.000 | -0.000 |
| -11.877 | -11.877 | 34.969 | -0.000 | 4.106 | -0.000 |
| -0.440 | -0.440 | 7.961 | -0.000 | 0.000 | 22.607 |
| 2566.514 | 1481.685 | 962.162 | 0.000 | 0.000 | 0.009 |
| 2513.367 | 1530.847 | 970.264 | 0.000 | 0.000 | 0.009 |
| 104.698 | 99.553 | 223.020 | -0.000 | -0.000 | -0.000 |
| 1492.634 | 2836.717 | 223.519 | 16.825 | 0.000 | 0.007 |
| -0.000 | 0.000 | -0.000 | -0.000 | 62.737 | 0.000 |
| 2444.026 | 1393.733 | 872.663 | 0.000 | 0.000 | 20.070 |
| -1424.414 | -2779.582 | -152.773 | -0.000 | -0.000 | -0.007 |
| 83.397 | 237.791 | 100.481 | -0.000 | 0.000 | 0.000 |
| 104.698 | 99.553 | 223.020 | 0.000 | 0.000 | -0.000 |
| -1492.634 | -2836.717 | -223.519 | 16.825 | -0.000 | -0.007 |
| -0.000 | -0.000 | 0.000 | 0.000 | 62.737 | 0.000 |
| -2444.026 | -1393.733 | -872.663 | -0.000 | -0.000 | 20.053 |
| 268.972 | 338.378 | 148.121 | 0.000 | 0.000 | 0.000 |
| 83.397 | 237.791 | 100.481 | -0.000 | -0.000 | -0.000 |
| 334.330 | 244.205 | 218.159 | 0.000 | 0.000 | 0.001 |
| 148.672 | 283.275 | 22.150 | 16.908 | 0.000 | 0.001 |
| 244.283 | 139.305 | 87.223 | 0.000 | 30.814 | 0.001 |
| 244.142 | 139.224 | 87.173 | 0.000 | 0.000 | 20.100 |
| -98.957 | -281.621 | -129.544 | -0.000 | -0.000 | -0.001 |
| -175.685 | -5.832 | 10.187 | -0.000 | -0.000 | -0.001 |
| 104.698 | 99.553 | 223.020 | 0.000 | 0.000 | 0.000 |
| -148.672 | -283.275 | -22.150 | 16.908 | -0.000 | -0.001 |
| -244.283 | -139.305 | -87.223 | -0.000 | 30.814 | -0.001 |
| -244.141 | -139.224 | -87.173 | -0.000 | -0.000 | 20.099 |
| 120.339 | 92.682 | 90.614 | 0.000 | 0.000 | 0.000 |
| 83.398 | 237.791 | 100.481 | -0.000 | -0.000 | -0.000 |
| 30.232 | 38.871 | 48.652 | 0.000 | 0.000 | 0.000 |
| 12.357 | 27.013 | 1.300 | 19.934 | 0.000 | 0.000 |
| 23.728 | 13.531 | 8.472 | 0.000 | 31.729 | 0.000 |
| 16.747 | 29.734 | 15.830 | 0.000 | 0.000 | 14.296 |
| 113.539 | 68.635 | 81.815 | -0.000 | -0.000 | -0.000 |
| 35.918 | 41.175 | 45.823 | -0.000 | -0.000 | -0.000 |
| -7.068 | -22.531 | 29.808 | -0.000 | -0.000 | -0.000 |
| -12.357 | -27.013 | -1.300 | 19.934 | -0.000 | -0.000 |
| -23.728 | -13.531 | -8.472 | -0.000 | 31.729 | -0.000 |
| -16.747 | -29.734 | -15.830 | -0.000 | -0.000 | 14.296 |
| 227.332 | 61.525 | 67.819 | -0.000 | 3.872 | 0.000 |
| -1784.572 | 363.523 | -1892.423 | 0.000 | 434.524 | -0.000 |
| -480.081 | -45.235 | -135.475 | -0.000 | -24.088 | -0.000 |
| -1791.179 | 310.077 | -1873.570 | 23.356 | 430.857 | -4.186 |
| -1908.663 | 2393.912 | -1568.012 | -0.000 | 794.409 | -0.000 |
| -509.190 | -73.922 | -267.567 | -5.520 | -27.725 | 51.023 |
| 227.332 | 61.525 | 67.819 | 0.000 | 3.872 | -0.000 |
| 549.040 | 223.943 | 296.684 | 0.000 | 29.153 | 0.000 |
| 2102.696 | -477.208 | 2011.510 | 0.000 | -923.692 | 0.000 |
| 1791.179 | -310.077 | 1873.570 | 23.356 | -430.857 | -4.186 |
| 1944.820 | -2383.521 | 1551.073 | 0.000 | -901.992 | 0.000 |
| 509.190 | 73.922 | 267.567 | -5.520 | 27.725 | 51.023 |
| 95.587 | 31.920 | 3.948 | -0.000 | 0.183 | -0.000 |
| 63.562 | 256.773 | 59.484 | -0.000 | -7.746 | 0.000 |
| -177.642 | 2.053 | -143.161 | -0.000 | 41.678 | -0.000 |
| -57.677 | 388.574 | -2.181 | 14.746 | 68.382 | 0.560 |
| -22.043 | -7.849 | -11.660 | 0.000 | 98.414 | -0.000 |
| -207.112 | 43.553 | -211.173 | 5.163 | 91.723 | -20.636 |
| 155.819 | -51.058 | 206.909 | 0.000 | -90.812 | 0.000 |
| 179.243 | 19.519 | 160.120 | -0.000 | -37.867 | 0.000 |
| 163.453 | -64.811 | 195.079 | 0.000 | -39.262 | 0.000 |
| 57.675 | -388.575 | 2.177 | 14.747 | -68.381 | 0.560 |
| 217.421 | -40.753 | 225.002 | 0.000 | -122.883 | 0.000 |
| 207.112 | -43.553 | 211.173 | 5.163 | -91.723 | -20.636 |
| 62.431 | 28.294 | 8.748 | -0.000 | 8.295 | -0.000 |
| 35.647 | 154.364 | 27.767 | -0.000 | 0.697 | -0.000 |
| 25.298 | 28.031 | 129.611 | -0.000 | 3.502 | -0.000 |
| -14.234 | 1.768 | -13.751 | 28.218 | 2.779 | -3.823 |
| -9.380 | 5.393 | -26.763 | -0.000 | 7.084 | -0.000 |
| -12.829 | 2.259 | -12.514 | -1.402 | 4.987 | 18.648 |
| 106.287 | 11.614 | 14.528 | 0.000 | 0.698 | 0.000 |
| 44.690 | 152.098 | 31.897 | 0.000 | 1.552 | 0.000 |
| 26.413 | 7.703 | 53.246 | 0.000 | 3.062 | 0.000 |
| 14.234 | -1.768 | 13.751 | 28.218 | -2.779 | -3.823 |
| 7.591 | 4.568 | 17.327 | -0.000 | 32.406 | 0.000 |
| 12.829 | -2.259 | 12.514 | -1.402 | -4.987 | 18.648 |
| 149.230 | 71.128 | 179.557 | -0.000 | 300.971 | 0.000 |
| 73.951 | 152.181 | 182.774 | 0.000 | 287.351 | 0.000 |
| 95.399 | -208.314 | 145.323 | 0.000 | 268.689 | 0.000 |
| 0.000 | -0.000 | -0.000 | 85.835 | -0.000 | -7.747 |
| -4.848 | -4.506 | -0.935 | 0.000 | 111.258 | -0.000 |
| 65.750 | -243.192 | 3.774 | -0.614 | 277.138 | 45.201 |
| 312.701 | 73.168 | 81.950 | -0.000 | 27.710 | 0.000 |
| 1121.215 | 1107.811 | 168.851 | -0.000 | -13.207 | 0.000 |
| -36.249 | 277.887 | 137.431 | -0.000 | -285.526 | -0.000 |
| -0.000 | -0.000 | 0.000 | 85.834 | 0.000 | -7.747 |
| 154.010 | 1436.863 | -843.631 | 0.000 | 111.224 | -0.000 |
| -65.750 | 243.192 | -3.774 | -0.614 | -277.138 | 45.201 |
| 62.036 | -14.587 | 15.115 | -0.000 | 51.415 | -0.000 |
| 26.930 | 133.061 | 37.589 | 0.000 | 32.536 | 0.000 |
| 37.746 | 12.019 | 145.274 | 0.000 | 19.057 | -0.000 |
| -108.606 | -113.601 | -17.016 | 2.319 | 1.694 | -0.817 |
| 22.907 | -17.283 | -7.637 | 0.000 | 79.866 | 0.000 |
| 6.465 | -23.911 | 0.371 | -0.732 | 27.248 | 46.158 |
| 39.826 | -26.811 | -17.372 | -0.000 | -1.891 | -0.000 |
| 74.655 | 309.578 | 69.800 | -0.000 | -4.378 | 0.000 |
| -13.711 | -8.292 | 79.912 | 0.000 | -25.041 | -0.000 |
| 108.606 | 113.601 | 17.016 | 2.319 | -1.694 | -0.817 |
| -4.848 | -4.506 | -0.935 | -0.000 | 111.258 | 0.000 |
| -6.465 | 23.911 | -0.371 | -0.732 | -27.248 | 46.158 |
| 71.497 | 12.260 | 24.672 | 0.000 | 29.404 | -0.000 |
| 29.109 | 176.243 | 41.992 | 0.000 | 7.166 | -0.000 |
| 11.353 | 15.503 | 73.732 | 0.000 | 8.976 | -0.000 |
| 0.000 | 0.000 | 0.000 | 85.835 | 0.000 | -7.747 |
| 15.676 | 9.778 | 2.534 | 0.000 | 40.803 | 0.000 |
| -2.534 | -5.015 | 1.150 | -0.384 | 1.881 | 15.717 |
| 57.374 | 27.413 | 26.486 | -0.000 | 20.381 | -0.000 |
| -5.851 | 113.972 | 15.200 | -0.000 | 4.573 | -0.000 |
| 21.582 | 24.720 | 99.261 | -0.000 | 2.414 | 0.000 |
| -0.000 | -0.000 | -0.000 | 85.835 | -0.000 | -7.747 |
| 14.284 | 4.921 | 3.909 | -0.000 | 24.760 | 0.000 |
| 2.534 | 5.015 | -1.150 | -0.384 | -1.881 | 15.717 |
| -2525.468 | -72.721 | 832.498 | 0.000 | 1542.005 | -0.000 |
| -1026.130 | 493.503 | 338.329 | 0.000 | 469.149 | -0.000 |
| -2980.955 | -390.113 | 718.058 | 0.000 | 809.662 | -0.000 |
| -0.000 | 0.000 | 0.000 | 98.781 | 0.000 | -8.534 |
| -3041.346 | -436.879 | 610.063 | 0.000 | 838.522 | -0.000 |
| -3045.374 | -440.410 | 602.971 | 2.028 | 804.210 | 18.879 |
| 1322.551 | 665.151 | 11.436 | -0.000 | -471.568 | -0.000 |
| 1192.232 | 794.422 | 7.197 | 0.000 | -491.763 | -0.000 |
| 1116.532 | -257.358 | -88.618 | -0.000 | -469.347 | 0.000 |
| 0.000 | -0.000 | -0.000 | 98.781 | 0.000 | -8.534 |
| 1177.130 | 643.525 | -32.232 | -0.000 | -423.714 | -0.000 |
| 3045.374 | 440.410 | -602.971 | 2.028 | -804.210 | 18.879 |
| 21.067 | -38.219 | 43.641 | -0.000 | 74.680 | -0.000 |
| 93.750 | 344.494 | 88.461 | -0.000 | -5.096 | 0.000 |
| -52.211 | 98.962 | 208.108 | -0.000 | 38.084 | -0.000 |
| -105.982 | 32.333 | 26.986 | 57.376 | 46.031 | 0.785 |
| -114.888 | -58.928 | 1.459 | -0.000 | 122.087 | 0.000 |
| -105.848 | 32.292 | 26.952 | 1.129 | 45.973 | 61.052 |
| 387.007 | 91.138 | 106.315 | -0.000 | 32.457 | -0.000 |
| 144.888 | 145.554 | 41.855 | -0.000 | -38.868 | 0.000 |
| 199.285 | 109.877 | -266.423 | -0.000 | -31.634 | 0.000 |
| 105.982 | -32.333 | -26.986 | 57.376 | -46.031 | 0.785 |
| 307.881 | 47.379 | -53.523 | -0.000 | -44.552 | 0.000 |
| 105.848 | -32.292 | -26.952 | 1.129 | -45.973 | 61.052 |
| 122.103 | 15.184 | 39.400 | -0.000 | 30.614 | -0.000 |
| 33.410 | 70.831 | 53.413 | 0.000 | 13.307 | -0.000 |
| 28.086 | 46.905 | 110.208 | 0.000 | 12.773 | -0.000 |
| -12.847 | -4.966 | 0.878 | 51.831 | 5.454 | 0.570 |
| -8.181 | 0.724 | -1.767 | -0.000 | 70.120 | -0.000 |
| -17.379 | -2.037 | 5.029 | 2.556 | 9.996 | 14.625 |
| 154.255 | 23.251 | 38.263 | 0.000 | 19.625 | -0.000 |
| 61.306 | 206.589 | 69.647 | -0.000 | -0.325 | 0.000 |
| 49.259 | 45.440 | 158.005 | -0.000 | -8.235 | -0.000 |
| 12.847 | 4.966 | -0.878 | 51.831 | -5.454 | 0.570 |
| 18.361 | 12.445 | -3.191 | 0.000 | 57.737 | 0.000 |
| 17.379 | 2.037 | -5.029 | 2.556 | -9.996 | 14.625 |
| 76.892 | 74.398 | 74.398 | 0.000 | 0.000 | 0.000 |
| 74.398 | 76.892 | 74.398 | 0.000 | 0.000 | 0.000 |
| 74.398 | 74.398 | 76.892 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -1.767 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -1.767 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -1.767 |
| 76.892 | 74.398 | 74.398 | -0.000 | 0.000 | -0.000 |
| 74.398 | 76.892 | 74.398 | 0.000 | -0.000 | 0.000 |
| 74.398 | 74.398 | 76.892 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | -1.767 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -1.767 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | -1.767 |
| 76.904 | 74.392 | 74.392 | 0.000 | -0.000 | -0.000 |
| 74.392 | 76.904 | 74.392 | -0.000 | 0.000 | 0.000 |
| 74.392 | 74.392 | 76.904 | 0.000 | 0.000 | 0.000 |
| 0.007 | -0.004 | -0.004 | -1.763 | 0.000 | -0.000 |
| -0.004 | 0.007 | -0.004 | 0.000 | -1.763 | -0.000 |
| -0.004 | -0.004 | 0.007 | 0.000 | 0.000 | -1.763 |
| 76.893 | 74.398 | 74.398 | -0.000 | 0.000 | 0.000 |
| 74.398 | 76.893 | 74.398 | 0.000 | -0.000 | 0.000 |
| 74.398 | 74.398 | 76.893 | -0.000 | -0.000 | 0.000 |
| -0.007 | 0.004 | 0.004 | -1.763 | -0.000 | 0.000 |
| 0.004 | -0.007 | 0.004 | -0.000 | -1.763 | 0.000 |
| 0.004 | 0.004 | -0.007 | -0.000 | -0.000 | -1.763 |
| 78.137 | 73.776 | 73.776 | 0.000 | 0.000 | 0.000 |
| 73.776 | 78.137 | 73.776 | -0.000 | 0.000 | -0.000 |
| 73.776 | 73.776 | 78.137 | 0.000 | 0.000 | -0.000 |
| 0.733 | -0.367 | -0.367 | -1.531 | -0.000 | 0.000 |
| -0.367 | 0.733 | -0.367 | 0.000 | -1.531 | 0.000 |
| -0.367 | -0.367 | 0.733 | 0.000 | -0.000 | -1.531 |
| 77.047 | 74.320 | 74.320 | -0.000 | 0.000 | -0.000 |
| 74.320 | 77.047 | 74.320 | 0.000 | -0.000 | -0.000 |
| 74.320 | 74.320 | 77.047 | -0.000 | -0.000 | -0.000 |
| -0.733 | 0.367 | 0.367 | -1.531 | -0.000 | 0.000 |
| 0.367 | -0.733 | 0.367 | 0.000 | -1.531 | -0.000 |
| 0.367 | 0.367 | -0.733 | -0.000 | -0.000 | -1.531 |
| -268.354 | -268.573 | -268.573 | -0.000 | -0.000 | -0.000 |
| -268.573 | -268.354 | -268.573 | -0.000 | -0.000 | -0.000 |
| -268.573 | -268.573 | -268.354 | -0.000 | -0.000 | -0.000 |
| -290596.500 | -52909600.601 | -52909424.137 | -52909559.925 | -115577.539 | -115510.976 |
| -51323875.140 | 68807.653 | -51323662.059 | 51712.563 | -51323826.726 | 51730.827 |
| -53233939.711 | -53233889.384 | -12993.006 | -34394.921 | -34374.331 | -53233950.642 |
| -268.354 | -268.573 | -268.573 | 0.000 | 0.000 | 0.000 |
| -268.573 | -268.354 | -268.573 | 0.000 | 0.000 | 0.000 |
| -268.573 | -268.573 | -268.354 | 0.000 | 0.000 | -0.000 |
| 28587.204 | 47069910.935 | 47069987.812 | -47070366.037 | 6447.685 | -6440.531 |
| 45429166.902 | 19620.273 | 45429263.326 | 40869.287 | -45429539.114 | -40852.686 |
| 45311592.517 | 45311999.502 | 190326.958 | 98266.891 | -98323.329 | -45312179.240 |
| -268.353 | -268.573 | -268.573 | 0.000 | 0.000 | -0.000 |
| -268.573 | -268.353 | -268.573 | 0.000 | 0.000 | -0.000 |
| -268.573 | -268.573 | -268.353 | 0.000 | 0.000 | -0.000 |
| 2111.458 | -3979220.278 | -3979285.102 | -3979293.459 | 24.369 | 28.599 |
| -4097479.754 | -12566.921 | -4097421.292 | -434.932 | -4097534.530 | -445.193 |
| -4408246.393 | -4408261.133 | 517.087 | 1542.534 | 1542.780 | -4408266.267 |
| -268.354 | -268.574 | -268.574 | -0.000 | -0.000 | 0.000 |
| -268.574 | -268.354 | -268.574 | -0.000 | -0.000 | 0.000 |
| -268.574 | -268.574 | -268.354 | -0.000 | -0.000 | 0.000 |
| -39928.967 | 4447525.447 | 4447776.291 | -4448117.012 | -5324.529 | 5324.605 |
| 4542541.994 | -15436.549 | 4541010.773 | 36975.799 | -4542525.874 | -36973.972 |
| 5017274.998 | 5017189.061 | -2451.015 | 6555.867 | -6558.061 | -5017432.816 |
| -268.100 | -268.692 | -268.692 | -0.000 | -0.000 | -0.000 |
| -268.692 | -268.100 | -268.692 | -0.000 | -0.000 | -0.000 |
| -268.692 | -268.692 | -268.100 | -0.000 | -0.000 | -0.000 |
| -166.711 | -458344.683 | -458308.953 | -458368.859 | 253.268 | 252.445 |
| -454987.960 | 1644.843 | -454985.475 | 260.817 | -455015.026 | 259.914 |
| -462453.936 | -462453.328 | 283.337 | -1283.426 | -1283.680 | -462484.720 |
| -268.360 | -268.579 | -268.579 | -0.000 | -0.000 | 0.000 |
| -268.579 | -268.360 | -268.579 | -0.000 | -0.000 | 0.000 |
| -268.579 | -268.579 | -268.360 | -0.000 | -0.000 | 0.000 |
| 228.308 | 447411.601 | 447295.925 | -447455.280 | -365.315 | 364.332 |
| 409339.990 | -4215.247 | 409277.209 | -1970.033 | -409396.305 | 1971.206 |
| 467345.172 | 467332.580 | -942.507 | -2811.440 | 2812.927 | -467364.500 |
| 139233.436 | 94087.529 | 518750.003 | -266706.691 | 362188.283 | 37702.018 |
| 228663.897 | 186139.891 | 201869.440 | -20528.810 | 469374.200 | 26379.118 |
| -377332.757 | 31793.020 | 99605.079 | -682775.040 | 464534.716 | 425124.860 |
| -79266.842 | 227796.806 | -243583.395 | -759067.653 | 510615.402 | 466106.957 |
| 152623.588 | 92808.021 | 115291.265 | -201218.723 | 625343.958 | 123146.770 |
| 112898.166 | 249367.884 | 2116785.413 | -41654.263 | -371295.899 | -322.678 |
| -1066816.975 | -683004.212 | -2641908.926 | 518628.931 | 956683.367 | -403695.985 |
| -323103.380 | -128124.756 | -1254108.793 | 116251.756 | -93438.138 | -153621.462 |
| 311123.033 | -158712.030 | -342166.884 | 293818.080 | -426625.048 | -162510.197 |
| -264288.838 | -285091.553 | -1941013.359 | 611562.529 | 485608.060 | -501315.755 |
| -64233.817 | 47554.095 | 472266.722 | 238412.022 | -863423.672 | -150057.730 |
| -60836.085 | -237646.085 | 806033.993 | 20159.199 | -835188.142 | 101001.233 |
| 4724.659 | 3612.954 | -11196.877 | -33845.140 | 69220.134 | 29158.005 |
| 47370.751 | 27602.448 | 108519.090 | -55875.191 | 446.648 | 35154.302 |
| -17575.258 | -10316.477 | 38825.172 | -94985.047 | 34640.083 | 58180.883 |
| 5270.162 | 11155.678 | 52042.231 | -6714.991 | 34141.407 | -8675.208 |
| 39097.246 | 13911.088 | 56720.483 | -36310.407 | 20265.129 | 32618.862 |
| -47041.328 | -1815.834 | 24339.106 | -51073.964 | 46764.082 | 44858.117 |
| 90041.812 | -1440.634 | -37439.670 | 12072.744 | -70630.160 | -620.021 |
| 63926.399 | 27592.771 | 91454.270 | 67062.697 | -120651.942 | -34677.602 |
| 547.247 | 7503.065 | -32403.892 | 30867.109 | -61307.333 | -25293.900 |
| 23816.725 | -11146.586 | -68366.545 | 28467.606 | -18988.646 | -34800.265 |
| -33300.362 | -38856.642 | 37565.675 | 81794.464 | -52379.084 | -52437.016 |
| -27452.638 | -9085.516 | -38658.604 | 49350.441 | -39937.821 | -29374.418 |
| -5766.016 | -4837.365 | -17513.655 | -6431.750 | 15926.159 | 4744.941 |
| 1785.986 | 1109.611 | 5604.728 | -2589.262 | 4013.466 | 2178.407 |
| 1658.400 | 2078.977 | 1683.539 | -1536.593 | 4804.521 | 1041.980 |
| -13210.168 | -7988.260 | -24935.985 | -4108.072 | 19851.564 | 1823.658 |
| 4372.140 | 1442.557 | 6441.602 | -3842.654 | 1817.617 | 2372.333 |
| 5570.527 | 5669.069 | 21949.428 | -6241.498 | -5800.383 | 3173.676 |
| 120.362 | -493.039 | 8376.697 | 4567.792 | -8915.588 | -3454.757 |
| -6882.326 | -4125.876 | -14358.238 | 4341.460 | 3093.370 | -3582.144 |
| 674.490 | 1378.564 | 7541.364 | 4351.696 | -9469.431 | -3913.467 |
| -12569.184 | -12457.147 | -41403.225 | 3975.046 | 19826.827 | -2981.209 |
| 6255.414 | 3414.608 | -552.457 | -486.708 | -6079.665 | -117.690 |
| -6089.291 | -2584.396 | -19020.855 | -253.728 | 2424.871 | 401.021 |
| -19059552.792 | -56003902.494 | -10992911.250 | -25959.143 | 24063442.743 | 877427.346 |
| -16921317.058 | -51488682.578 | -7246919.953 | -1522090.790 | 22048890.860 | 2219482.532 |
| -16530608.679 | -53822179.359 | -5651901.154 | 704404.089 | 22257581.630 | 2036091.693 |
| -88.901 | -157.886 | -482.531 | 5927.161 | 104.004 | -966.032 |
| -1265.300 | -1158.258 | -3402.416 | 15.410 | 2858.158 | -26.689 |
| -16199145.388 | -51064887.029 | -3916767.991 | -394870.607 | 22117180.060 | 2762783.544 |
| 3070.871 | 3375.262 | 2195.880 | 7.789 | -332.484 | 8.458 |
| 3741.978 | 10155.657 | 7128.119 | 62.664 | -1191.590 | 39.584 |
| 3071.693 | 6633.862 | 19148.844 | 114.710 | -3294.567 | 29.569 |
| 16170693.115 | 46227129.641 | 8759774.609 | -1406988.062 | -21779196.205 | -532755.986 |
| 18514336.844 | 54899147.433 | 16059600.509 | 913116.960 | -24599529.514 | -2036458.028 |
| 43.988 | 105.715 | 228.422 | -957.382 | -54.599 | 3919.735 |
| -1614378.322 | -6133205.539 | -650178.025 | -220836.411 | 2331442.116 | 153325.097 |
| -1666615.293 | -6458810.906 | -1268469.443 | -40049.124 | 2443917.648 | 150271.983 |
| -1824053.190 | -4849920.865 | -935239.085 | -15356.842 | 2187383.322 | 175719.990 |
| -93.623 | -169.847 | -519.959 | 5918.663 | 109.148 | -964.577 |
| -1265.834 | -1160.965 | -3412.744 | 14.641 | 2858.465 | -26.602 |
| -1466601.080 | -5080647.485 | -558972.077 | -37617.195 | 2247972.867 | 240516.480 |
| 3070.892 | 3376.747 | 2206.814 | 8.519 | -333.635 | 8.379 |
| 3744.300 | 10162.684 | 7198.570 | 67.754 | -1199.802 | 39.043 |
| 3094.714 | 6689.503 | 19329.934 | 130.890 | -3320.301 | 27.878 |
| 1544515.300 | 4957106.545 | 749289.292 | -17325.933 | -2197882.747 | -113892.551 |
| 1798182.404 | 4724222.254 | 837641.369 | 30814.361 | -2154240.932 | -190026.416 |
| 46.167 | 111.360 | 246.593 | -955.819 | -57.033 | 3918.407 |
| -170585.886 | -534889.833 | -100211.360 | -272.105 | 229919.487 | 18804.933 |
| -151706.288 | -496206.671 | -50847.899 | -7256.663 | 218900.974 | 19648.418 |
| -181871.181 | -411722.339 | -68974.103 | -5770.964 | 203517.252 | 18502.902 |
| -151.969 | -316.615 | -979.480 | 5764.266 | 172.896 | -935.792 |
| -1271.335 | -1194.392 | -3544.095 | 4.592 | 2860.584 | -25.400 |
| 18.204 | 40.178 | 18.351 | -968.856 | -25.973 | 3919.250 |
| 3069.789 | 3393.963 | 2350.011 | 18.199 | -348.315 | 7.260 |
| 3721.018 | 10127.359 | 8099.821 | 143.474 | -1296.133 | 28.131 |
| 183691.705 | 479345.051 | 104899.818 | 8850.063 | -217546.010 | -19086.935 |
| 193941.646 | 382708.597 | 83618.175 | 8017.303 | -197819.249 | -21444.341 |
| 205461.788 | 502981.107 | 72267.603 | 24136.915 | -226882.052 | -27132.704 |
| 71.994 | 177.139 | 457.543 | -932.138 | -85.787 | 3890.241 |
| 2542.150 | -584.786 | -584.785 | 0.000 | 0.000 | -0.000 |
| -584.786 | 2542.150 | -584.785 | 0.001 | 0.000 | -0.000 |
| -584.785 | -584.785 | 2542.150 | 0.000 | 0.001 | -0.000 |
| -0.000 | 0.000 | 0.001 | 5830.234 | 0.001 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 5830.234 | -0.000 |
| 0.000 | 0.000 | -0.001 | 0.000 | 0.000 | 5895.794 |
| 2542.149 | -584.785 | -584.785 | -0.001 | -0.001 | 0.000 |
| -584.785 | 2542.149 | -584.785 | -0.001 | -0.001 | -0.000 |
| -584.785 | -584.785 | 2542.148 | -0.001 | -0.001 | 0.000 |
| 0.000 | -0.000 | -0.001 | 5830.233 | -0.001 | 0.000 |
| -0.000 | 0.000 | -0.001 | -0.000 | 5830.233 | 0.000 |
| -0.000 | -0.000 | 0.001 | -0.000 | -0.000 | 5895.794 |
| 2542.151 | -584.786 | -584.786 | 0.000 | 0.000 | -0.000 |
| -584.786 | 2542.151 | -584.786 | 0.000 | 0.000 | -0.000 |
| -584.786 | -584.786 | 2542.151 | 0.000 | 0.000 | 0.000 |
| -0.003 | 0.001 | 0.002 | 5830.233 | -0.000 | 0.000 |
| 0.001 | -0.003 | 0.002 | 0.000 | 5830.233 | 0.000 |
| 0.001 | 0.002 | -0.003 | 0.000 | 0.000 | 5830.233 |
| 2542.144 | -584.783 | -584.783 | -0.000 | -0.000 | 0.000 |
| -584.783 | 2542.144 | -584.783 | -0.000 | -0.000 | -0.000 |
| -584.783 | -584.783 | 2542.144 | -0.000 | -0.000 | 0.000 |
| 0.003 | -0.001 | -0.003 | 5830.233 | -0.000 | -0.000 |
| -0.001 | 0.003 | -0.002 | -0.000 | 5830.233 | 0.000 |
| -0.001 | -0.002 | 0.003 | -0.000 | -0.000 | 5830.233 |
| 2542.200 | -584.807 | -584.807 | 0.000 | 0.000 | -0.000 |
| -584.807 | 2542.200 | -584.807 | 0.000 | 0.000 | -0.000 |
| -584.807 | -584.807 | 2542.200 | 0.000 | 0.000 | -0.000 |
| -0.029 | 0.014 | 0.024 | 5830.233 | 0.000 | -0.000 |
| 0.014 | -0.029 | 0.024 | 0.000 | 5830.233 | 0.000 |
| 0.014 | 0.024 | -0.029 | 0.000 | 0.000 | 5830.233 |
| 2542.098 | -584.764 | -584.764 | -0.000 | -0.000 | 0.000 |
| -584.764 | 2542.098 | -584.764 | -0.000 | -0.000 | -0.000 |
| -584.764 | -584.764 | 2542.098 | -0.000 | -0.000 | 0.000 |
| 0.029 | -0.014 | -0.024 | 5830.233 | -0.000 | -0.000 |
| -0.014 | 0.029 | -0.024 | -0.000 | 5830.233 | 0.000 |
| -0.014 | -0.024 | 0.029 | -0.000 | -0.000 | 5830.233 |
| 93673.500 | 82615.692 | 82615.693 | 0.000 | -0.000 | 0.000 |
| 82615.693 | 93673.499 | 82615.693 | -0.000 | -0.000 | 0.000 |
| 82615.693 | 82615.692 | 93673.500 | -0.000 | -0.000 | 0.000 |
| 0.056 | -0.029 | -0.029 | 2104.980 | -0.000 | 0.000 |
| -0.029 | 0.056 | -0.030 | 0.000 | 2104.982 | 0.000 |
| -0.029 | -0.030 | 0.057 | 0.000 | 0.000 | 2104.980 |
| 93671.834 | 82615.863 | 82615.863 | -0.000 | -0.000 | -0.000 |
| 82615.863 | 93671.834 | 82615.863 | -0.000 | 0.000 | 0.000 |
| 82615.863 | 82615.863 | 93671.833 | 0.000 | 0.000 | -0.000 |
| -0.056 | 0.029 | 0.030 | 2104.981 | 0.000 | -0.000 |
| 0.029 | -0.056 | 0.029 | -0.000 | 2104.981 | -0.000 |
| 0.029 | 0.030 | -0.057 | -0.000 | 0.000 | 2104.981 |
| 93676.765 | 82617.047 | 82617.047 | 0.000 | 0.000 | -0.000 |
| 82617.047 | 93676.765 | 82617.047 | -0.000 | -0.000 | 0.000 |
| 82617.047 | 82617.047 | 93676.765 | -0.000 | -0.000 | 0.000 |
| 0.566 | -0.289 | -0.292 | 2104.299 | -0.000 | 0.000 |
| -0.289 | 0.566 | -0.292 | 0.000 | 2104.300 | 0.000 |
| -0.289 | -0.292 | 0.566 | 0.000 | -0.000 | 2104.299 |
| 93670.223 | 82613.682 | 82613.682 | 0.000 | -0.000 | -0.000 |
| 82613.682 | 93670.223 | 82613.682 | -0.000 | -0.000 | -0.000 |
| 82613.682 | 82613.682 | 93670.223 | 0.000 | -0.000 | 0.000 |
| -0.566 | 0.289 | 0.292 | 2104.299 | 0.000 | -0.000 |
| 0.289 | -0.566 | 0.292 | -0.000 | 2104.300 | -0.000 |
| 0.289 | 0.292 | -0.566 | -0.000 | 0.000 | 2104.299 |
| 93697.643 | 82636.467 | 82636.467 | 0.000 | 0.000 | 0.000 |
| 82636.467 | 93697.643 | 82636.467 | 0.000 | -0.000 | -0.000 |
| 82636.467 | 82636.467 | 93697.643 | -0.000 | -0.000 | 0.000 |
| -0.109 | -0.018 | -0.020 | 2104.876 | -0.000 | 0.000 |
| -0.018 | -0.109 | -0.020 | 0.000 | 2104.876 | 0.000 |
| -0.018 | -0.020 | -0.109 | 0.000 | 0.000 | 2104.876 |
| 93647.870 | 82594.999 | 82594.999 | 0.000 | -0.000 | -0.000 |
| 82594.999 | 93647.870 | 82594.999 | -0.000 | -0.000 | -0.000 |
| 82594.999 | 82594.999 | 93647.870 | -0.000 | -0.000 | -0.000 |
| 0.109 | 0.018 | 0.020 | 2104.876 | 0.000 | -0.000 |
| 0.018 | 0.109 | 0.020 | -0.000 | 2104.876 | -0.000 |
| 0.018 | 0.020 | 0.109 | -0.000 | -0.000 | 2104.876 |
| 593.262 | 336.528 | 336.528 | 0.000 | -0.000 | -0.000 |
| 336.528 | 593.262 | 336.528 | -0.000 | 0.000 | -0.000 |
| 336.528 | 336.528 | 593.262 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -355.305 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -355.305 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -355.305 |
| 593.262 | 336.528 | 336.528 | -0.000 | -0.000 | 0.000 |
| 336.528 | 593.262 | 336.528 | -0.000 | -0.000 | 0.000 |
| 336.528 | 336.528 | 593.262 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -355.305 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -355.305 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 0.000 | -355.305 |
| 593.262 | 336.529 | 336.529 | 0.000 | 0.000 | -0.000 |
| 336.529 | 593.262 | 336.529 | 0.000 | 0.000 | -0.000 |
| 336.529 | 336.529 | 593.262 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 74.831 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 74.831 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 74.831 |
| 593.261 | 336.528 | 336.528 | -0.000 | -0.000 | 0.000 |
| 336.528 | 593.261 | 336.528 | -0.000 | -0.000 | 0.000 |
| 336.528 | 336.528 | 593.261 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 74.831 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 74.831 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 74.831 |
| 593.263 | 336.530 | 336.530 | -0.000 | 0.000 | -0.000 |
| 336.530 | 593.263 | 336.530 | 0.000 | -0.000 | -0.000 |
| 336.530 | 336.530 | 593.263 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.001 | -0.000 | -355.305 | -0.000 | -0.000 |
| 0.001 | 0.000 | -0.000 | -0.000 | -355.305 | -0.000 |
| 0.001 | -0.000 | 0.000 | -0.000 | 0.000 | -355.305 |
| 593.260 | 336.527 | 336.527 | -0.000 | -0.000 | 0.000 |
| 336.527 | 593.260 | 336.527 | -0.000 | -0.000 | 0.000 |
| 336.527 | 336.527 | 593.260 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.001 | 0.000 | -355.305 | 0.000 | 0.000 |
| -0.001 | -0.000 | 0.000 | 0.000 | -355.305 | 0.000 |
| -0.001 | 0.000 | -0.000 | 0.000 | 0.000 | -355.305 |
| 711.547 | 526.906 | 526.906 | 0.000 | -0.000 | -0.000 |
| 526.906 | 711.547 | 526.906 | 0.000 | 0.000 | 0.000 |
| 526.906 | 526.906 | 711.547 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 172.568 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 172.568 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 172.568 |
| 711.547 | 526.906 | 526.906 | -0.000 | -0.000 | 0.000 |
| 526.906 | 711.547 | 526.906 | -0.000 | -0.000 | 0.000 |
| 526.906 | 526.906 | 711.547 | -0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 172.568 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 172.568 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 172.568 |
| 711.547 | 526.906 | 526.906 | 0.000 | 0.000 | -0.000 |
| 526.906 | 711.547 | 526.906 | -0.000 | -0.000 | 0.000 |
| 526.906 | 526.906 | 711.547 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 172.568 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 172.568 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 172.568 |
| 711.546 | 526.906 | 526.906 | 0.000 | -0.000 | 0.000 |
| 526.906 | 711.546 | 526.906 | 0.000 | -0.000 | 0.000 |
| 526.906 | 526.906 | 711.546 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 172.568 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 172.568 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 172.568 |
| 711.548 | 526.908 | 526.908 | 0.000 | 0.000 | -0.000 |
| 526.908 | 711.548 | 526.908 | -0.000 | 0.000 | -0.000 |
| 526.908 | 526.908 | 711.548 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 172.568 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 172.568 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 172.568 |
| 711.545 | 526.905 | 526.905 | -0.000 | -0.000 | 0.000 |
| 526.905 | 711.545 | 526.905 | 0.000 | -0.000 | -0.000 |
| 526.905 | 526.905 | 711.545 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 172.568 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 172.568 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 172.568 |
| -36565524.562 | 825443.594 | 793189.978 | 187996.316 | 1128647.866 | 794305.936 |
| -35081318.596 | 920020.470 | 883278.001 | 26257.792 | 1104405.306 | 501511.178 |
| -35172869.784 | 585892.845 | 863659.463 | 330846.253 | 935150.472 | 598369.280 |
| -34515742.680 | 1630216.005 | 825643.957 | -236796.139 | 998181.390 | 415519.178 |
| -33973156.311 | 966129.136 | 646138.383 | -78094.214 | 1232343.642 | 432940.002 |
| -35050971.557 | 556295.736 | 709309.953 | 94391.827 | 1241497.197 | 604826.306 |
| 35400023.925 | -1483576.929 | -1008645.996 | 167727.945 | -1194486.437 | -477644.557 |
| 35444741.646 | -817093.808 | -923336.778 | -81767.349 | -1177830.079 | -501300.496 |
| 34756360.428 | -465464.506 | -648859.456 | -40363.325 | -1227367.913 | -342151.304 |
| 35420377.815 | -1047038.180 | -887303.045 | 6341.176 | -1070133.935 | -667760.221 |
| 38876734.087 | -50590.169 | -766654.417 | -166271.790 | -1115862.964 | -740924.466 |
| 36567305.013 | -330930.947 | -442758.050 | -66009.103 | -1917054.964 | -846576.538 |
| -4001995.889 | 166637.831 | 60557.527 | -8087.043 | 116689.617 | 45987.382 |
| -3594537.355 | 89320.323 | 95005.028 | 9381.393 | 118817.249 | 52443.942 |
| -3448900.664 | 72144.840 | 84076.556 | -9791.193 | 113600.374 | 52193.952 |
| -3645392.113 | 71143.711 | 86219.947 | -4222.922 | 139363.325 | 68630.849 |
| -3563055.485 | -21558.442 | 101586.033 | 105201.949 | 43523.433 | -16034.879 |
| -2719659.544 | 126600.433 | 80440.710 | -1130.855 | 111448.811 | 22695.465 |
| 3352838.286 | -93197.308 | -76277.696 | -2311.740 | -108428.138 | -51132.365 |
| 3601810.352 | -130636.368 | -103032.871 | 5693.016 | -118232.273 | -44769.720 |
| 3684232.168 | -99789.867 | -100897.193 | -6189.033 | -120259.819 | -50621.368 |
| 3533923.929 | -107875.230 | -95031.706 | -1620.719 | -108060.960 | -41034.347 |
| 3561051.989 | -207325.243 | -117652.255 | 28315.252 | -90264.951 | -60925.371 |
| 3522353.232 | -163281.053 | -99054.163 | 29090.368 | -102011.518 | -86346.190 |
| -343576.095 | 17699.357 | 10931.098 | -2334.625 | 10174.347 | 5035.276 |
| -356472.943 | -9120.400 | 328.282 | 7944.662 | 12230.552 | 7569.035 |
| -326162.339 | 3354.041 | 8413.117 | 281.749 | 10792.548 | 4801.146 |
| -383005.384 | 8089.265 | 9561.386 | -517.923 | 14646.933 | 2051.278 |
| -349063.633 | 7060.412 | 7343.986 | 130.757 | 11432.693 | 4345.049 |
| -345438.500 | 14079.427 | 8674.511 | -1800.732 | 11567.577 | 4193.557 |
| 381753.017 | -12354.936 | -6816.633 | 449.323 | -15144.420 | 729.475 |
| 333751.586 | -10907.655 | -7564.006 | 306.953 | -13295.964 | -271.425 |
| 338221.772 | -32786.465 | -9914.497 | 7698.528 | -6146.242 | -15545.491 |
| 361359.072 | -10536.506 | -10296.942 | 267.572 | -12943.297 | -4030.210 |
| 342810.160 | -17871.274 | -8431.023 | 2729.771 | -11878.828 | -4328.639 |
| 351886.745 | -13461.676 | -7709.862 | 118.745 | -11496.024 | -4612.936 |
| 283.976 | 47.731 | 93.114 | -0.000 | 0.000 | 0.000 |
| 47.731 | 283.976 | 93.114 | -0.000 | 0.000 | 0.000 |
| 93.114 | 93.114 | 235.575 | -0.000 | 0.000 | 0.000 |
| -7.032 | -7.032 | -26.414 | 60.952 | -0.000 | 0.000 |
| -7.032 | -7.032 | -26.414 | -0.000 | 60.952 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 75.743 |
| 153.090 | 43.266 | 59.180 | 0.000 | 0.000 | -0.000 |
| 43.266 | 153.091 | 59.180 | 0.000 | 0.000 | -0.000 |
| 11.965 | 11.965 | 141.697 | 0.000 | 0.000 | -0.000 |
| 7.032 | 7.032 | 26.414 | 60.952 | 0.000 | -0.000 |
| 7.032 | 7.032 | 26.414 | 0.000 | 60.952 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 75.743 |
| 283.976 | 47.731 | 93.114 | -0.000 | 0.000 | -0.000 |
| 47.731 | 283.976 | 93.114 | 0.000 | 0.000 | -0.000 |
| 21.187 | 21.187 | 130.057 | -0.000 | -0.000 | 0.000 |
| -0.373 | -0.373 | -10.564 | 33.838 | -0.000 | 0.000 |
| -0.373 | -0.373 | -10.564 | -0.000 | 33.838 | 0.000 |
| -1.242 | -1.242 | -4.666 | -0.000 | -0.000 | 49.039 |
| 86.412 | -22.436 | 7.486 | 0.000 | 0.000 | -0.000 |
| -22.436 | 86.412 | 7.486 | 0.000 | 0.000 | -0.000 |
| 93.114 | 93.114 | 235.575 | -0.000 | -0.000 | -0.000 |
| 0.373 | 0.373 | 10.564 | 33.838 | 0.000 | -0.000 |
| 0.373 | 0.373 | 10.564 | 0.000 | 33.838 | -0.000 |
| 1.242 | 1.242 | 4.666 | 0.000 | 0.000 | 49.039 |
| 65.296 | 6.707 | 13.800 | -0.000 | -0.000 | 0.000 |
| 6.707 | 65.296 | 13.800 | -0.000 | -0.000 | 0.000 |
| 93.114 | 93.114 | 235.576 | -0.000 | 0.000 | 0.000 |
| 0.115 | 0.115 | -1.211 | 15.434 | -0.000 | 0.000 |
| 0.115 | 0.115 | -1.211 | -0.000 | 15.434 | 0.000 |
| -0.124 | -0.124 | -0.465 | -0.000 | -0.000 | 49.160 |
| 65.019 | 6.601 | 15.642 | 0.000 | 0.000 | -0.000 |
| 6.601 | 65.019 | 15.642 | 0.000 | 0.000 | -0.000 |
| 10.333 | 10.333 | 28.049 | -0.000 | -0.000 | -0.000 |
| -0.115 | -0.115 | 1.211 | 15.434 | 0.000 | -0.000 |
| -0.115 | -0.115 | 1.211 | 0.000 | 15.434 | -0.000 |
| 0.124 | 0.124 | 0.465 | 0.000 | 0.000 | 49.160 |
| -1477.069 | -1524.407 | -619.324 | 0.104 | 0.143 | -0.007 |
| -1524.348 | -1477.127 | -619.324 | 0.104 | 0.143 | -0.007 |
| -628.567 | -628.564 | 2645.396 | -0.001 | 0.001 | -0.001 |
| -0.000 | -0.000 | -0.000 | 113.809 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 113.809 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 70.839 |
| 776.110 | 623.101 | -2533.667 | 0.004 | 0.002 | 0.001 |
| 623.105 | 776.105 | -2533.669 | 0.004 | 0.002 | 0.001 |
| 87.407 | 87.407 | 237.728 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 113.809 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 113.809 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 70.839 |
| -72.800 | -70.951 | 432.028 | 0.007 | 0.010 | -0.001 |
| -70.947 | -72.803 | 432.028 | 0.007 | 0.010 | -0.001 |
| -68.217 | -68.221 | 516.648 | 0.007 | 0.010 | -0.001 |
| -85.106 | -85.108 | 402.887 | 50.496 | 0.005 | -0.000 |
| -85.125 | -85.127 | 402.722 | 0.003 | 50.516 | -0.000 |
| -65.954 | -65.954 | 253.658 | -0.000 | -0.000 | 36.595 |
| 80.759 | 77.105 | -433.193 | -0.006 | -0.009 | 0.000 |
| 77.102 | 80.762 | -433.192 | -0.006 | -0.009 | 0.000 |
| 174.397 | 174.404 | 193.567 | -0.012 | -0.016 | 0.001 |
| 85.107 | 85.109 | -402.887 | 50.488 | -0.005 | 0.000 |
| 85.107 | 85.109 | -402.887 | -0.003 | 50.486 | 0.000 |
| 65.954 | 65.953 | -253.659 | 0.000 | 0.000 | 36.595 |
| 16.730 | -19.070 | 56.058 | 0.000 | 0.001 | -0.000 |
| -19.070 | 16.730 | 56.058 | 0.000 | 0.001 | -0.000 |
| 87.407 | 87.407 | 237.726 | 0.000 | 0.000 | -0.000 |
| -1.953 | -1.953 | 42.214 | 26.958 | 0.001 | -0.000 |
| -1.953 | -1.953 | 42.214 | 0.000 | 26.958 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 70.839 |
| 18.050 | 8.564 | -18.448 | -0.001 | -0.001 | 0.000 |
| 1.209 | 21.688 | -24.848 | -0.000 | -0.001 | 0.000 |
| 22.970 | 22.971 | 50.100 | -0.001 | -0.001 | 0.000 |
| 1.953 | 1.953 | -42.214 | 26.957 | -0.001 | 0.000 |
| 1.953 | 1.953 | -42.214 | -0.000 | 26.956 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 70.839 |
| 1325315.259 | 5006473.247 | 370368.015 | 91658.023 | -2681885.584 | -23354.374 |
| 838270.334 | 4610709.699 | 511310.248 | -100240.595 | -2487155.673 | 10306.734 |
| 1276566.345 | 4415971.654 | 159214.984 | 238185.191 | -2433274.063 | 19144.314 |
| 1167190.416 | 4603131.190 | 570040.151 | 7839.293 | -2399937.960 | 6743.906 |
| 1182396.252 | 4623792.196 | 369712.261 | 28639.944 | -2574744.768 | 25068.208 |
| 1283073.101 | 4825786.508 | -96203.755 | 47748.627 | -2911677.174 | -35431.385 |
| -1254059.749 | -4676388.877 | -1076375.477 | -17065.339 | 2302730.162 | -50175.308 |
| -1274574.184 | -4778862.423 | -443370.179 | -3940.488 | 2520215.668 | -19574.047 |
| -1039091.334 | -4843305.801 | -459781.986 | -109586.834 | 2938336.002 | -44364.091 |
| -1534298.223 | -4991234.259 | -237763.003 | 7845.530 | 2729550.645 | -100324.566 |
| -349968.960 | -3643964.774 | 528350.245 | -15201.530 | 2718404.122 | -38996.454 |
| -1106185.434 | -4561287.975 | -347752.783 | 47531.731 | 2720948.091 | 353.061 |
| 107132.442 | 464749.363 | 51512.855 | 7970.312 | -248626.201 | -2823.327 |
| 137545.063 | 488201.080 | 51515.444 | 6468.344 | -258077.363 | -1143.836 |
| 146195.619 | 488385.819 | 71651.840 | 24693.007 | -242953.830 | 18094.986 |
| 120882.670 | 482916.662 | 73897.227 | 801.241 | -235580.637 | -1263.232 |
| 209015.192 | 542362.938 | 148503.893 | 5523.916 | -236282.692 | 15032.637 |
| 127580.081 | 473250.630 | 39094.545 | 9856.605 | -231545.067 | 8782.557 |
| -126027.505 | -474987.286 | -64986.193 | -2948.287 | 247600.460 | -433.585 |
| -106916.561 | -465788.022 | -33774.942 | 747.495 | 266853.533 | -4990.164 |
| -153876.278 | -502065.785 | -50662.459 | -4789.463 | 270491.395 | -4434.844 |
| -133690.950 | -479767.642 | -51110.369 | 5157.538 | 252729.613 | -7278.408 |
| -114091.538 | -468488.081 | -11450.824 | -5407.037 | 273022.534 | -6370.597 |
| -110598.074 | -466765.036 | -44834.182 | -2056.012 | 257846.318 | -4753.334 |
| 10844.473 | 46662.644 | 5798.147 | -628.006 | -23924.857 | -196.270 |
| 21266.245 | 54064.906 | 10100.583 | 501.894 | -26013.799 | 1653.944 |
| 9261.701 | 43348.250 | 4778.732 | -282.152 | -23454.326 | -38.168 |
| 11460.991 | 48719.048 | 2069.219 | -767.944 | -26968.939 | 909.742 |
| 13829.045 | 50620.509 | 8371.511 | -1188.349 | -25133.454 | 108.450 |
| 6578.059 | 38651.549 | 10134.078 | -169.098 | -16680.976 | 348.212 |
| -13499.374 | -48613.623 | -4034.943 | -772.193 | 26056.129 | -562.593 |
| -14870.198 | -49727.863 | -7491.152 | -176.004 | 25002.186 | -229.403 |
| -22504.107 | -53519.754 | -14236.666 | -866.651 | 22651.387 | -2278.630 |
| -10741.866 | -45175.262 | -4430.584 | 388.626 | 24790.055 | 223.412 |
| -18299.257 | -51420.540 | -3356.872 | -1144.249 | 26128.519 | -1211.414 |
| -11556.155 | -48049.530 | -3355.965 | -422.045 | 26886.744 | -405.605 |
| 223.847 | 127.662 | 127.662 | 0.000 | 0.000 | 0.000 |
| 127.662 | 223.847 | 127.662 | 0.000 | 0.000 | 0.000 |
| 127.662 | 127.662 | 223.847 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 11.534 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 11.534 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 11.534 |
| 223.847 | 127.662 | 127.662 | -0.000 | -0.000 | -0.000 |
| 127.662 | 223.847 | 127.662 | -0.000 | -0.000 | -0.000 |
| 127.662 | 127.662 | 223.847 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 11.534 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 11.534 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 11.534 |
| 223.847 | 127.662 | 127.662 | 0.000 | 0.000 | 0.000 |
| 127.662 | 223.847 | 127.662 | 0.000 | 0.000 | 0.000 |
| 127.662 | 127.662 | 223.847 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 11.534 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 11.534 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 11.534 |
| 223.846 | 127.662 | 127.662 | -0.000 | -0.000 | -0.000 |
| 127.662 | 223.846 | 127.662 | -0.000 | -0.000 | -0.000 |
| 127.662 | 127.662 | 223.846 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 11.534 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 11.534 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 11.534 |
| 223.850 | 127.663 | 127.663 | 0.000 | 0.000 | 0.000 |
| 127.663 | 223.850 | 127.663 | 0.000 | 0.000 | 0.000 |
| 127.663 | 127.663 | 223.850 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.001 | 0.001 | 11.534 | 0.000 | -0.000 |
| 0.001 | -0.000 | 0.001 | 0.000 | 11.534 | 0.000 |
| 0.001 | 0.001 | -0.000 | 0.000 | 0.000 | 11.534 |
| 223.844 | 127.661 | 127.661 | -0.000 | -0.000 | -0.000 |
| 127.661 | 223.844 | 127.661 | -0.000 | -0.000 | -0.000 |
| 127.661 | 127.661 | 223.844 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.001 | -0.001 | 11.534 | -0.000 | -0.000 |
| -0.001 | 0.000 | -0.001 | -0.000 | 11.534 | -0.000 |
| -0.001 | -0.001 | 0.000 | -0.000 | -0.000 | 11.534 |
| -896711791.093 | -885967983.634 | -891371906.266 | -125963.584 | 593462.187 | 470366.407 |
| -816411925.360 | -879621226.160 | -812084516.403 | 253188.406 | 428665.669 | -237053.595 |
| -881054032.024 | -895810787.241 | -862115931.196 | 377241.152 | -61198.222 | 528563.626 |
| -0.000 | -0.000 | -0.000 | -0.001 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.001 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.000 | -0.001 |
| -1410.399 | 23.915 | 23.915 | 0.000 | -0.000 | -0.000 |
| 23.915 | -1410.399 | 23.915 | -0.000 | -0.000 | -0.000 |
| 23.915 | 23.915 | -1410.399 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.001 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.001 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -0.000 | -0.001 |
| -1410.387 | 23.915 | 23.915 | 0.000 | -0.000 | 0.000 |
| 23.915 | -1410.387 | 23.915 | -0.000 | -0.000 | -0.000 |
| 23.915 | 23.915 | -1410.387 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.001 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.001 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | -0.000 | -0.001 |
| 88934675.685 | 87268287.431 | 89385064.415 | -77687.219 | 27301.791 | 179698.689 |
| 89204213.925 | 81803954.638 | 89357680.796 | -19228.055 | -9466.434 | -36925.468 |
| 86906363.121 | 88448586.687 | 89653181.641 | -26849.535 | 37196.460 | 5363.756 |
| 0.000 | 0.000 | -0.000 | -0.001 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.001 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.001 |
| -1410.292 | 23.915 | 23.915 | -0.000 | -0.000 | 0.000 |
| 23.915 | -1410.292 | 23.915 | -0.000 | 0.000 | -0.000 |
| 23.915 | 23.915 | -1410.292 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.001 | -0.001 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.001 | -0.000 | -0.001 | -0.000 |
| -0.000 | 0.001 | -0.000 | -0.000 | 0.000 | -0.001 |
| -1410.504 | 23.915 | 23.915 | -0.000 | 0.000 | 0.000 |
| 23.915 | -1410.504 | 23.915 | 0.000 | -0.000 | 0.000 |
| 23.915 | 23.915 | -1410.504 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.001 | -0.001 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.001 | 0.000 | -0.001 | 0.000 |
| 0.000 | -0.001 | 0.000 | 0.000 | -0.000 | -0.001 |
| 158.977 | 148.558 | 148.558 | -0.000 | -0.000 | 0.000 |
| 148.558 | 158.977 | 148.558 | -0.000 | -0.000 | -0.000 |
| 148.558 | 148.558 | 158.977 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 21.488 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 21.488 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 21.488 |
| 158.977 | 148.558 | 148.558 | 0.000 | 0.000 | 0.000 |
| 148.558 | 158.977 | 148.558 | 0.000 | 0.000 | 0.000 |
| 148.558 | 148.558 | 158.977 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 21.488 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 21.488 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 21.488 |
| 158.977 | 148.558 | 148.558 | -0.000 | -0.000 | -0.000 |
| 148.558 | 158.977 | 148.558 | 0.000 | -0.000 | -0.000 |
| 148.558 | 148.558 | 158.977 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 21.488 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 21.488 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 21.488 |
| 158.976 | 148.558 | 148.558 | 0.000 | 0.000 | 0.000 |
| 148.558 | 158.976 | 148.558 | 0.000 | 0.000 | -0.000 |
| 148.558 | 148.558 | 158.976 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 21.488 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 21.488 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 21.488 |
| 158.977 | 148.558 | 148.558 | -0.000 | -0.000 | -0.000 |
| 148.558 | 158.977 | 148.558 | -0.000 | -0.000 | 0.000 |
| 148.558 | 148.558 | 158.977 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 21.488 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 21.488 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 21.488 |
| 158.976 | 148.557 | 148.557 | 0.000 | 0.000 | 0.000 |
| 148.557 | 158.976 | 148.557 | 0.000 | 0.000 | 0.000 |
| 148.557 | 148.557 | 158.976 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 21.488 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 21.488 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 21.488 |
| -438.727 | -561.678 | -561.678 | 0.000 | -0.000 | -0.000 |
| -561.678 | -438.727 | -561.678 | -0.000 | 0.000 | -0.000 |
| -561.678 | -561.678 | -438.727 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -604.971 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -604.971 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -604.971 |
| -438.727 | -561.678 | -561.678 | 0.000 | 0.000 | -0.000 |
| -561.678 | -438.727 | -561.678 | 0.000 | 0.000 | -0.000 |
| -561.678 | -561.678 | -438.727 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -604.971 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -604.971 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -604.971 |
| -438.726 | -561.677 | -561.677 | 0.000 | 0.000 | -0.000 |
| -561.677 | -438.726 | -561.677 | 0.000 | 0.000 | -0.000 |
| -561.677 | -561.677 | -438.726 | -0.000 | -0.000 | 0.000 |
| 0.001 | 0.002 | 0.001 | -604.971 | -0.000 | -0.000 |
| 0.002 | 0.001 | 0.001 | -0.000 | -604.971 | 0.000 |
| 0.002 | 0.001 | 0.001 | -0.000 | -0.000 | -604.971 |
| -438.728 | -561.680 | -561.680 | 0.000 | 0.000 | 0.000 |
| -561.680 | -438.728 | -561.680 | 0.000 | 0.000 | 0.000 |
| -561.680 | -561.680 | -438.728 | 0.000 | 0.000 | -0.000 |
| -0.001 | -0.002 | -0.001 | -604.971 | 0.000 | -0.000 |
| -0.002 | -0.001 | -0.001 | 0.000 | -604.971 | 0.000 |
| -0.002 | -0.001 | -0.001 | 0.000 | -0.000 | -604.971 |
| -438.721 | -561.664 | -561.664 | 0.000 | 0.000 | -0.000 |
| -561.664 | -438.721 | -561.664 | 0.000 | 0.000 | 0.000 |
| -561.664 | -561.664 | -438.721 | 0.000 | 0.000 | -0.000 |
| 0.015 | 0.016 | 0.015 | -604.971 | 0.000 | 0.000 |
| 0.016 | 0.015 | 0.015 | 0.000 | -604.971 | 0.000 |
| 0.016 | 0.015 | 0.015 | 0.000 | -0.000 | -604.971 |
| -438.733 | -561.692 | -561.692 | -0.000 | 0.000 | -0.000 |
| -561.692 | -438.733 | -561.692 | 0.000 | -0.000 | -0.000 |
| -561.692 | -561.692 | -438.733 | -0.000 | -0.000 | -0.000 |
| -0.015 | -0.016 | -0.015 | -604.971 | -0.000 | -0.000 |
| -0.016 | -0.015 | -0.015 | -0.000 | -604.971 | 0.000 |
| -0.016 | -0.015 | -0.015 | -0.000 | 0.000 | -604.971 |
| 1004.987 | -370.723 | -370.723 | -0.000 | -0.000 | -0.000 |
| -370.723 | 1004.987 | -370.723 | -0.000 | -0.000 | -0.000 |
| -370.723 | -370.723 | 1004.987 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 382.602 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 382.602 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 382.602 |
| 1004.987 | -370.723 | -370.723 | 0.000 | 0.000 | 0.000 |
| -370.723 | 1004.987 | -370.723 | 0.000 | 0.000 | 0.000 |
| -370.723 | -370.723 | 1004.987 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 382.602 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 382.602 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 382.602 |
| 1004.987 | -370.723 | -370.723 | -0.000 | -0.000 | -0.000 |
| -370.723 | 1004.987 | -370.723 | -0.000 | -0.000 | -0.000 |
| -370.723 | -370.723 | 1004.987 | -0.000 | -0.000 | -0.000 |
| 0.005 | -0.002 | -0.002 | -2062.694 | -0.000 | -0.000 |
| -0.002 | 0.005 | -0.002 | -0.000 | -2062.694 | -0.000 |
| -0.002 | -0.002 | 0.005 | -0.000 | -0.000 | -2062.694 |
| 1004.986 | -370.723 | -370.723 | 0.000 | 0.000 | 0.000 |
| -370.723 | 1004.986 | -370.723 | 0.000 | 0.000 | 0.000 |
| -370.723 | -370.723 | 1004.986 | 0.000 | 0.000 | 0.000 |
| -0.005 | 0.002 | 0.002 | -2062.694 | 0.000 | 0.000 |
| 0.002 | -0.005 | 0.002 | 0.000 | -2062.694 | 0.000 |
| 0.002 | 0.002 | -0.005 | 0.000 | 0.000 | -2062.694 |
| 1004.991 | -370.723 | -370.723 | -0.000 | -0.000 | -0.000 |
| -370.723 | 1004.991 | -370.723 | -0.000 | -0.000 | -0.000 |
| -370.723 | -370.723 | 1004.991 | -0.000 | -0.000 | -0.000 |
| 0.047 | -0.017 | -0.022 | -2062.693 | -0.000 | -0.000 |
| -0.017 | 0.047 | -0.022 | -0.000 | -2062.693 | -0.000 |
| -0.017 | -0.022 | 0.047 | -0.000 | -0.000 | -2062.693 |
| 1004.982 | -370.722 | -370.722 | 0.000 | 0.000 | 0.000 |
| -370.722 | 1004.982 | -370.722 | 0.000 | 0.000 | 0.000 |
| -370.722 | -370.722 | 1004.982 | 0.000 | 0.000 | 0.000 |
| -0.047 | 0.017 | 0.022 | -2062.693 | 0.000 | 0.000 |
| 0.017 | -0.047 | 0.022 | 0.000 | -2062.693 | 0.000 |
| 0.017 | 0.022 | -0.047 | 0.000 | 0.000 | -2062.693 |
| -194.071 | -172.856 | -172.856 | 0.000 | 0.000 | 0.000 |
| -172.856 | -194.071 | -172.856 | 0.000 | 0.000 | 0.000 |
| -172.856 | -172.856 | -194.071 | 0.000 | 0.000 | 0.000 |
| -710686038.293 | -664887890.724 | -660743481.244 | -10138555.005 | 54955665.981 | 47986637.598 |
| -701775434.311 | -735252927.972 | -700041182.038 | 46593763.825 | -16552187.581 | 44254396.903 |
| -661705857.244 | -660162253.882 | -711788861.626 | 53561534.218 | 51178238.285 | -10484108.694 |
| -194.071 | -172.856 | -172.856 | -0.000 | -0.000 | -0.000 |
| -172.856 | -194.071 | -172.856 | -0.000 | -0.000 | -0.000 |
| -172.856 | -172.856 | -194.071 | -0.000 | -0.000 | -0.000 |
| 713155472.994 | 736867487.640 | 740072816.577 | -70590701.506 | 7143646.146 | 5964266.253 |
| 718503699.598 | 695757334.100 | 723821630.934 | 4012809.486 | -71577023.898 | 5641999.507 |
| 751863107.410 | 750963138.959 | 729392162.781 | 9607623.520 | 8654086.380 | -64193471.914 |
| -194.071 | -172.855 | -172.855 | 0.000 | 0.000 | 0.000 |
| -172.855 | -194.071 | -172.855 | 0.000 | 0.000 | 0.000 |
| -172.855 | -172.855 | -194.071 | 0.000 | 0.000 | 0.000 |
| -73007831.658 | -69859190.959 | -69380748.978 | -2310159.163 | 4346985.182 | 4216079.781 |
| -68740248.492 | -72140820.240 | -68504030.180 | 4523909.111 | -2297907.083 | 4357103.132 |
| -68461037.865 | -68258805.027 | -72495198.994 | 4723809.940 | 4554999.310 | -1912113.570 |
| -194.072 | -172.856 | -172.856 | -0.000 | -0.000 | -0.000 |
| -172.856 | -194.072 | -172.856 | -0.000 | -0.000 | -0.000 |
| -172.856 | -172.856 | -194.072 | -0.000 | -0.000 | -0.000 |
| 73761142.143 | 75012807.929 | 75326187.125 | -6453954.274 | 699230.372 | -622359.926 |
| 75737745.689 | 74452879.433 | 75776400.857 | 1083613.402 | -6435004.250 | -417110.033 |
| 76520614.502 | 76650506.259 | 74717134.072 | 794046.374 | -585426.204 | -6587979.852 |
| -194.066 | -172.851 | -172.851 | 0.000 | 0.000 | 0.000 |
| -172.851 | -194.066 | -172.851 | 0.000 | 0.000 | 0.000 |
| -172.851 | -172.851 | -194.066 | 0.000 | 0.000 | 0.000 |
| -7242990.393 | -6911995.333 | -6906869.013 | -271741.621 | 391681.892 | 342917.732 |
| -6798152.258 | -7132069.728 | -6822109.167 | 425943.057 | -287230.529 | 313741.270 |
| -6949003.651 | -6950257.024 | -7244710.359 | 412359.907 | 298857.290 | -298742.455 |
| -194.076 | -172.861 | -172.861 | -0.000 | -0.000 | -0.000 |
| -172.861 | -194.076 | -172.861 | -0.000 | -0.000 | -0.000 |
| -172.861 | -172.861 | -194.076 | -0.000 | -0.000 | -0.000 |
| 3211166.359 | 3138835.920 | 3138835.741 | 89532.987 | 0.005 | 0.005 |
| 3138835.920 | 3211166.359 | 3138835.741 | 0.005 | 89532.987 | 0.005 |
| 3138835.920 | 3138835.741 | 3211166.359 | 0.005 | 0.005 | 89532.987 |
| 710.821 | 266.379 | 216.257 | -0.000 | -0.000 | -0.000 |
| 266.379 | 710.821 | 216.257 | 0.000 | 0.000 | -0.000 |
| 216.257 | 216.257 | 521.244 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 115.350 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 115.350 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 36.905 |
| 710.821 | 266.379 | 216.257 | 0.000 | -0.000 | -0.000 |
| 266.379 | 710.821 | 216.257 | 0.000 | -0.000 | -0.000 |
| 216.257 | 216.257 | 521.244 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 115.350 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 115.350 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 36.905 |
| 710.821 | 266.379 | 216.257 | -0.000 | 0.000 | -0.000 |
| 266.379 | 710.821 | 216.257 | 0.000 | 0.000 | -0.000 |
| 216.257 | 216.257 | 521.245 | -0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 115.350 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 115.350 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 36.905 |
| 710.821 | 266.379 | 216.257 | 0.000 | -0.000 | 0.000 |
| 266.379 | 710.821 | 216.257 | 0.000 | -0.000 | -0.000 |
| 216.257 | 216.257 | 521.244 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 115.350 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 115.350 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 36.905 |
| 710.824 | 266.380 | 216.258 | -0.000 | 0.000 | -0.000 |
| 266.380 | 710.824 | 216.258 | -0.000 | 0.000 | -0.000 |
| 216.258 | 216.258 | 521.248 | -0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 115.350 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 115.350 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 36.905 |
| 710.818 | 266.378 | 216.256 | 0.000 | 0.000 | -0.000 |
| 266.378 | 710.818 | 216.256 | 0.000 | -0.000 | -0.000 |
| 216.256 | 216.256 | 521.240 | 0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 115.350 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 115.350 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 36.905 |
| 173.933 | 173.219 | 137.159 | -0.000 | -0.000 | 0.000 |
| 173.219 | 173.933 | 137.159 | -0.000 | -0.000 | 0.000 |
| 137.159 | 137.159 | 820.760 | -0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 68.975 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 68.975 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 19.386 |
| 173.933 | 173.219 | 137.159 | 0.000 | 0.000 | -0.000 |
| 173.219 | 173.933 | 137.159 | 0.000 | 0.000 | 0.000 |
| 137.159 | 137.159 | 820.760 | 0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 68.975 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 68.975 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 19.386 |
| 173.933 | 173.219 | 137.159 | -0.000 | -0.000 | -0.000 |
| 173.219 | 173.933 | 137.159 | -0.000 | -0.000 | -0.000 |
| 137.159 | 137.159 | 820.760 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 68.975 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 68.975 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 19.386 |
| 173.933 | 173.219 | 137.159 | 0.000 | 0.000 | 0.000 |
| 173.219 | 173.933 | 137.159 | 0.000 | 0.000 | 0.000 |
| 137.159 | 137.159 | 820.760 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 68.975 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 68.975 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 19.386 |
| 173.933 | 173.220 | 137.159 | -0.000 | -0.000 | -0.000 |
| 173.220 | 173.933 | 137.159 | -0.000 | -0.000 | -0.000 |
| 137.159 | 137.159 | 820.763 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 68.975 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 68.975 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 19.386 |
| 173.932 | 173.218 | 137.158 | 0.000 | 0.000 | -0.000 |
| 173.218 | 173.932 | 137.158 | 0.000 | 0.000 | 0.000 |
| 137.158 | 137.158 | 820.757 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 68.975 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 68.975 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 19.386 |
| 643.933 | 84.000 | 84.000 | 0.000 | -0.000 | 0.000 |
| 84.000 | 643.933 | 84.000 | 0.000 | -0.000 | -0.000 |
| 84.000 | 84.000 | 643.933 | 0.000 | -0.000 | 0.000 |
| 194.673 | 194.673 | 194.673 | -161.722 | -0.000 | 0.000 |
| 194.673 | 194.673 | 194.673 | 0.000 | -161.722 | 0.000 |
| 194.673 | 194.673 | 194.673 | -0.000 | -0.000 | -161.722 |
| 335.723 | -25.895 | -25.895 | 0.000 | 0.000 | 0.000 |
| -25.895 | 335.723 | -25.895 | -0.000 | 0.000 | -0.000 |
| -25.895 | -25.895 | 335.723 | 0.000 | 0.000 | 0.000 |
| -194.673 | -194.673 | -194.673 | -161.722 | 0.000 | -0.000 |
| -194.673 | -194.673 | -194.673 | 0.000 | -161.722 | -0.000 |
| -194.673 | -194.673 | -194.673 | -0.000 | 0.000 | -161.722 |
| 492.330 | 60.745 | 60.745 | -0.000 | -0.000 | -0.000 |
| 60.745 | 492.330 | 60.745 | 0.000 | -0.000 | -0.000 |
| 60.745 | 60.745 | 492.330 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 129.394 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 129.394 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 129.394 |
| 643.933 | 84.000 | 84.000 | -0.000 | -0.000 | -0.000 |
| 84.000 | 643.933 | 84.000 | -0.000 | 0.000 | 0.000 |
| 84.000 | 84.000 | 643.933 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 129.394 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 129.394 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 129.394 |
| 455.794 | 51.905 | 51.905 | 0.000 | -0.000 | -0.000 |
| 51.905 | 455.794 | 51.905 | -0.000 | -0.000 | -0.000 |
| 51.905 | 51.905 | 455.794 | -0.000 | -0.000 | -0.000 |
| 902693.021 | 8192698.741 | 4099023.269 | -10949206.818 | 24163.650 | -53541.145 |
| 4280312.184 | 691479.352 | 9183442.998 | -24322.375 | -10176404.148 | 11160.222 |
| 8092253.066 | 3960227.058 | 1361758.437 | 15442.804 | -23308.868 | -10182327.576 |
| 453.158 | 48.164 | 48.164 | -0.000 | 0.000 | 0.000 |
| 48.164 | 453.158 | 48.164 | 0.000 | 0.000 | 0.000 |
| 48.164 | 48.164 | 453.158 | -0.000 | 0.000 | -0.000 |
| -761679.287 | -9051200.606 | -4260873.627 | -10396451.280 | -12715.149 | -21736.764 |
| -3922192.877 | -252519.660 | -9111123.001 | -161.804 | -9634610.450 | 4300.473 |
| -8643223.707 | -4140801.666 | -949379.572 | 15482.655 | 28146.719 | -10362645.437 |
| 515.779 | 473.146 | 2008.543 | 24.206 | 0.000 | 0.000 |
| 508.668 | 570.995 | 1848.280 | -20.475 | -0.000 | 0.000 |
| 86.267 | 86.267 | 557.506 | -0.000 | 0.000 | -0.000 |
| 376.022 | 327.614 | 2030.762 | 78.632 | 0.000 | -0.000 |
| 351.818 | 351.818 | 2030.762 | -0.000 | 78.632 | 24.204 |
| 348.627 | 348.627 | 2012.340 | -0.000 | 24.210 | 21.670 |
| 233.392 | 86.063 | 86.267 | 24.842 | -0.000 | 0.000 |
| -224.757 | -180.762 | -2015.826 | -24.214 | -0.000 | -0.000 |
| 86.267 | 86.267 | 557.506 | 0.000 | -0.000 | 0.000 |
| -327.614 | -376.022 | -2030.762 | 78.632 | 0.000 | -0.000 |
| -351.818 | -351.818 | -2030.762 | 0.000 | 78.632 | 24.204 |
| -348.627 | -348.627 | -2012.339 | 0.000 | 24.210 | 21.670 |
| 233.392 | 86.063 | 86.267 | 24.842 | -0.000 | -0.000 |
| 128.439 | 213.440 | 145.105 | -24.463 | -0.000 | 0.000 |
| 66.183 | 66.183 | 288.267 | -0.000 | -0.000 | -0.000 |
| 24.842 | -24.842 | 0.000 | 101.802 | -0.000 | -0.000 |
| 48.152 | 48.152 | 212.454 | -0.000 | 60.790 | 18.655 |
| 35.841 | 35.841 | 151.479 | 0.000 | 19.697 | 40.985 |
| 143.957 | 69.142 | -149.903 | 20.146 | -0.000 | -0.000 |
| 86.825 | 152.900 | -151.507 | -24.348 | 0.000 | -0.000 |
| 86.267 | 86.267 | 557.506 | 0.000 | -0.000 | 0.000 |
| -29.498 | -66.807 | -212.454 | 60.790 | -0.000 | -0.000 |
| -48.152 | -48.152 | -212.454 | 0.000 | 60.790 | 18.655 |
| -35.841 | -35.841 | -151.479 | -0.000 | 19.697 | 40.985 |
| 199.119 | 106.906 | 42.304 | 24.507 | -0.000 | -0.000 |
| 106.906 | 199.119 | 42.304 | -24.507 | 0.000 | -0.000 |
| 3.977 | 3.977 | 18.438 | -0.000 | 0.000 | 0.000 |
| 6.708 | 9.433 | 39.002 | 2.014 | 0.000 | -0.000 |
| 11.993 | 11.992 | 69.226 | -0.000 | 22.822 | 22.668 |
| 7.269 | 7.269 | 34.992 | 0.000 | 5.368 | 43.506 |
| 184.973 | 97.930 | 10.729 | 19.872 | -0.000 | -0.000 |
| 104.687 | 192.151 | 15.346 | -24.478 | -0.000 | 0.000 |
| -3.974 | -3.974 | -18.419 | 0.000 | -0.000 | -0.000 |
| 24.842 | -24.842 | -0.000 | 101.802 | 0.000 | 0.000 |
| -11.993 | -11.992 | -69.226 | 0.000 | 22.822 | 22.668 |
| -7.269 | -7.269 | -34.992 | -0.000 | 5.368 | 43.506 |
| 196.280 | 83.971 | -102.089 | -24.387 | 0.000 | 0.000 |
| 82.829 | 226.516 | 84.439 | 24.148 | 0.000 | 0.000 |
| -16.522 | -16.522 | -166.076 | 0.000 | -0.000 | 0.000 |
| -24.148 | 24.148 | 0.000 | 101.882 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 101.882 | -24.148 |
| -9.074 | -9.074 | -146.758 | -0.000 | -24.375 | 56.906 |
| 226.516 | 82.829 | 84.439 | -24.148 | 0.000 | -0.000 |
| 82.829 | 226.516 | 84.439 | 24.148 | 0.000 | 0.000 |
| 14.754 | 14.754 | 166.074 | -0.000 | -0.000 | -0.000 |
| 15.638 | 15.638 | 166.075 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 101.882 | -24.148 |
| 9.074 | 9.074 | 146.758 | 0.000 | -24.375 | 56.906 |
| 226.516 | 82.829 | 84.439 | -24.148 | -0.000 | 0.000 |
| 98.256 | 182.985 | 13.405 | 21.398 | 0.000 | 0.000 |
| -2.450 | -2.450 | -16.624 | 0.000 | 0.000 | 0.000 |
| -24.148 | 24.148 | 0.000 | 101.882 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 101.882 | -24.148 |
| 0.000 | 0.000 | 0.000 | -0.000 | -24.148 | 71.843 |
| 180.226 | 97.776 | 16.607 | 0.000 | 0.000 | -0.000 |
| 82.829 | 226.515 | 84.439 | 24.148 | -0.000 | -0.000 |
| 83.555 | 83.555 | 553.963 | 0.000 | 0.000 | -0.000 |
| -20.434 | 30.345 | 80.149 | 62.606 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 101.882 | -24.148 |
| -0.000 | -0.000 | -0.000 | 0.000 | -24.148 | 71.843 |
| 190.397 | 101.197 | 25.530 | -24.563 | 0.000 | 0.000 |
| 97.073 | 184.059 | 24.898 | 20.146 | 0.000 | 0.000 |
| -1.049 | -1.049 | -1.698 | 0.000 | -0.000 | 0.000 |
| -0.273 | -0.276 | -2.181 | -0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 101.882 | -24.148 |
| 0.000 | 0.000 | 0.000 | -0.000 | -24.148 | 71.843 |
| 181.268 | 89.764 | -8.554 | -14.898 | 0.000 | -0.000 |
| 101.736 | 190.328 | 30.286 | 24.567 | -0.000 | 0.000 |
| -0.773 | -0.773 | 1.501 | -0.000 | 0.000 | -0.000 |
| 0.275 | 0.274 | 2.181 | 0.002 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 101.882 | -24.148 |
| -0.000 | -0.000 | -0.000 | 0.000 | -24.148 | 71.843 |
| 661.631 | 236.562 | 527.257 | 0.000 | -378.709 | 0.001 |
| -703.902 | 4784.297 | 908.340 | -0.000 | -691.839 | -0.001 |
| 339.927 | 4588.962 | 690.053 | 0.000 | 616.646 | 0.001 |
| 731.295 | 4114.705 | 157.845 | 16.722 | 732.393 | 24.420 |
| 104.981 | -20.717 | 15.953 | -0.000 | 91.027 | 0.000 |
| 277.705 | 4586.948 | 607.102 | 14.833 | 569.545 | -32.637 |
| -382.246 | -5361.093 | -1223.475 | -0.000 | -289.539 | -0.001 |
| -13.097 | 464.334 | 98.398 | 0.000 | -22.761 | -0.000 |
| 55.511 | 104.071 | 249.733 | -0.000 | 15.953 | -0.000 |
| -731.295 | -4114.705 | -157.845 | 16.722 | -732.393 | 24.420 |
| -261.813 | -4596.925 | -586.842 | -0.000 | -557.270 | -0.001 |
| 564.304 | -4853.080 | -1002.897 | 42.007 | 612.301 | 554.942 |
| 138.843 | 510.392 | 172.582 | 0.000 | 60.795 | 0.000 |
| -6.462 | 393.777 | 50.143 | 0.000 | 55.654 | 0.000 |
| 106.931 | 501.420 | 185.514 | 0.000 | 93.765 | 0.000 |
| 58.406 | 513.895 | 126.141 | 13.842 | 40.596 | 7.916 |
| 104.981 | -20.717 | 15.953 | -0.000 | 91.027 | -0.000 |
| 41.443 | 11.678 | 40.282 | -17.909 | -33.987 | -80.172 |
| 21.638 | -433.765 | 49.063 | 0.000 | -24.591 | 0.000 |
| 36.938 | 97.008 | -36.350 | -0.000 | 95.717 | -0.000 |
| 109.241 | -427.218 | 7.194 | 0.000 | 104.110 | 0.000 |
| -58.405 | -513.895 | -126.141 | 13.842 | -40.596 | 7.916 |
| -25.181 | 19.406 | -31.488 | -0.000 | 74.030 | -0.000 |
| -41.443 | -11.678 | -40.282 | -17.909 | 33.987 | -80.172 |
| 181.728 | 44.999 | 83.812 | 0.000 | 61.151 | 0.000 |
| 2.394 | 43.923 | -16.221 | 0.000 | 9.136 | 0.000 |
| 44.047 | 42.339 | 147.018 | 0.000 | 10.221 | 0.000 |
| 3.481 | 42.178 | 10.671 | 25.786 | 0.915 | 10.239 |
| 64.382 | 21.860 | 31.258 | 0.000 | 48.796 | 0.000 |
| 2.229 | 16.448 | 7.071 | -8.455 | -5.238 | 32.017 |
| 119.832 | -56.592 | 59.712 | 0.000 | 48.687 | 0.000 |
| -27.823 | -48.699 | -17.241 | 0.000 | -6.446 | 0.000 |
| 62.879 | -40.160 | 106.936 | 0.000 | 39.908 | -0.000 |
| -3.481 | -42.178 | -10.671 | 25.786 | -0.915 | 10.239 |
| 54.763 | 9.161 | 12.474 | -0.000 | 61.271 | -0.000 |
| -2.229 | -16.448 | -7.071 | -8.455 | 5.238 | 32.017 |
| -545.945 | 73.740 | 786.396 | -0.000 | 0.000 | 0.000 |
| -576288246.901 | -7891086681.802 | 2575670382.230 | -17431618.414 | 2045550.347 | -25167030.466 |
| 786.062 | 786.062 | -953.347 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 44.251 | 0.000 | 0.000 |
| -0.001 | 0.002 | -0.000 | 0.000 | 44.251 | 0.000 |
| -5738004134.669 | -2399377894.752 | 2734610978.912 | -5243727.809 | -10931771.901 | -2970658517.835 |
| -545.943 | 73.740 | 786.394 | 0.000 | -0.000 | -0.000 |
| 3198240695.527 | 1954764466.536 | -934552609.432 | 1684022.924 | 31335915.914 | -8374442.446 |
| 786.064 | 786.064 | -953.350 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 44.250 | -0.000 | 0.000 |
| 0.001 | -0.002 | 0.000 | -0.000 | 44.250 | -0.000 |
| 5436603066.322 | 2383368939.224 | -2913200332.337 | -16199930.263 | 26045165.623 | -2785525690.831 |
| -329936466.443 | -296085443.985 | 321335561.435 | 9991346.397 | 1356777.940 | -2567431.562 |
| -49651285.956 | -826759678.376 | 241383155.782 | -938878.768 | -113963.258 | 9433783.452 |
| 786.054 | 786.054 | -953.331 | 0.000 | 0.000 | -0.000 |
| 0.001 | 0.000 | -0.001 | 44.251 | -0.000 | -0.000 |
| -0.002 | 0.003 | -0.001 | 0.000 | 44.251 | 0.000 |
| -642127330.324 | -252500717.094 | 244623478.917 | 1601828.391 | 3144934.497 | -330370894.530 |
| 58852136.779 | 766696624.818 | -271375640.744 | -2886737.807 | 642994.963 | -3148711.266 |
| 311861571.580 | 273453192.795 | -308273435.853 | -328913.020 | -1241115.404 | -1698234.542 |
| 786.071 | 786.071 | -953.366 | -0.000 | -0.000 | 0.000 |
| -0.001 | -0.000 | 0.001 | 44.251 | -0.000 | 0.000 |
| 0.002 | -0.003 | 0.001 | -0.000 | 44.251 | -0.000 |
| 629944680.217 | 266636061.758 | -240101134.601 | -340710.365 | -443312.258 | -333956042.160 |
| -30254693.477 | -28341452.044 | 29128728.605 | -1542.619 | 103151.239 | 8963.537 |
| -5167125.957 | -79358912.883 | 25791021.480 | -65387.872 | -9552.188 | -141194.162 |
| 785.975 | 785.975 | -953.175 | 0.000 | 0.000 | 0.000 |
| 0.029 | -0.023 | -0.006 | 44.251 | 0.000 | 0.000 |
| 0.000 | 0.005 | -0.006 | -0.000 | 44.251 | -0.000 |
| -61690522.167 | -24943818.230 | 26568902.970 | -161556.426 | -228516.840 | -31080435.843 |
| 2863273.337 | 87040049.736 | -21940113.498 | 196218.626 | -3422.261 | -990394.464 |
| 31058094.024 | 27690304.542 | -31047702.026 | 23724.174 | -153945.057 | -110350.459 |
| 786.150 | 786.150 | -953.522 | 0.000 | -0.000 | -0.000 |
| -0.029 | 0.023 | 0.006 | 44.251 | -0.000 | -0.000 |
| -0.000 | -0.005 | 0.006 | 0.000 | 44.251 | -0.000 |
| 63994186.099 | 26227176.119 | -23881290.180 | -76303.170 | 186222.430 | -35209598.794 |
| 399.635 | 114.929 | 114.929 | 0.000 | 0.000 | -0.000 |
| 114.929 | 399.635 | 114.929 | 0.000 | -0.000 | -0.000 |
| 114.929 | 114.929 | 399.635 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -127.168 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -127.168 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -127.168 |
| 399.635 | 114.929 | 114.929 | 0.000 | -0.000 | 0.000 |
| 114.929 | 399.635 | 114.929 | 0.000 | -0.000 | 0.000 |
| 114.929 | 114.929 | 399.634 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -127.168 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -127.168 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -127.168 |
| 399.635 | 114.929 | 114.929 | 0.000 | -0.000 | -0.000 |
| 114.929 | 399.635 | 114.929 | 0.000 | -0.000 | -0.000 |
| 114.929 | 114.929 | 399.635 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -16.366 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -16.366 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -16.366 |
| 399.634 | 114.929 | 114.929 | -0.000 | 0.000 | 0.000 |
| 114.929 | 399.634 | 114.929 | 0.000 | 0.000 | -0.000 |
| 114.929 | 114.929 | 399.634 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -16.366 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -16.366 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -16.366 |
| 399.637 | 114.930 | 114.930 | 0.000 | -0.000 | -0.000 |
| 114.930 | 399.637 | 114.930 | 0.000 | -0.000 | -0.000 |
| 114.930 | 114.930 | 399.637 | 0.000 | 0.000 | -0.000 |
| 0.001 | 0.000 | -0.000 | -127.168 | 0.000 | 0.000 |
| 0.000 | 0.001 | -0.000 | 0.000 | -127.168 | -0.000 |
| 0.000 | -0.000 | 0.001 | 0.000 | 0.000 | -127.168 |
| 399.632 | 114.929 | 114.929 | -0.000 | 0.000 | 0.000 |
| 114.929 | 399.632 | 114.929 | -0.000 | 0.000 | 0.000 |
| 114.929 | 114.929 | 399.632 | -0.000 | 0.000 | 0.000 |
| -0.001 | -0.000 | 0.000 | -127.168 | -0.000 | 0.000 |
| -0.000 | -0.001 | 0.000 | 0.000 | -127.168 | 0.000 |
| -0.000 | 0.000 | -0.001 | 0.000 | -0.000 | -127.168 |
| 189.725 | 30.352 | 70.528 | 0.000 | 0.000 | -0.000 |
| 280.497 | 144.719 | 91.137 | -0.000 | 0.000 | 0.001 |
| 600.240 | -7.981 | 26.822 | -0.000 | 0.000 | 0.004 |
| 193.757 | -17.432 | 85.005 | 63.892 | 0.000 | -0.001 |
| 199.055 | -17.909 | 87.329 | 0.000 | 58.223 | -0.001 |
| 185.578 | 12.203 | 98.600 | 0.000 | 0.000 | -403.767 |
| -225.909 | 0.890 | -68.247 | -0.000 | -0.000 | 0.001 |
| -277.092 | 98.854 | -90.736 | 0.000 | -0.000 | -0.001 |
| -387.768 | -148.472 | -171.909 | 0.000 | -0.000 | 0.001 |
| -193.757 | 17.432 | -85.005 | 63.892 | -0.000 | 0.001 |
| -199.055 | 17.909 | -87.329 | -0.000 | 58.223 | 0.001 |
| -185.352 | -12.098 | -98.395 | -0.000 | -0.000 | -403.633 |
| 140.821 | 45.171 | 20.043 | 0.000 | 0.000 | -0.000 |
| 54.043 | 167.112 | 53.396 | -0.000 | 0.000 | 0.000 |
| -34.197 | 13.079 | 34.162 | 0.000 | 0.000 | -0.000 |
| 52.753 | -4.746 | 23.144 | 44.801 | 0.000 | -0.000 |
| 27.641 | 2.209 | 8.900 | -0.000 | 12.388 | -0.000 |
| 54.687 | -4.920 | 23.992 | 0.000 | 0.000 | 45.155 |
| 22.259 | 14.806 | -50.171 | -0.000 | -0.000 | 0.000 |
| 54.043 | 167.111 | 53.396 | 0.000 | -0.000 | 0.000 |
| 5.565 | 5.052 | 17.140 | 0.000 | -0.000 | 0.000 |
| -52.753 | 4.746 | -23.144 | 44.801 | -0.000 | 0.000 |
| -17.960 | 0.383 | 12.904 | 0.000 | 9.920 | -0.000 |
| -54.690 | 4.920 | -23.993 | -0.000 | -0.000 | 45.154 |
| 19.569 | 5.723 | 14.642 | -0.000 | 0.000 | -0.000 |
| 37.610 | 137.458 | 37.879 | 0.000 | 0.000 | -0.000 |
| 2.142 | 4.840 | 26.083 | 0.000 | 0.000 | -0.000 |
| -0.676 | 0.266 | -5.164 | -1.000 | -0.000 | -0.000 |
| 2.004 | -0.091 | -0.554 | -0.000 | 12.813 | -0.000 |
| -0.454 | 0.226 | -4.597 | -0.000 | -0.000 | 6.353 |
| 14.893 | 0.998 | 5.637 | -0.000 | -0.000 | 0.000 |
| 30.840 | 136.189 | 34.407 | -0.000 | -0.000 | 0.000 |
| -1.329 | 5.105 | 26.820 | -0.000 | -0.000 | 0.000 |
| 0.676 | -0.266 | 5.164 | -1.000 | 0.000 | 0.000 |
| -2.004 | 0.091 | 0.554 | 0.000 | 12.813 | 0.000 |
| 0.454 | -0.226 | 4.597 | 0.000 | 0.000 | 6.353 |
| 302.136 | 254.113 | 254.113 | -0.000 | 0.000 | 0.000 |
| 254.113 | 302.136 | 254.113 | -0.000 | 0.000 | -0.000 |
| 254.113 | 254.113 | 302.136 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -76.528 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -76.528 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -76.528 |
| 302.136 | 254.113 | 254.113 | 0.000 | -0.000 | -0.000 |
| 254.113 | 302.136 | 254.113 | 0.000 | -0.000 | 0.000 |
| 254.113 | 254.113 | 302.136 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | -76.528 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -76.528 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | -76.528 |
| 298.251 | 256.056 | 256.056 | 0.000 | 0.000 | 0.000 |
| 256.056 | 298.251 | 256.056 | 0.000 | 0.000 | -0.000 |
| 256.056 | 256.056 | 298.251 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 19.834 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 19.834 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 19.834 |
| 302.136 | 254.113 | 254.113 | 0.000 | -0.000 | -0.000 |
| 254.113 | 302.136 | 254.113 | 0.000 | -0.000 | 0.000 |
| 254.113 | 254.113 | 302.136 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 19.834 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 19.834 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 19.834 |
| 298.252 | 256.056 | 256.056 | 0.000 | 0.000 | 0.000 |
| 256.056 | 298.252 | 256.056 | 0.000 | 0.000 | 0.000 |
| 256.056 | 256.056 | 298.252 | 0.000 | 0.000 | 0.000 |
| 0.002 | -0.002 | -0.003 | -162.538 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 19.834 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 19.834 |
| 298.250 | 256.055 | 256.055 | -0.000 | -0.000 | -0.000 |
| 256.055 | 298.250 | 256.055 | -0.000 | -0.000 | 0.000 |
| 256.055 | 256.055 | 298.250 | -0.000 | -0.000 | -0.000 |
| -0.002 | 0.002 | 0.003 | -162.538 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 19.834 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 19.834 |
| 76431.005 | 68343.818 | 68343.818 | 0.000 | -0.000 | 0.000 |
| 68343.818 | 76431.005 | 68343.818 | 0.000 | -0.000 | 0.000 |
| 68343.818 | 68343.818 | 76431.005 | 0.000 | -0.000 | 0.000 |
| 0.059 | -0.029 | -0.030 | -23.284 | -0.000 | -0.000 |
| -0.030 | 0.059 | -0.029 | -0.000 | -23.284 | -0.000 |
| -0.030 | -0.030 | 0.059 | -0.000 | 0.000 | -23.284 |
| 76430.505 | 68343.404 | 68343.405 | -0.000 | -0.000 | -0.000 |
| 68343.404 | 76430.505 | 68343.405 | -0.000 | 0.000 | 0.000 |
| 68343.404 | 68343.404 | 76430.505 | -0.000 | 0.000 | -0.000 |
| -0.059 | 0.030 | 0.030 | -23.284 | 0.000 | 0.000 |
| 0.029 | -0.058 | 0.029 | 0.000 | -23.284 | 0.000 |
| 0.030 | 0.030 | -0.059 | 0.000 | 0.000 | -23.284 |
| 76433.268 | 68345.674 | 68345.674 | 0.000 | 0.000 | -0.000 |
| 68345.674 | 76433.268 | 68345.674 | -0.000 | 0.000 | 0.000 |
| 68345.674 | 68345.674 | 76433.268 | 0.000 | 0.000 | 0.000 |
| 0.584 | -0.298 | -0.301 | -23.762 | 0.000 | -0.000 |
| -0.298 | 0.584 | -0.301 | -0.000 | -23.762 | -0.000 |
| -0.298 | -0.301 | 0.584 | -0.000 | 0.000 | -23.762 |
| 76425.366 | 68342.988 | 68342.988 | -0.000 | 0.000 | -0.000 |
| 68342.988 | 76425.366 | 68342.988 | 0.000 | -0.000 | -0.000 |
| 68342.988 | 68342.988 | 76425.366 | -0.000 | -0.000 | -0.000 |
| -0.584 | 0.298 | 0.301 | -23.762 | -0.000 | 0.000 |
| 0.298 | -0.584 | 0.301 | 0.000 | -23.762 | 0.000 |
| 0.298 | 0.301 | -0.584 | 0.000 | 0.000 | -23.762 |
| 76455.621 | 68364.362 | 68364.362 | 0.000 | 0.000 | 0.000 |
| 68364.362 | 76455.621 | 68364.362 | 0.000 | 0.000 | 0.000 |
| 68364.362 | 68364.362 | 76455.621 | 0.000 | 0.000 | 0.000 |
| -0.040 | -0.053 | -0.057 | -23.371 | 0.000 | 0.000 |
| -0.053 | -0.040 | -0.057 | 0.000 | -23.371 | 0.000 |
| -0.053 | -0.057 | -0.040 | 0.000 | 0.000 | -23.371 |
| 76405.952 | 68322.830 | 68322.830 | -0.000 | -0.000 | -0.000 |
| 68322.830 | 76405.952 | 68322.830 | -0.000 | -0.000 | -0.000 |
| 68322.830 | 68322.830 | 76405.952 | -0.000 | -0.000 | -0.000 |
| 0.040 | 0.053 | 0.057 | -23.371 | -0.000 | -0.000 |
| 0.053 | 0.040 | 0.057 | 0.000 | -23.371 | 0.000 |
| 0.053 | 0.057 | 0.040 | -0.000 | -0.000 | -23.371 |
| 3.419 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | 3.419 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 3.419 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 3.419 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | 3.419 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 3.419 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 3.419 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | 3.419 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 3.419 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 352.537 | 260.631 | 260.631 | 0.000 | -0.000 | -0.000 |
| 260.631 | 352.537 | 260.631 | 0.000 | -0.000 | -0.000 |
| 260.631 | 260.631 | 352.537 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -103.703 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -103.703 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -103.703 |
| 352.537 | 260.631 | 260.631 | -0.000 | -0.000 | 0.000 |
| 260.631 | 352.537 | 260.631 | -0.000 | -0.000 | 0.000 |
| 260.631 | 260.631 | 352.537 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -103.703 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -103.703 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | -103.703 |
| 352.537 | 260.631 | 260.631 | 0.000 | -0.000 | -0.000 |
| 260.631 | 352.537 | 260.631 | 0.000 | -0.000 | -0.000 |
| 260.631 | 260.631 | 352.537 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -103.703 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -103.703 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | -103.703 |
| 352.536 | 260.631 | 260.631 | 0.000 | 0.000 | 0.000 |
| 260.631 | 352.536 | 260.631 | 0.000 | 0.000 | 0.000 |
| 260.631 | 260.631 | 352.536 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -103.703 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -103.703 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.000 | -103.703 |
| 352.538 | 260.632 | 260.632 | 0.000 | 0.000 | -0.000 |
| 260.632 | 352.538 | 260.632 | 0.000 | -0.000 | -0.000 |
| 260.632 | 260.632 | 352.538 | 0.000 | -0.000 | -0.000 |
| 0.001 | 0.000 | -0.000 | -103.703 | -0.000 | -0.000 |
| 0.000 | 0.001 | -0.000 | 0.000 | -103.703 | -0.000 |
| 0.000 | -0.000 | 0.001 | -0.000 | -0.000 | -103.703 |
| 352.535 | 260.630 | 260.630 | -0.000 | 0.000 | 0.000 |
| 260.630 | 352.535 | 260.630 | 0.000 | 0.000 | 0.000 |
| 260.630 | 260.630 | 352.535 | -0.000 | 0.000 | 0.000 |
| -0.001 | -0.000 | 0.000 | -103.703 | 0.000 | 0.000 |
| -0.000 | -0.001 | 0.000 | -0.000 | -103.703 | 0.000 |
| -0.000 | 0.000 | -0.001 | -0.000 | -0.000 | -103.703 |
| 315.035 | 204.751 | 204.751 | -0.000 | -0.000 | -0.000 |
| 204.751 | 315.035 | 204.751 | -0.000 | -0.000 | -0.000 |
| 204.751 | 204.751 | 315.035 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 25.407 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 25.407 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 25.407 |
| 315.035 | 204.751 | 204.751 | -0.000 | 0.000 | 0.000 |
| 204.751 | 315.035 | 204.751 | 0.000 | 0.000 | 0.000 |
| 204.751 | 204.751 | 315.035 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 25.407 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 25.407 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 25.407 |
| 315.035 | 204.751 | 204.751 | 0.000 | -0.000 | -0.000 |
| 204.751 | 315.035 | 204.751 | 0.000 | -0.000 | -0.000 |
| 204.751 | 204.751 | 315.035 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 25.407 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 25.407 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -0.000 | 25.407 |
| 315.035 | 204.751 | 204.751 | 0.000 | 0.000 | 0.000 |
| 204.751 | 315.035 | 204.751 | 0.000 | 0.000 | 0.000 |
| 204.751 | 204.751 | 315.035 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 25.407 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 25.407 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 25.407 |
| 315.036 | 204.751 | 204.751 | -0.000 | -0.000 | -0.000 |
| 204.751 | 315.036 | 204.751 | 0.000 | -0.000 | -0.000 |
| 204.751 | 204.751 | 315.036 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 25.407 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 25.407 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 0.000 | 25.407 |
| 315.034 | 204.750 | 204.750 | 0.000 | 0.000 | 0.000 |
| 204.750 | 315.034 | 204.750 | 0.000 | 0.000 | 0.000 |
| 204.750 | 204.750 | 315.034 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 25.407 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 25.407 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 25.407 |
| 352.180 | 260.862 | 214.806 | -0.000 | 0.000 | 0.000 |
| 260.862 | 352.180 | 214.806 | -0.000 | 0.000 | 0.000 |
| 232.407 | 232.407 | 207.428 | -0.000 | 0.000 | -0.000 |
| 0.001 | 0.001 | -0.000 | -101.671 | 0.000 | -0.000 |
| -0.005 | -0.004 | 0.001 | -0.000 | -440.323 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | -10.291 |
| 270.769 | 231.117 | 226.878 | 0.000 | -0.000 | 0.000 |
| 260.862 | 352.180 | 214.806 | 0.000 | -0.000 | 0.000 |
| 222.751 | 222.751 | 209.525 | 0.000 | -0.000 | 0.000 |
| -0.001 | -0.001 | 0.000 | -101.671 | -0.000 | -0.000 |
| 0.005 | 0.005 | -0.001 | 0.000 | -440.323 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -0.000 | -10.291 |
| 247.325 | 279.222 | 224.200 | -0.000 | 0.000 | 0.000 |
| 260.862 | 352.180 | 214.806 | -0.000 | 0.000 | -0.000 |
| 222.751 | 222.751 | 209.525 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -101.671 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 12.079 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 45.659 |
| 352.180 | 260.862 | 214.806 | 0.000 | -0.000 | -0.000 |
| 231.118 | 270.769 | 226.878 | 0.000 | -0.000 | 0.000 |
| 232.407 | 232.407 | 207.427 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -101.671 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 12.079 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 45.659 |
| 247.326 | 279.222 | 224.201 | -0.000 | 0.000 | 0.000 |
| 260.863 | 352.181 | 214.807 | 0.000 | 0.000 | -0.000 |
| 222.752 | 222.752 | 209.526 | -0.000 | 0.000 | -0.000 |
| 3768769.533 | 3771161.444 | 13593185.243 | -11321257.941 | -91127.233 | -48891.812 |
| 3737130.594 | -1160619.045 | 25655317.224 | 74817.644 | -848593.649 | 100749.581 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 45.659 |
| 247.325 | 279.223 | 224.199 | 0.000 | -0.000 | 0.000 |
| 279.221 | 247.326 | 224.199 | 0.000 | -0.000 | 0.000 |
| 232.407 | 232.407 | 207.427 | 0.000 | -0.000 | 0.000 |
| -4071065.367 | -3831640.019 | -14199878.413 | -10879103.753 | -21323.667 | -51542.158 |
| -3364735.652 | 1350344.250 | -26265624.651 | -159264.920 | -734674.835 | 113540.838 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 45.659 |
| -0.351 | 0.174 | 0.174 | 0.000 | 0.000 | 0.000 |
| 0.174 | -0.351 | 0.174 | 0.000 | 0.000 | 0.000 |
| 0.174 | 0.174 | -0.351 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.015 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.015 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.015 |
| -0.351 | 0.174 | 0.174 | -0.000 | -0.000 | -0.000 |
| 0.174 | -0.351 | 0.174 | -0.000 | -0.000 | -0.000 |
| 0.174 | 0.174 | -0.351 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.015 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.015 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.015 |
| -0.351 | 0.174 | 0.174 | 0.000 | 0.000 | 0.000 |
| 0.174 | -0.351 | 0.174 | 0.000 | 0.000 | 0.000 |
| 0.174 | 0.174 | -0.351 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.015 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.015 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.015 |
| -0.351 | 0.174 | 0.174 | -0.000 | -0.000 | -0.000 |
| 0.174 | -0.351 | 0.174 | -0.000 | -0.000 | -0.000 |
| 0.174 | 0.174 | -0.351 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.015 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.015 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.015 |
| -0.354 | 0.171 | 0.171 | 0.000 | 0.000 | 0.000 |
| 0.171 | -0.354 | 0.171 | 0.000 | 0.000 | 0.000 |
| 0.171 | 0.171 | -0.354 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.015 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.015 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.015 |
| -0.018 | 0.013 | 0.014 | -0.001 | 0.000 | 0.000 |
| -0.000 | 0.001 | 0.001 | -0.000 | 0.000 | -0.000 |
| -0.013 | -0.014 | 0.002 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.015 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.015 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.015 |
| 293.645 | 275.538 | 275.538 | 0.000 | 0.000 | 0.000 |
| 275.538 | 293.645 | 275.538 | 0.000 | 0.000 | 0.000 |
| 275.538 | 275.538 | 293.645 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 28.203 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 28.203 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 28.203 |
| 293.645 | 275.538 | 275.538 | -0.000 | -0.000 | -0.000 |
| 275.538 | 293.645 | 275.538 | -0.000 | -0.000 | -0.000 |
| 275.538 | 275.538 | 293.645 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 28.203 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 28.203 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 28.203 |
| 293.645 | 275.538 | 275.538 | -0.000 | -0.000 | -0.000 |
| 275.538 | 293.645 | 275.538 | -0.000 | 0.000 | 0.000 |
| 275.538 | 275.538 | 293.645 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 28.203 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 28.203 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 28.203 |
| 293.645 | 275.538 | 275.538 | -0.000 | -0.000 | 0.000 |
| 275.538 | 293.645 | 275.538 | 0.000 | -0.000 | -0.000 |
| 275.538 | 275.538 | 293.645 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 28.203 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 28.203 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 28.203 |
| 293.646 | 275.539 | 275.539 | 0.000 | 0.000 | 0.000 |
| 275.539 | 293.646 | 275.539 | 0.000 | 0.000 | 0.000 |
| 275.539 | 275.539 | 293.646 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 28.203 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 28.203 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 28.203 |
| 293.645 | 275.537 | 275.537 | -0.000 | -0.000 | -0.000 |
| 275.537 | 293.645 | 275.537 | 0.000 | -0.000 | -0.000 |
| 275.537 | 275.537 | 293.645 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 28.203 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 28.203 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 28.203 |
| 341.889 | 262.743 | 262.743 | -0.000 | 0.000 | 0.000 |
| 262.743 | 341.889 | 262.743 | 0.000 | 0.000 | 0.000 |
| 262.743 | 262.743 | 341.889 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 50.083 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 50.083 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 50.083 |
| 341.785 | 262.795 | 262.795 | -0.000 | -0.000 | -0.000 |
| 262.795 | 341.785 | 262.795 | -0.000 | -0.000 | -0.000 |
| 262.795 | 262.795 | 341.785 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 50.083 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 50.083 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 50.083 |
| 341.889 | 262.743 | 262.743 | -0.000 | 0.000 | 0.000 |
| 262.743 | 341.889 | 262.743 | 0.000 | 0.000 | 0.000 |
| 262.743 | 262.743 | 341.889 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 50.083 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 50.083 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 50.083 |
| 341.889 | 262.742 | 262.742 | -0.000 | -0.000 | -0.000 |
| 262.742 | 341.889 | 262.742 | -0.000 | -0.000 | -0.000 |
| 262.742 | 262.742 | 341.889 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 50.083 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 50.083 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 50.083 |
| 341.890 | 262.743 | 262.743 | 0.000 | 0.000 | 0.000 |
| 262.743 | 341.890 | 262.743 | 0.000 | 0.000 | 0.000 |
| 262.743 | 262.743 | 341.890 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 50.083 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 50.083 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 50.083 |
| 341.783 | 262.794 | 262.794 | -0.000 | -0.000 | -0.000 |
| 262.794 | 341.783 | 262.794 | -0.000 | -0.000 | -0.000 |
| 262.794 | 262.794 | 341.783 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 50.083 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 50.083 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 50.083 |
| 3.419 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | 3.419 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 3.419 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 3.419 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | 3.419 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 3.419 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 3.419 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | 3.419 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | 3.419 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 371.619 | 244.616 | 244.616 | -0.000 | -0.000 | 0.000 |
| 244.616 | 371.619 | 244.616 | 0.000 | -0.000 | -0.000 |
| 244.616 | 244.616 | 371.619 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 47.487 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 47.487 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 47.487 |
| 371.619 | 244.616 | 244.616 | -0.000 | 0.000 | -0.000 |
| 244.616 | 371.619 | 244.616 | 0.000 | 0.000 | 0.000 |
| 244.616 | 244.616 | 371.619 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 47.487 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 47.487 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 0.000 | 47.487 |
| 371.619 | 244.616 | 244.616 | 0.000 | 0.000 | 0.000 |
| 244.616 | 371.619 | 244.616 | 0.000 | -0.000 | -0.000 |
| 244.616 | 244.616 | 371.619 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 47.487 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 47.487 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 47.487 |
| 371.619 | 244.616 | 244.616 | -0.000 | 0.000 | -0.000 |
| 244.616 | 371.619 | 244.616 | -0.000 | -0.000 | 0.000 |
| 244.616 | 244.616 | 371.619 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 47.487 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 47.487 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 47.487 |
| 371.621 | 244.617 | 244.617 | 0.000 | -0.000 | 0.000 |
| 244.617 | 371.621 | 244.617 | 0.000 | 0.000 | 0.000 |
| 244.617 | 244.617 | 371.621 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 47.487 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 47.487 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 47.487 |
| 371.618 | 244.615 | 244.615 | -0.000 | 0.000 | -0.000 |
| 244.615 | 371.618 | 244.615 | -0.000 | -0.000 | 0.000 |
| 244.615 | 244.615 | 371.618 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 47.487 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 47.487 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 47.487 |
| 310.413 | 271.536 | 262.772 | -0.000 | -0.000 | 0.000 |
| 274.661 | 307.288 | 262.772 | -0.000 | -0.000 | -0.000 |
| 262.772 | 262.772 | 319.177 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 13.777 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 13.777 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 19.438 |
| 310.413 | 271.536 | 262.772 | 0.000 | 0.000 | -0.000 |
| 271.536 | 310.413 | 262.772 | 0.000 | 0.000 | 0.000 |
| 262.772 | 262.772 | 319.177 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 13.777 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 13.777 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -0.000 | 19.438 |
| 310.413 | 271.537 | 262.772 | -0.000 | 0.000 | 0.000 |
| 274.661 | 307.288 | 262.772 | -0.000 | 0.000 | -0.000 |
| 262.772 | 262.772 | 319.177 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 13.777 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 13.777 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 16.313 |
| 310.413 | 271.536 | 262.772 | 0.000 | -0.000 | 0.000 |
| 274.661 | 307.288 | 262.772 | -0.000 | 0.000 | 0.000 |
| 262.772 | 262.772 | 319.177 | 0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 13.777 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 13.777 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 16.313 |
| 307.289 | 274.662 | 262.773 | 0.000 | -0.000 | -0.000 |
| 271.537 | 310.414 | 262.773 | -0.000 | -0.000 | -0.000 |
| 262.773 | 262.773 | 319.178 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 13.777 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 13.777 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 19.438 |
| 310.412 | 271.536 | 262.771 | 0.000 | 0.000 | 0.000 |
| 271.536 | 310.412 | 262.771 | 0.000 | 0.000 | 0.000 |
| 262.771 | 262.771 | 319.176 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 13.777 | -0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 13.777 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 19.438 |
| 26.752 | 26.773 | 84.439 | 0.000 | 0.000 | 0.000 |
| 26.773 | 26.752 | 84.439 | 0.000 | 0.000 | 0.000 |
| 84.421 | 84.421 | 398.137 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 45.922 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 45.922 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.011 |
| 36.367 | 17.158 | 84.439 | -0.000 | -0.000 | -0.000 |
| 17.158 | 36.367 | 84.439 | 0.000 | -0.000 | -0.000 |
| 84.421 | 84.421 | 398.136 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 45.922 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 45.922 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.011 |
| 26.752 | 26.773 | 84.439 | 0.000 | 0.000 | 0.000 |
| 17.158 | 36.367 | 84.439 | 0.000 | -0.000 | 0.000 |
| 84.421 | 84.421 | 398.137 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 45.922 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 45.922 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.011 |
| 26.752 | 26.773 | 84.439 | -0.000 | -0.000 | -0.000 |
| 26.773 | 26.752 | 84.439 | -0.000 | -0.000 | -0.000 |
| 84.421 | 84.421 | 398.136 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 45.922 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 45.922 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.011 |
| 26.752 | 26.773 | 84.439 | 0.000 | 0.000 | 0.000 |
| 26.773 | 26.752 | 84.439 | 0.000 | 0.000 | 0.000 |
| 84.421 | 84.421 | 398.138 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 45.922 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 45.922 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.011 |
| 36.367 | 17.158 | 84.438 | -0.000 | -0.000 | -0.000 |
| 17.158 | 36.367 | 84.438 | -0.000 | -0.000 | -0.000 |
| 84.421 | 84.421 | 398.135 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 45.922 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 45.922 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.011 |
| 123.906 | 117.142 | 117.142 | -0.000 | -0.000 | -0.000 |
| 117.142 | 123.906 | 117.142 | -0.000 | -0.000 | -0.000 |
| 117.142 | 117.142 | 123.906 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -97.103 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -97.103 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -0.000 | -97.103 |
| 123.906 | 117.142 | 117.142 | 0.000 | 0.000 | 0.000 |
| 117.142 | 123.906 | 117.142 | 0.000 | 0.000 | 0.000 |
| 117.142 | 117.142 | 123.906 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -97.103 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -97.103 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.000 | -97.103 |
| 123.906 | 117.143 | 117.143 | -0.000 | -0.000 | -0.000 |
| 117.143 | 123.906 | 117.143 | -0.000 | -0.000 | -0.000 |
| 117.143 | 117.143 | 123.906 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -97.103 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -97.103 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -97.103 |
| 123.906 | 117.142 | 117.142 | 0.000 | 0.000 | 0.000 |
| 117.142 | 123.906 | 117.142 | 0.000 | 0.000 | 0.000 |
| 117.142 | 117.142 | 123.906 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -97.103 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -97.103 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -97.103 |
| 123.906 | 117.143 | 117.143 | -0.000 | -0.000 | -0.000 |
| 117.143 | 123.906 | 117.143 | -0.000 | -0.000 | -0.000 |
| 117.143 | 117.143 | 123.906 | -0.000 | -0.000 | -0.000 |
| 21514655.240 | 21571275.155 | 21360738.892 | 1325199.346 | 178413.447 | -115882.922 |
| 21498894.606 | 21596536.561 | 21380953.165 | 123776.776 | 1446305.769 | -32967.622 |
| 21546852.626 | 21449893.290 | 21424127.388 | -729.455 | 109858.531 | 1376910.426 |
| 123.905 | 117.142 | 117.142 | 0.000 | 0.000 | 0.000 |
| 117.142 | 123.905 | 117.142 | 0.000 | 0.000 | 0.000 |
| 117.142 | 117.142 | 123.905 | 0.000 | 0.000 | 0.000 |
| -22003088.831 | -21302800.949 | -21258734.181 | 504574.719 | -115150.169 | 46957.911 |
| -21115716.338 | -21905759.227 | -20977463.961 | -61514.751 | 493292.965 | 85503.952 |
| -21076050.527 | -21339177.306 | -21883340.167 | -57846.359 | -80168.042 | 633160.505 |
| 270.271 | 237.599 | 190.475 | 0.000 | 0.000 | 0.000 |
| 237.599 | 270.271 | 190.475 | 0.000 | 0.000 | 0.000 |
| 190.475 | 190.475 | 212.525 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 21.939 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 21.939 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -66.909 |
| 270.271 | 237.599 | 190.475 | 0.000 | -0.000 | -0.000 |
| 237.599 | 270.271 | 190.475 | -0.000 | -0.000 | -0.000 |
| 190.475 | 190.475 | 212.525 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 21.939 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 21.939 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -66.909 |
| 270.271 | 237.599 | 190.475 | 0.000 | 0.000 | 0.000 |
| 237.599 | 270.271 | 190.475 | -0.000 | 0.000 | 0.000 |
| 190.475 | 190.475 | 212.525 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 22.110 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 22.110 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -66.909 |
| 270.271 | 237.599 | 190.475 | -0.000 | -0.000 | -0.000 |
| 237.599 | 270.271 | 190.475 | -0.000 | -0.000 | -0.000 |
| 190.475 | 190.475 | 212.525 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 22.110 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 22.110 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -66.909 |
| 270.272 | 237.600 | 190.476 | 0.000 | 0.000 | 0.000 |
| 237.600 | 270.272 | 190.476 | 0.000 | 0.000 | 0.000 |
| 190.476 | 190.476 | 212.526 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 22.110 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 22.110 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -66.909 |
| 270.270 | 237.598 | 190.474 | -0.000 | -0.000 | -0.000 |
| 237.598 | 270.270 | 190.474 | -0.000 | -0.000 | -0.000 |
| 190.474 | 190.474 | 212.524 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 22.110 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 22.110 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -66.909 |
| 52.586 | 11.677 | 101.040 | 0.000 | -0.000 | -0.000 |
| 11.677 | 52.586 | 101.040 | 0.000 | -0.000 | 0.000 |
| 101.040 | 101.040 | 497.108 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -19.929 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | -19.929 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.002 |
| 52.586 | 11.677 | 101.040 | -0.000 | 0.000 | 0.000 |
| 11.677 | 52.586 | 101.040 | -0.000 | 0.000 | 0.000 |
| 101.040 | 101.040 | 497.108 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -19.929 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | -19.929 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.002 |
| 52.586 | 11.677 | 101.040 | 0.000 | -0.000 | 0.000 |
| 11.677 | 52.586 | 101.040 | 0.000 | -0.000 | -0.000 |
| 101.040 | 101.040 | 497.109 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -19.929 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -19.929 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.002 |
| 52.586 | 11.677 | 101.040 | 0.000 | 0.000 | 0.000 |
| 11.677 | 52.586 | 101.040 | -0.000 | 0.000 | 0.000 |
| 101.040 | 101.040 | 497.108 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -19.929 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | -19.929 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 0.002 |
| 55.609 | 8.655 | 101.041 | 0.000 | -0.000 | -0.000 |
| 8.655 | 55.609 | 101.041 | 0.000 | -0.000 | -0.000 |
| 101.041 | 101.041 | 497.110 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | -19.929 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -19.929 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.002 |
| -17198327.384 | -16272422.185 | -17074879.057 | -157091.402 | 17908.243 | 40040.351 |
| -16412300.232 | -16478532.895 | -18293214.322 | -48722.838 | -100724.393 | 15376.297 |
| 101.040 | 101.040 | 497.106 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -19.929 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | -19.929 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 0.002 |
| 3997778847.511 | 4180520954.936 | 4301669180.765 | 145959440.904 | -5765056.663 | -9609881.110 |
| 228.215 | 174.663 | 179.086 | 7.187 | 0.000 | -0.000 |
| -426.638 | -426.638 | 22.124 | -0.000 | -0.000 | -0.000 |
| -443.260 | -905.437 | -40.895 | -202.409 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 3.120 | -7.187 |
| 0.000 | 0.000 | 0.000 | -0.000 | -7.187 | -26.776 |
| -4075705839.602 | -4495663417.353 | -4605953569.305 | -53845270.424 | 743385.589 | -3535953.632 |
| 228.215 | 174.663 | 179.086 | 7.187 | -0.000 | 0.000 |
| 767.121 | 767.121 | 94.518 | 0.000 | -0.000 | 0.000 |
| -7.187 | 7.187 | -0.000 | 3.120 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 3.120 | -7.187 |
| -0.000 | -0.000 | -0.000 | 0.000 | -7.187 | -26.776 |
| -223.555 | -43.729 | 158.765 | 47.273 | 0.000 | -0.000 |
| 228.215 | 174.664 | 179.086 | 7.187 | 0.000 | -0.000 |
| 110.554 | 110.554 | 54.701 | -0.000 | 0.000 | 0.000 |
| -50.087 | -220.170 | -8.195 | -76.433 | 0.000 | -0.000 |
| 405089663.050 | 435019718.841 | 466266914.556 | -26693091.892 | 14916690.421 | -13624710.353 |
| -0.000 | 0.000 | -0.000 | -0.000 | -7.187 | -26.776 |
| -229.464 | 37.934 | 161.062 | 85.042 | -0.000 | 0.000 |
| 228.215 | 174.663 | 179.086 | 7.187 | 0.000 | -0.000 |
| 229.929 | 229.929 | 61.940 | 0.000 | -0.000 | -0.000 |
| -7.187 | 7.187 | -0.000 | 3.120 | -0.000 | 0.000 |
| -404709899.355 | -434703886.343 | -469031788.307 | 27492985.880 | 14542927.501 | -13518185.289 |
| 0.000 | -0.000 | 0.000 | 0.000 | -7.187 | -26.776 |
| 40286866.667 | 42022719.717 | 43396434.592 | 1913230.776 | -75522.535 | -46869.629 |
| 228.216 | 174.664 | 179.086 | 7.187 | 0.000 | 0.000 |
| 181.823 | 181.823 | 59.023 | -0.000 | 0.000 | -0.000 |
| 40846219.739 | 44759651.564 | 46263235.501 | 716510.824 | -52725.044 | -2793.556 |
| 39837009.609 | 43999458.890 | 46576081.806 | -3271190.760 | 1517449.948 | -1567483.795 |
| 39901001.069 | 43866318.153 | 46876318.702 | -3255999.846 | -1473770.814 | 1668686.307 |
| -40887583.774 | -44800299.832 | -46280871.895 | -737975.292 | 45872.489 | 27190.433 |
| -40428722.016 | -41851410.289 | -42767912.541 | -1763237.782 | 109245.118 | -38759.636 |
| 181.822 | 181.822 | 59.023 | -0.000 | -0.000 | 0.000 |
| -40422988.850 | -41939764.430 | -43195360.482 | -1757649.034 | 151782.324 | 17499.352 |
| -39939746.466 | -43922980.314 | -46705328.307 | 3234748.310 | 1386388.767 | -1660693.452 |
| -39926784.925 | -43850281.218 | -46569209.993 | 3209235.888 | -1589968.100 | 1536038.210 |
| 220.744 | 168.650 | 168.650 | -0.000 | 0.000 | 0.000 |
| 168.650 | 220.744 | 168.650 | -0.000 | 0.000 | -0.000 |
| 168.650 | 168.650 | 220.744 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -40.830 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -40.830 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -40.830 |
| 220.744 | 168.650 | 168.650 | 0.000 | -0.000 | -0.000 |
| 168.650 | 220.744 | 168.650 | 0.000 | -0.000 | 0.000 |
| 168.650 | 168.650 | 220.744 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -40.830 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -40.830 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -40.830 |
| 220.744 | 168.650 | 168.650 | -0.000 | 0.000 | -0.000 |
| 168.650 | 220.744 | 168.650 | -0.000 | 0.000 | 0.000 |
| 168.650 | 168.650 | 220.744 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -40.830 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -40.830 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -40.830 |
| 220.744 | 168.650 | 168.650 | -0.000 | -0.000 | 0.000 |
| 168.650 | 220.744 | 168.650 | 0.000 | -0.000 | 0.000 |
| 168.650 | 168.650 | 220.744 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -40.830 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -40.830 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -40.830 |
| 220.745 | 168.650 | 168.650 | 0.000 | 0.000 | 0.000 |
| 168.650 | 220.745 | 168.650 | -0.000 | 0.000 | -0.000 |
| 168.650 | 168.650 | 220.745 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -40.830 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -40.830 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -40.830 |
| 220.743 | 168.649 | 168.649 | -0.000 | -0.000 | -0.000 |
| 168.649 | 220.743 | 168.649 | -0.000 | -0.000 | -0.000 |
| 168.649 | 168.649 | 220.743 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -40.830 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -40.830 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -40.830 |
| 3.950 | 7.143 | 0.053 | 0.000 | 0.000 | 0.000 |
| 6.588 | 8.116 | 0.035 | 0.000 | -0.000 | 0.000 |
| 0.038 | 0.576 | 24.848 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.162 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.015 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 5.547 |
| 3.950 | 7.143 | 0.053 | -0.000 | 0.000 | -0.000 |
| 6.588 | 8.116 | 0.035 | -0.000 | 0.000 | -0.000 |
| 0.038 | 0.575 | 24.848 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.162 | -0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | -0.015 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 5.547 |
| 3.950 | 7.143 | 0.053 | -0.000 | -0.000 | 0.000 |
| 6.588 | 8.114 | 0.035 | -0.000 | -0.000 | 0.000 |
| 0.038 | 0.575 | 24.848 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.162 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.015 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 5.547 |
| 3.950 | 7.143 | 0.053 | -0.000 | -0.000 | -0.000 |
| 6.588 | 8.117 | 0.035 | -0.000 | -0.000 | -0.000 |
| 0.038 | 0.575 | 24.848 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.162 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -0.015 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 5.547 |
| 3.950 | 7.143 | 0.053 | 0.000 | 0.000 | 0.000 |
| 6.588 | 8.099 | 0.035 | 0.000 | 0.000 | 0.000 |
| 0.038 | 0.575 | 24.846 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.162 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.015 | 0.000 |
| 0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 5.547 |
| 3.950 | 7.143 | 0.053 | -0.000 | -0.000 | -0.000 |
| 6.588 | 8.133 | 0.036 | -0.000 | -0.000 | -0.000 |
| 0.038 | 0.575 | 24.850 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.162 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -0.015 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 5.547 |
| 36.836 | 3.855 | 66.872 | -0.000 | 0.000 | 0.000 |
| 3.855 | 36.836 | 66.872 | 0.000 | -0.000 | 0.000 |
| 66.872 | 66.872 | 509.236 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -19.232 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -19.232 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -0.004 |
| 36.836 | 3.855 | 66.872 | -0.000 | -0.000 | 0.000 |
| 3.855 | 36.836 | 66.872 | -0.000 | -0.000 | 0.000 |
| 66.872 | 66.872 | 509.236 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -19.232 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -19.232 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | -0.004 |
| 36.836 | 3.855 | 66.872 | 0.000 | 0.000 | -0.000 |
| 3.855 | 36.836 | 66.872 | 0.000 | 0.000 | -0.000 |
| 66.872 | 66.872 | 509.237 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -19.232 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -19.232 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.004 |
| 36.836 | 3.855 | 66.872 | 0.000 | -0.000 | -0.000 |
| 3.855 | 36.836 | 66.872 | -0.000 | -0.000 | -0.000 |
| 66.872 | 66.872 | 509.236 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -19.232 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -19.232 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -0.004 |
| 36.836 | 3.855 | 66.872 | 0.000 | 0.000 | 0.000 |
| 3.855 | 36.836 | 66.872 | 0.000 | 0.000 | 0.000 |
| 66.872 | 66.872 | 509.238 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -19.232 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -19.232 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.004 |
| 36.835 | 3.855 | 66.872 | -0.000 | -0.000 | -0.000 |
| 3.855 | 36.835 | 66.872 | 0.000 | -0.000 | -0.000 |
| 66.872 | 66.872 | 509.234 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -19.232 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -19.232 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.004 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.001 | -0.001 | -0.000 | -0.000 | 0.000 | 0.000 |
| -0.001 | -0.001 | -0.000 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 0.001 | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.001 | 0.001 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| -0.001 | -0.001 | -0.000 | -0.000 | 0.000 | 0.000 |
| -0.001 | -0.001 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 0.000 |
| 0.001 | 0.001 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.001 | 0.001 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 8.061 | 0.310 | 0.055 | 0.000 | 0.000 | 0.003 |
| 0.434 | 7.599 | 1.343 | 0.000 | 0.000 | 0.003 |
| -0.012 | 0.875 | 0.132 | 0.000 | 0.000 | 0.003 |
| 0.006 | -0.215 | -0.038 | 1.331 | 0.000 | 0.003 |
| 0.119 | 0.046 | 0.008 | -0.002 | 0.404 | 0.013 |
| -0.028 | 0.378 | 0.066 | -0.000 | 0.001 | 0.247 |
| 8.057 | 0.749 | 0.131 | -0.000 | -0.000 | -0.003 |
| 0.421 | 8.018 | 1.416 | -0.000 | -0.000 | -0.003 |
| -0.029 | 1.267 | 0.200 | -0.000 | -0.000 | -0.003 |
| -0.006 | 0.215 | 0.038 | 1.331 | -0.000 | -0.003 |
| -0.118 | -0.045 | -0.008 | 0.002 | 0.388 | -0.013 |
| 0.028 | -0.378 | -0.066 | 0.000 | -0.001 | 0.221 |
| 8.121 | 0.121 | 0.022 | 0.000 | 0.000 | 0.001 |
| -0.104 | 4.891 | 0.872 | -0.000 | 0.001 | 0.001 |
| -0.074 | 1.288 | 0.204 | -0.000 | 0.003 | 0.001 |
| 0.001 | -0.021 | -0.004 | 1.395 | 0.000 | 0.000 |
| 0.022 | -0.032 | -0.006 | 0.000 | 0.327 | 0.000 |
| -0.003 | 0.042 | 0.007 | -0.000 | 0.000 | 0.501 |
| 8.121 | 0.161 | 0.029 | -0.000 | -0.000 | -0.001 |
| -0.113 | 4.730 | 0.844 | 0.000 | -0.001 | -0.001 |
| -0.095 | 0.806 | 0.120 | 0.000 | -0.001 | -0.001 |
| 0.005 | 0.009 | 0.002 | 0.325 | -0.002 | -0.002 |
| -0.022 | 0.032 | 0.006 | -0.000 | 0.326 | -0.000 |
| 0.003 | -0.042 | -0.007 | 0.000 | -0.000 | 0.498 |
| 8.125 | 0.049 | 0.009 | -0.000 | 0.000 | 0.000 |
| -0.117 | 4.863 | 0.867 | -0.000 | 0.000 | 0.000 |
| -0.156 | 1.434 | 0.230 | 0.000 | 0.000 | 0.000 |
| -0.001 | -0.001 | -0.000 | 0.295 | 0.000 | 0.000 |
| 0.001 | -0.001 | -0.000 | -0.000 | 0.330 | 0.000 |
| -0.002 | 0.001 | 0.000 | -0.000 | 0.000 | 0.498 |
| 8.111 | 0.034 | 0.006 | 0.000 | -0.000 | -0.000 |
| -0.103 | 4.844 | 0.864 | 0.000 | -0.000 | -0.000 |
| -0.092 | 0.910 | 0.138 | 0.000 | -0.000 | -0.000 |
| -0.004 | 0.003 | 0.001 | 0.294 | -0.000 | -0.000 |
| -0.001 | 0.001 | 0.000 | 0.000 | 0.330 | -0.000 |
| 0.002 | -0.001 | -0.000 | 0.000 | -0.000 | 0.498 |
| 134.138 | 23.204 | 8.452 | 0.000 | -0.000 | 0.000 |
| 259.505 | -329.392 | 33.656 | 0.000 | -0.000 | 0.000 |
| 237.969 | -320.211 | 10.530 | 0.000 | -0.000 | 0.000 |
| 402.328 | -51.489 | 13.133 | 4.123 | -0.000 | -0.000 |
| 318.570 | -281.167 | 1.616 | 0.000 | -10.997 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 28.266 |
| 134.138 | 23.204 | 8.452 | -0.000 | 0.000 | -0.000 |
| -233.673 | 396.361 | -13.980 | -0.000 | 0.000 | -0.000 |
| -316.123 | 289.693 | 20.314 | 0.000 | 0.000 | -0.000 |
| -402.328 | 51.489 | -13.133 | 4.123 | -0.000 | 0.000 |
| -318.570 | 281.168 | -1.616 | -0.000 | -10.997 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 0.000 | 28.266 |
| 63.320 | -20.015 | 3.857 | -0.000 | -0.000 | 0.000 |
| -21.722 | 7.336 | 7.080 | 0.000 | -0.000 | 0.000 |
| 32.872 | -18.962 | 18.398 | 0.000 | -0.000 | -0.000 |
| 23.426 | -32.955 | -0.893 | 4.440 | -0.000 | -0.000 |
| -34.358 | -16.790 | -1.784 | 0.000 | -0.087 | 0.000 |
| 31.486 | -28.083 | 0.026 | 0.000 | -0.000 | 8.269 |
| 18.724 | 0.418 | 0.280 | -0.000 | 0.000 | -0.000 |
| 22.743 | 81.813 | 10.842 | -0.000 | 0.000 | 0.000 |
| -24.264 | 45.918 | 17.176 | -0.000 | 0.000 | -0.000 |
| -23.426 | 32.955 | 0.893 | 4.440 | 0.000 | -0.000 |
| 34.358 | 16.790 | 1.784 | -0.000 | -0.087 | -0.000 |
| -31.486 | 28.083 | -0.026 | -0.000 | 0.000 | 8.269 |
| 31.888 | 3.639 | 1.865 | -0.000 | 0.000 | 0.000 |
| 11.663 | 24.713 | 9.473 | 0.000 | 0.000 | 0.000 |
| 3.052 | 9.305 | 19.413 | -0.000 | -0.000 | 0.000 |
| 6.136 | 0.135 | 0.214 | 4.384 | -0.000 | -0.000 |
| 1.935 | -3.199 | 0.085 | 0.000 | -0.964 | 0.000 |
| -0.889 | -0.434 | -0.046 | 0.000 | -0.000 | 11.337 |
| 28.711 | 12.154 | 0.894 | -0.000 | 0.000 | -0.000 |
| 5.182 | 31.036 | 9.208 | -0.000 | 0.000 | 0.000 |
| -1.761 | 12.931 | 19.330 | 0.000 | 0.000 | -0.000 |
| -6.136 | -0.135 | -0.214 | 4.384 | -0.000 | -0.000 |
| -1.935 | 3.199 | -0.085 | -0.000 | -0.964 | 0.000 |
| 0.889 | 0.434 | 0.046 | -0.000 | 0.000 | 11.337 |
| 28.870 | 5.193 | 9.255 | -0.000 | 0.000 | -0.000 |
| 19.701 | 102.635 | 8.077 | -0.000 | 0.000 | -0.000 |
| 5.652 | 11.027 | 31.270 | -0.000 | 0.000 | -0.000 |
| -0.001 | 0.001 | -0.000 | 5.538 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 10.139 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 7.761 |
| 28.462 | 9.210 | 9.164 | 0.000 | -0.000 | -0.000 |
| -1.815 | 59.969 | 7.860 | 0.000 | -0.000 | -0.000 |
| 6.187 | 8.131 | 31.029 | 0.000 | 0.000 | -0.000 |
| 0.001 | -0.001 | 0.000 | 5.538 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 10.139 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 7.761 |
| 24.263 | 8.751 | 7.345 | -0.000 | 0.000 | 0.000 |
| 8.466 | 52.108 | 8.594 | 0.000 | -0.000 | -0.000 |
| 11.370 | 8.077 | 38.679 | -0.000 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 5.538 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 10.139 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 7.761 |
| 120.987 | 19.701 | 11.370 | -0.000 | 0.000 | 0.000 |
| 8.463 | 52.126 | 8.594 | -0.000 | -0.000 | -0.000 |
| 11.370 | 8.077 | 38.679 | -0.000 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | 5.538 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 10.139 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 7.761 |
| 24.254 | 8.750 | 7.344 | -0.000 | -0.000 | 0.000 |
| 5.865 | 53.024 | 8.540 | 0.000 | -0.000 | -0.000 |
| 5.823 | 8.534 | 30.251 | -0.000 | 0.000 | 0.000 |
| -0.005 | -0.003 | -0.000 | 4.577 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 6.650 | 0.000 |
| -0.020 | -0.047 | -0.000 | -0.000 | -0.000 | 6.742 |
| 28.454 | 9.209 | 9.164 | 0.000 | -0.000 | -0.000 |
| 5.869 | 53.036 | 8.540 | 0.000 | -0.000 | -0.000 |
| 11.371 | 8.077 | 38.682 | 0.000 | -0.000 | 0.000 |
| 0.005 | 0.003 | 0.000 | 4.577 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | 6.650 | -0.000 |
| 0.020 | 0.047 | 0.000 | -0.000 | 0.000 | 6.742 |
| 26.543 | 3.676 | 11.263 | 0.000 | 1.361 | 0.000 |
| -56.183 | 192.330 | 96.536 | -0.000 | 392.517 | -0.000 |
| -209.183 | 206.913 | 138.900 | -0.000 | 321.244 | -0.000 |
| 5.651 | -12.627 | -22.677 | -0.423 | 343.378 | 1.017 |
| -0.720 | 0.373 | -1.397 | 0.000 | 12.012 | 0.000 |
| -162.224 | 92.117 | 88.904 | -0.755 | 416.766 | 0.195 |
| 164.809 | -89.463 | -84.786 | 0.001 | -418.761 | 0.001 |
| 173.232 | -152.061 | -192.587 | 0.000 | -342.015 | 0.000 |
| 243.062 | 199.740 | 140.314 | 0.002 | -354.312 | 0.002 |
| -5.651 | 12.627 | 22.677 | -0.422 | -343.378 | 1.017 |
| 161.526 | -91.949 | -84.363 | 0.001 | -412.170 | 0.001 |
| 170.409 | 298.475 | 0.948 | -66.883 | -503.755 | 24.462 |
| -9.741 | 11.520 | 13.654 | -0.000 | 39.284 | -0.000 |
| -12.040 | -5.106 | 6.558 | -0.000 | 39.314 | -0.000 |
| 11.598 | 7.602 | 34.439 | 0.000 | -1.397 | 0.000 |
| -0.000 | -0.000 | -0.000 | 8.595 | 0.000 | 0.642 |
| -7.724 | -46.738 | 6.796 | -0.001 | 47.468 | -0.001 |
| -17.116 | -26.377 | -3.100 | 0.676 | 49.455 | 2.918 |
| 11.383 | 18.440 | -6.954 | 0.001 | -54.018 | 0.001 |
| -0.508 | 22.075 | -3.375 | 0.000 | -41.301 | 0.000 |
| 20.336 | -15.347 | -14.355 | 0.000 | -32.443 | 0.000 |
| 0.000 | 0.000 | 0.000 | 8.595 | -0.000 | 0.642 |
| 13.185 | -8.215 | -6.097 | 0.000 | -36.637 | 0.000 |
| 11.978 | 29.321 | 2.242 | -0.202 | -49.597 | -0.845 |
| 0.951 | 1.875 | 3.206 | -0.000 | 4.256 | -0.000 |
| -1.843 | 0.363 | -0.801 | -0.000 | 4.457 | -0.000 |
| -0.917 | -3.078 | 3.143 | -0.000 | 4.421 | -0.000 |
| -0.519 | -1.212 | -0.051 | 2.166 | 4.359 | 0.402 |
| -0.699 | 2.015 | 2.487 | -0.000 | 4.467 | -0.000 |
| -1.288 | 2.238 | 1.021 | -0.297 | 1.037 | 1.628 |
| 8.475 | 0.963 | 2.569 | 0.000 | 0.067 | 0.000 |
| 1.831 | 6.110 | -0.233 | 0.000 | -3.813 | 0.000 |
| 4.899 | -0.068 | 11.418 | 0.000 | -2.334 | 0.000 |
| 1.823 | -2.216 | -1.367 | 1.431 | -3.110 | -0.031 |
| 1.934 | -2.913 | -2.192 | 0.000 | -3.030 | 0.000 |
| 1.288 | -2.238 | -1.021 | -0.297 | -1.037 | 1.628 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 22.673 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 22.673 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 22.671 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 22.674 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 22.658 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 22.682 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | -0.000 |
| 2.841 | 2.854 | 6.694 | -0.000 | -0.000 | -0.000 |
| 2.854 | 2.841 | 6.694 | -0.000 | -0.000 | -0.000 |
| 5.867 | 5.867 | 8.673 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.056 | 4.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.056 | -0.000 | 4.000 | -0.000 |
| 0.000 | 0.000 | 0.001 | -0.000 | -0.000 | 2.776 |
| 2.841 | 2.854 | 6.692 | 0.000 | -0.000 | 0.000 |
| 2.854 | 2.841 | 6.692 | 0.000 | 0.000 | 0.000 |
| 5.867 | 5.867 | 8.672 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.056 | 4.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.056 | 0.000 | 4.000 | 0.000 |
| -0.000 | -0.000 | -0.001 | 0.000 | -0.000 | 2.776 |
| 2.841 | 2.854 | 6.693 | -0.000 | -0.000 | -0.000 |
| 2.854 | 2.841 | 6.693 | -0.000 | -0.000 | -0.000 |
| 5.867 | 5.867 | 8.670 | -0.000 | 0.000 | -0.000 |
| -2870011.149 | -2099293.491 | -9589846.024 | 137845.830 | -114735.713 | 142074.631 |
| -2178531.240 | -3025449.487 | -9710181.687 | -74389.443 | 60963.280 | 130544.071 |
| 0.000 | 0.000 | 0.000 | -0.000 | 0.000 | 2.776 |
| 2.841 | 2.854 | 6.693 | 0.000 | 0.000 | 0.000 |
| 2.854 | 2.841 | 6.693 | 0.000 | -0.000 | 0.000 |
| 5.867 | 5.867 | 8.675 | 0.000 | 0.000 | 0.000 |
| 2999982.137 | 2208995.167 | 9659553.865 | 149997.492 | -11527.622 | -35104.339 |
| 2200978.576 | 2918251.634 | 9598200.370 | 1615.825 | 153253.918 | -67147.424 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 2.776 |
| 2.841 | 2.854 | 6.693 | -0.000 | -0.000 | -0.000 |
| 2.854 | 2.841 | 6.693 | -0.000 | -0.000 | -0.000 |
| 5.867 | 5.867 | 8.649 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 4.260 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 4.260 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 2.776 |
| 2.841 | 2.854 | 6.693 | 0.000 | 0.000 | 0.000 |
| 2.854 | 2.841 | 6.693 | 0.000 | -0.000 | 0.000 |
| 5.867 | 5.867 | 8.700 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 4.260 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 4.260 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 2.776 |
| 2.295 | 2.278 | 1.483 | -0.000 | 0.000 | 0.000 |
| 2.278 | 2.295 | 1.483 | -0.000 | -0.000 | 0.000 |
| 2.398 | 2.398 | -3.037 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 2.698 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 2.698 | 0.000 |
| 0.000 | 0.000 | 0.001 | 0.000 | -0.000 | 2.279 |
| 2.295 | 2.278 | 1.481 | 0.000 | 0.000 | -0.000 |
| 2.278 | 2.295 | 1.481 | 0.000 | -0.000 | -0.000 |
| 2.398 | 2.398 | -3.038 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 2.698 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 2.698 | -0.000 |
| -0.000 | -0.000 | -0.001 | -0.000 | -0.000 | 2.279 |
| 2.295 | 2.278 | 1.482 | 0.000 | -0.000 | 0.000 |
| 2.278 | 2.295 | 1.482 | -0.000 | -0.000 | 0.000 |
| 2.398 | 2.398 | -3.039 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.007 | 2.506 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.007 | -0.000 | 2.506 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 2.279 |
| 2.295 | 2.278 | 1.482 | -0.000 | 0.000 | -0.000 |
| 2.278 | 2.295 | 1.482 | 0.000 | 0.000 | -0.000 |
| 2.398 | 2.398 | -3.035 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.007 | 2.506 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.007 | -0.000 | 2.506 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 2.279 |
| 2.295 | 2.278 | 1.482 | 0.000 | -0.000 | 0.000 |
| 2.278 | 2.295 | 1.482 | -0.000 | 0.000 | 0.000 |
| 2.398 | 2.398 | -3.058 | 0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.005 | 2.506 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.005 | -0.000 | 2.506 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 2.279 |
| 2.295 | 2.278 | 1.482 | 0.000 | -0.000 | 0.000 |
| 2.278 | 2.295 | 1.482 | 0.000 | 0.000 | 0.000 |
| 2.398 | 2.398 | -3.016 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.005 | 2.506 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.005 | 0.000 | 2.506 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 2.279 |
| 2.074 | 2.076 | 1.603 | 0.000 | 0.000 | -0.000 |
| 2.076 | 2.074 | 1.603 | 0.000 | 0.000 | -0.000 |
| 1.603 | 1.603 | 27.763 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.001 | 0.748 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.001 | 0.000 | 0.748 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 2.073 |
| 2.074 | 2.076 | 1.603 | -0.000 | -0.000 | 0.000 |
| 2.076 | 2.074 | 1.603 | -0.000 | -0.000 | 0.000 |
| 1.603 | 1.603 | 27.763 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.001 | 0.748 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.001 | -0.000 | 0.748 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 2.073 |
| 2.074 | 2.076 | 1.603 | 0.000 | 0.000 | -0.000 |
| 2.076 | 2.074 | 1.603 | 0.000 | 0.000 | -0.000 |
| 1.603 | 1.603 | 27.761 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.896 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.896 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 2.073 |
| 2.074 | 2.076 | 1.603 | -0.000 | -0.000 | 0.000 |
| 2.076 | 2.074 | 1.603 | -0.000 | -0.000 | 0.000 |
| 1.603 | 1.603 | 27.765 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.896 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 0.896 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 2.073 |
| 2.074 | 2.076 | 1.603 | 0.000 | 0.000 | -0.000 |
| 2.076 | 2.074 | 1.603 | 0.000 | 0.000 | -0.000 |
| 1.603 | 1.603 | 27.745 | 0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 0.896 | 0.000 | -0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 0.896 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 2.073 |
| 2.074 | 2.076 | 1.603 | -0.000 | -0.000 | 0.000 |
| 2.076 | 2.074 | 1.603 | -0.000 | -0.000 | 0.000 |
| 1.603 | 1.603 | 27.781 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 0.896 | -0.000 | 0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 0.896 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 2.073 |
| 7.877 | 63.799 | 8.081 | 0.002 | 0.000 | 0.000 |
| -82.402 | 93.564 | -36.159 | 0.446 | -0.000 | -0.000 |
| 3.123 | 83.341 | 8.754 | 0.003 | 0.000 | 0.000 |
| -85.474 | 85.733 | -41.398 | 1.965 | 0.000 | -0.000 |
| 0.069 | 60.167 | 4.904 | 0.001 | 3.604 | -0.000 |
| -66.185 | 66.271 | -32.065 | 0.677 | 0.000 | -0.200 |
| 39.129 | -30.846 | 18.099 | -0.102 | 0.000 | -0.000 |
| 86.491 | -76.623 | 45.579 | -0.436 | 0.000 | 0.000 |
| 29.302 | -14.863 | 15.616 | -0.084 | 0.000 | -0.000 |
| 24.476 | -23.763 | 11.918 | 4.221 | -0.000 | 0.000 |
| 66.194 | -66.282 | 32.069 | -0.678 | 1.894 | -0.000 |
| 66.185 | -66.271 | 32.065 | -0.677 | -0.000 | -0.200 |
| -3.301 | 10.632 | -2.539 | 0.026 | -0.000 | -0.000 |
| -1.032 | 13.392 | 3.368 | 0.005 | 0.000 | -0.000 |
| 3.250 | 5.561 | 2.537 | 0.000 | -0.000 | 0.000 |
| -50518.859 | -4787495.580 | -7595107.887 | -1628708.121 | -0.000 | -0.000 |
| 0.007 | 6.376 | 0.520 | 0.000 | 3.603 | -0.000 |
| -6.620 | 6.628 | -3.207 | 0.068 | 0.000 | -0.200 |
| 7.816 | -3.609 | 2.596 | -0.000 | -0.000 | -0.000 |
| 2.275 | 5.308 | 4.584 | -0.005 | -0.000 | -0.000 |
| 10.338 | -3.374 | 5.822 | -0.044 | 0.000 | 0.000 |
| -0.003 | -2.926 | -0.239 | 4.532 | -0.000 | 0.000 |
| -0.007 | -6.376 | -0.520 | -0.000 | 3.603 | -0.000 |
| 3.461 | -3.488 | 1.675 | -0.011 | -0.000 | -0.198 |
| 4.374 | 3.040 | 1.187 | 0.044 | -0.000 | -0.000 |
| 0.273 | 9.918 | 3.825 | 0.000 | 0.000 | -0.000 |
| 2.829 | 4.760 | 2.234 | 0.001 | -0.000 | 0.000 |
| -5051.814 | -478747.292 | -759517.279 | -162867.011 | -0.000 | -0.000 |
| -0.491 | 0.617 | -0.228 | 0.002 | 1.789 | -0.000 |
| 0.001 | 0.649 | 0.053 | 0.000 | -0.000 | -0.181 |
| 5.980 | 1.277 | 1.953 | -0.005 | -0.000 | 0.000 |
| 0.272 | 9.358 | 3.779 | -0.000 | -0.000 | 0.000 |
| 3.267 | 3.931 | 2.415 | -0.000 | -0.000 | 0.000 |
| 5051.814 | 478747.301 | 759517.305 | -162867.008 | -0.000 | 0.000 |
| 0.837 | -0.846 | 0.405 | -0.011 | 1.887 | -0.000 |
| 0.548 | -0.766 | 0.248 | -0.003 | -0.000 | -0.200 |
| 2.380 | 0.804 | 3.791 | -0.000 | -0.000 | -0.000 |
| 0.804 | 2.380 | 3.791 | 0.000 | -0.000 | -0.000 |
| 3.030 | 3.030 | 44.128 | -0.000 | -0.000 | -0.000 |
| 0.199 | 0.199 | 40.589 | 2.009 | -0.000 | -0.000 |
| 0.199 | 0.199 | 40.589 | -0.000 | 2.009 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 0.788 |
| 2.175 | 0.604 | -37.374 | 0.000 | 0.000 | 0.000 |
| 0.604 | 2.175 | -37.374 | 0.000 | 0.000 | 0.000 |
| 2.632 | 2.632 | -37.040 | 0.000 | 0.000 | 0.000 |
| -0.199 | -0.199 | -40.589 | 2.009 | 0.000 | 0.000 |
| -0.199 | -0.199 | -40.589 | 0.000 | 2.009 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 0.788 |
| 2.395 | 0.823 | 7.351 | -0.000 | -0.000 | -0.000 |
| 0.804 | 2.380 | 3.791 | -0.000 | -0.000 | -0.000 |
| 2.851 | 2.851 | 7.601 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | 2.010 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 2.010 | -0.000 |
| 0.020 | 0.020 | 4.123 | -0.000 | -0.000 | 0.786 |
| 2.354 | 0.783 | -0.898 | 0.000 | 0.000 | 0.000 |
| 0.943 | 2.194 | -0.887 | 0.000 | 0.000 | 0.000 |
| 3.405 | 3.405 | 120.424 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 2.010 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | 2.010 | 0.000 |
| -0.020 | -0.020 | -4.123 | 0.000 | 0.000 | 0.786 |
| 2.349 | 0.818 | 2.097 | -0.000 | -0.000 | -0.000 |
| 1.594 | 1.588 | 3.623 | -0.000 | -0.000 | -0.000 |
| 2.833 | 2.833 | 3.933 | -0.000 | -0.000 | -0.000 |
| 0.004 | 0.004 | 0.864 | 2.007 | -0.000 | -0.000 |
| 0.006 | 0.006 | 1.186 | -0.000 | 1.768 | -0.000 |
| 0.004 | 0.004 | 0.715 | -0.000 | -0.000 | 0.373 |
| 2.371 | 0.800 | 2.543 | 0.000 | 0.000 | 0.000 |
| 0.804 | 2.380 | 3.791 | 0.000 | 0.000 | 0.000 |
| 2.829 | 2.829 | 3.155 | 0.000 | 0.000 | 0.000 |
| -0.004 | -0.004 | -0.864 | 2.007 | 0.000 | 0.000 |
| -0.006 | -0.006 | -1.186 | 0.000 | 1.768 | 0.000 |
| -0.004 | -0.004 | -0.715 | 0.000 | 0.000 | 0.373 |
| 37.296 | 28.949 | 18.817 | -0.000 | 0.000 | 0.000 |
| 29.002 | 37.198 | 25.104 | -0.000 | 0.000 | 0.000 |
| -20.259 | 12.110 | 38.541 | -0.000 | 0.000 | 0.000 |
| -1.224 | 1.217 | 13.629 | -6.622 | 0.000 | 0.000 |
| -30.851 | 2.348 | 23.089 | -0.000 | 8.556 | 0.000 |
| -30.843 | 2.347 | 23.085 | -0.000 | -0.000 | 27.326 |
| 66.874 | 26.013 | -11.082 | 0.000 | -0.000 | -0.000 |
| 29.002 | 37.198 | 25.105 | -0.000 | -0.000 | 0.000 |
| 41.427 | 7.416 | -7.620 | 0.000 | -0.000 | -0.000 |
| 1.224 | -1.217 | -13.629 | -6.622 | -0.000 | 0.000 |
| 30.851 | -2.348 | -23.089 | 0.000 | 8.556 | -0.000 |
| 30.843 | -2.347 | -23.085 | 0.000 | -0.000 | 27.326 |
| 32.948 | 28.594 | 14.313 | 0.000 | 0.000 | 0.000 |
| 25.328 | 36.213 | 13.544 | -0.000 | 0.000 | -0.000 |
| 7.500 | 9.997 | 17.767 | -0.000 | 0.000 | -0.000 |
| -3.143 | 0.273 | 2.735 | 8.349 | 0.000 | -0.000 |
| -3.163 | 0.239 | 2.348 | -0.000 | 8.532 | 0.000 |
| -3.085 | 0.235 | 2.309 | -0.000 | 0.000 | 27.326 |
| 36.897 | 28.116 | 9.343 | -0.000 | -0.000 | 0.000 |
| 31.497 | 35.744 | 8.928 | 0.000 | -0.000 | 0.000 |
| 13.669 | 9.528 | 13.155 | -0.000 | -0.000 | -0.000 |
| 3.143 | -0.273 | -2.735 | 8.349 | -0.000 | -0.000 |
| 3.163 | -0.239 | -2.348 | 0.000 | 8.532 | 0.000 |
| 3.085 | -0.235 | -2.309 | 0.000 | -0.000 | 27.326 |
| 35.723 | 28.383 | 12.236 | -0.000 | 0.000 | -0.000 |
| 28.103 | 36.001 | 11.467 | -0.000 | 0.000 | -0.000 |
| 10.275 | 9.785 | 15.671 | -0.000 | 0.000 | -0.000 |
| -0.290 | 0.022 | 0.211 | 7.961 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 9.501 | 0.000 |
| -0.310 | 0.023 | 0.231 | -0.000 | -0.000 | 27.325 |
| 36.341 | 28.337 | 11.775 | -0.000 | -0.000 | -0.000 |
| 28.722 | 35.956 | 11.006 | -0.000 | -0.000 | 0.000 |
| 10.893 | 9.739 | 15.230 | 0.000 | -0.000 | -0.000 |
| 0.290 | -0.022 | -0.211 | 7.961 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 9.501 | 0.000 |
| 0.310 | -0.023 | -0.231 | 0.000 | -0.000 | 27.325 |
| 90.130 | 69.976 | 69.976 | -0.000 | -0.000 | -0.000 |
| 69.976 | 90.130 | 69.976 | -0.000 | -0.000 | -0.000 |
| 69.976 | 69.976 | 90.130 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 34.874 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 34.874 | -0.000 |
| -0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 34.874 |
| 746.053 | 737.157 | 737.157 | -0.000 | 0.001 | 0.000 |
| 737.157 | 746.052 | 737.156 | -0.000 | 0.001 | 0.000 |
| 737.157 | 737.156 | 746.053 | -0.000 | 0.001 | 0.000 |
| 0.000 | -0.000 | -0.000 | 34.874 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 34.874 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -0.000 | 34.874 |
| 90.130 | 69.976 | 69.976 | -0.000 | -0.000 | -0.000 |
| 69.976 | 90.130 | 69.976 | -0.000 | 0.000 | -0.000 |
| 69.976 | 69.976 | 90.130 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 34.874 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 34.874 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 34.874 |
| -96.471 | -98.026 | -98.026 | 0.000 | 0.002 | 0.000 |
| -98.026 | -96.471 | -98.026 | 0.000 | 0.002 | 0.000 |
| -98.026 | -98.026 | -96.471 | 0.000 | 0.002 | 0.000 |
| 0.000 | 0.000 | -0.000 | 34.874 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | 34.874 | 0.000 |
| 0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 34.874 |
| 90.130 | 69.977 | 69.977 | -0.000 | -0.000 | -0.000 |
| 69.977 | 90.130 | 69.977 | 0.000 | 0.000 | -0.000 |
| 69.977 | 69.977 | 90.130 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 34.874 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | 34.874 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 34.874 |
| 28.718 | 30.559 | 30.559 | 0.000 | 0.000 | 0.000 |
| 30.559 | 28.718 | 30.559 | 0.000 | 0.000 | 0.000 |
| 30.559 | 30.559 | 28.718 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 34.874 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 34.874 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | 34.874 |
| -18.729 | -29.241 | 9.680 | 0.000 | 0.000 | -0.000 |
| -33.452 | 22.502 | 25.838 | 0.000 | -0.000 | -0.000 |
| 3.917 | 20.950 | 16.799 | 0.000 | -0.000 | -0.000 |
| -0.006 | -0.112 | -0.188 | 19.959 | 0.000 | -0.000 |
| 141680969.052 | 117275003.536 | 7480688.939 | 0.574 | -35118615.942 | 0.336 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -14.437 |
| -7.115 | -7.960 | 2.918 | 0.000 | 0.000 | 0.000 |
| -34.216 | 21.169 | 26.408 | -0.000 | 0.000 | 0.000 |
| 3.930 | 21.193 | 17.204 | -0.000 | -0.000 | -0.000 |
| 0.006 | 0.112 | 0.188 | 19.959 | -0.000 | -0.000 |
| -141680969.028 | -117275003.404 | -7480688.429 | -0.318 | -35118615.856 | 0.355 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -14.437 |
| -19.071 | -29.839 | 9.936 | 0.000 | 0.000 | -0.000 |
| -33.794 | 21.903 | 26.094 | -0.000 | -0.000 | -0.000 |
| 6.823 | 26.935 | 16.348 | 0.000 | -0.000 | -0.000 |
| -0.001 | -0.011 | -0.019 | 19.960 | -0.000 | -0.000 |
| -0.039 | -0.787 | -1.320 | -0.000 | -15.763 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | -14.437 |
| -19.151 | -29.976 | 9.993 | -0.000 | -0.000 | 0.000 |
| -33.874 | 21.768 | 26.152 | -0.000 | -0.000 | -0.000 |
| 6.747 | 26.803 | 16.406 | -0.000 | 0.000 | 0.000 |
| 0.001 | 0.011 | 0.019 | 19.960 | 0.000 | -0.000 |
| 0.039 | 0.787 | 1.320 | 0.000 | -15.763 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | -14.437 |
| -19.085 | -29.881 | 9.962 | 0.000 | 0.000 | -0.000 |
| -33.813 | 21.854 | 26.112 | 0.000 | 0.000 | 0.000 |
| 6.788 | 26.871 | 16.369 | 0.000 | -0.000 | -0.000 |
| 0.000 | -0.001 | -0.002 | 19.960 | 0.000 | -0.000 |
| 1416810.444 | 1172750.369 | 74805.231 | 0.016 | -351190.937 | -0.006 |
| 1270997.250 | 1114984.520 | 76681.426 | -0.008 | -0.001 | -129822.386 |
| -19.138 | -29.934 | 9.968 | -0.000 | -0.000 | 0.000 |
| -33.855 | 21.817 | 26.133 | -0.000 | -0.000 | 0.000 |
| 6.781 | 26.867 | 16.386 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.001 | 0.002 | 19.960 | -0.000 | 0.000 |
| -1416810.442 | -1172750.371 | -74805.246 | -0.014 | -351190.934 | -0.006 |
| -1270997.250 | -1114984.520 | -76681.426 | 0.008 | -0.001 | -129822.386 |
| 85.634 | 46.382 | 168.018 | -0.000 | -0.000 | 0.000 |
| 46.382 | 85.634 | 168.018 | -0.000 | -0.000 | 0.000 |
| 168.018 | 168.018 | 447.049 | -0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -43.970 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -43.970 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 19.626 |
| 85.634 | 46.382 | 168.018 | 0.000 | 0.000 | -0.000 |
| 46.382 | 85.634 | 168.018 | 0.000 | 0.000 | 0.000 |
| 168.018 | 168.018 | 447.049 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -43.970 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -43.970 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 19.626 |
| 85.634 | 46.382 | 168.018 | -0.000 | -0.000 | -0.000 |
| 46.382 | 85.634 | 168.018 | -0.000 | -0.000 | -0.000 |
| 168.018 | 168.018 | 447.049 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -43.970 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -43.970 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 19.626 |
| 85.634 | 46.382 | 168.018 | 0.000 | 0.000 | 0.000 |
| 46.382 | 85.634 | 168.018 | 0.000 | -0.000 | 0.000 |
| 168.018 | 168.018 | 447.048 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -43.970 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -43.970 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 19.626 |
| 85.635 | 46.382 | 168.019 | -0.000 | -0.000 | -0.000 |
| 46.382 | 85.635 | 168.019 | -0.000 | -0.000 | 0.000 |
| 168.018 | 168.018 | 447.051 | -0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -43.970 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -43.970 | 0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 19.626 |
| 85.634 | 46.381 | 168.018 | 0.000 | 0.000 | 0.000 |
| 46.381 | 85.634 | 168.018 | 0.000 | 0.000 | -0.000 |
| 168.018 | 168.018 | 447.046 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | -43.970 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -43.970 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 19.626 |
| 3401101324.621 | 3565262322.371 | 4185873244.342 | 910795.162 | 917128.829 | 15228562.692 |
| 222.079 | 196.744 | 120.696 | -0.000 | -0.000 | 0.000 |
| 120.696 | 120.696 | 140.819 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -5.820 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -5.820 | 0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 0.000 | -305.999 |
| 196.744 | 222.079 | 120.696 | -0.000 | 0.000 | -0.000 |
| 222.079 | 196.744 | 120.696 | -0.000 | -0.000 | -0.000 |
| 120.696 | 120.696 | 140.819 | -0.000 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -5.820 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -5.820 | -0.000 |
| -0.000 | 0.000 | -0.000 | -0.000 | 0.000 | -305.999 |
| 196.744 | 222.079 | 120.696 | 0.000 | -0.000 | 0.000 |
| 222.079 | 196.744 | 120.696 | 0.000 | -0.000 | 0.000 |
| 120.696 | 120.696 | 140.819 | 0.000 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -5.820 | -0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | 0.000 | -5.820 | 0.000 |
| 395722196.910 | 350446220.614 | 400399194.819 | -412519.305 | -515133.806 | 22755025.285 |
| -309047524.206 | -516124471.428 | -526334473.466 | 0.035 | -0.000 | -0.001 |
| -341689386.541 | -356803555.519 | -415822129.510 | 437602.571 | 134381.811 | -1915306.189 |
| 120.696 | 120.696 | 140.819 | -0.000 | -0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | -5.820 | 0.000 | -0.000 |
| -0.000 | -0.000 | 0.000 | 0.000 | -5.820 | -0.000 |
| -395213419.910 | -352129691.082 | -399365369.415 | 381309.609 | -474961.285 | 21103695.251 |
| 34191506.719 | 35704456.119 | 41148949.324 | -31445.633 | -11276.590 | 121201.101 |
| 30904937.722 | 51612756.174 | 52633656.995 | 0.003 | -0.000 | 0.000 |
| 120.696 | 120.696 | 140.820 | 0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | -5.820 | 0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | -5.820 | 0.000 |
| 46435550.983 | 36081749.339 | 52633478.547 | 0.000 | -0.000 | 8966719.387 |
| -30904657.583 | -51612271.216 | -52633304.872 | -0.003 | 0.000 | -0.000 |
| -34035589.352 | -35880424.187 | -41143964.369 | 17815.818 | 36628.404 | -60396.706 |
| 120.696 | 120.696 | 140.818 | -0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | -5.820 | -0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | -5.820 | -0.000 |
| -46435550.984 | -36081749.340 | -52633478.547 | -0.000 | -0.000 | 8966719.388 |
| 2.197 | 2.257 | 3.821 | -0.000 | 0.000 | -0.000 |
| 2.257 | 2.197 | 3.821 | 0.000 | 0.000 | -0.000 |
| 1.606 | 1.606 | 59.797 | 0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -1.507 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -1.507 | -0.000 |
| 0.000 | 0.000 | 0.011 | -0.000 | -0.000 | 2.198 |
| 2.197 | 2.256 | 3.799 | 0.000 | -0.000 | 0.000 |
| 2.256 | 2.197 | 3.799 | -0.000 | 0.000 | 0.000 |
| 1.606 | 1.606 | 59.775 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -1.507 | -0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -1.507 | 0.000 |
| -0.000 | -0.000 | -0.011 | 0.000 | 0.000 | 2.198 |
| 2.197 | 2.257 | 3.811 | -0.000 | -0.000 | -0.000 |
| 2.257 | 2.197 | 3.811 | -0.000 | -0.000 | -0.000 |
| 1.606 | 1.606 | 59.785 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -1.507 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -1.507 | -0.000 |
| 0.000 | 0.000 | 0.001 | 0.000 | -0.000 | 2.198 |
| 2.197 | 2.257 | 3.809 | 0.000 | 0.000 | -0.000 |
| 2.257 | 2.197 | 3.809 | 0.000 | 0.000 | -0.000 |
| 1.606 | 1.606 | 59.787 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -1.507 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -1.507 | 0.000 |
| -0.000 | -0.000 | -0.001 | -0.000 | 0.000 | 2.198 |
| 2.197 | 2.257 | 3.810 | -0.000 | -0.000 | 0.000 |
| 2.257 | 2.197 | 3.810 | -0.000 | -0.000 | 0.000 |
| 1.606 | 1.606 | 59.768 | -0.000 | -0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -1.507 | 0.000 | -0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | -1.507 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | -0.000 | 2.198 |
| 2.197 | 2.257 | 3.810 | -0.000 | 0.000 | 0.000 |
| 2.257 | 2.197 | 3.810 | 0.000 | -0.000 | 0.000 |
| 1.606 | 1.606 | 59.803 | 0.000 | 0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -1.507 | -0.000 | 0.000 |
| -0.000 | -0.000 | 0.000 | -0.000 | -1.507 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 2.198 |
| 306.493 | 282.596 | 273.683 | 0.000 | -0.000 | 0.000 |
| 282.596 | 238.745 | 282.606 | 0.000 | -0.000 | 0.000 |
| 273.682 | 282.604 | 306.491 | 0.000 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 33.488 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 41.614 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 33.489 |
| 306.493 | 282.596 | 273.683 | -0.000 | 0.000 | -0.000 |
| 282.596 | 238.745 | 282.606 | -0.000 | 0.000 | -0.000 |
| 273.682 | 282.604 | 306.491 | -0.000 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 33.488 | 0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | -0.000 | 41.614 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 33.489 |
| 306.493 | 282.596 | 273.683 | -0.000 | -0.000 | -0.000 |
| 282.596 | 238.745 | 282.606 | -0.000 | -0.000 | 0.000 |
| 273.682 | 282.605 | 306.491 | -0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 33.488 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 41.614 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 33.489 |
| 306.492 | 282.596 | 273.683 | 0.000 | 0.000 | 0.000 |
| 282.596 | 238.745 | 282.606 | 0.000 | 0.000 | -0.000 |
| 273.682 | 282.604 | 306.491 | 0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 33.488 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | 0.000 | 41.614 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | 0.000 | 33.489 |
| 306.493 | 282.597 | 273.684 | -0.000 | -0.000 | 0.000 |
| 282.597 | 238.745 | 282.607 | -0.000 | -0.000 | 0.000 |
| 273.683 | 282.605 | 306.492 | 0.000 | -0.000 | 0.000 |
| 0.000 | 0.000 | 0.000 | 33.488 | -0.000 | 0.000 |
| -0.000 | 0.000 | -0.000 | 0.000 | 41.614 | 0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 33.489 |
| 306.492 | 282.595 | 273.682 | 0.000 | 0.000 | -0.000 |
| 282.595 | 238.745 | 282.605 | 0.000 | 0.000 | -0.000 |
| 273.681 | 282.604 | 306.490 | -0.000 | 0.000 | -0.000 |
| -0.000 | -0.000 | -0.000 | 33.488 | 0.000 | -0.000 |
| 0.000 | -0.000 | 0.000 | -0.000 | 41.614 | -0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 33.489 |
| 371.563 | 244.511 | 244.510 | -0.000 | 0.000 | -0.000 |
| 244.518 | 355.493 | 260.531 | -0.000 | 0.000 | -0.000 |
| 244.518 | 260.531 | 355.492 | -0.000 | -0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 63.463 | 0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 0.000 | 47.482 | -0.000 |
| -0.000 | -0.000 | -0.000 | 0.000 | -0.000 | 47.481 |
| 371.563 | 244.511 | 244.510 | -0.000 | -0.000 | 0.000 |
| 244.518 | 355.493 | 260.531 | -0.000 | -0.000 | 0.000 |
| 244.518 | 260.531 | 355.492 | -0.000 | 0.000 | -0.000 |
| 0.000 | -0.000 | -0.000 | 63.463 | 0.000 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 47.482 | 0.000 |
| 0.000 | 0.000 | -0.000 | -0.000 | 0.000 | 47.481 |
| 371.563 | 244.511 | 244.510 | -0.000 | 0.000 | -0.000 |
| 244.518 | 355.493 | 260.531 | 0.000 | -0.000 | -0.000 |
| 244.518 | 260.531 | 355.492 | 0.000 | -0.000 | -0.000 |
| -0.000 | 0.000 | 0.000 | 63.463 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 47.482 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 47.481 |
| 371.563 | 244.511 | 244.510 | -0.000 | 0.000 | 0.000 |
| 244.518 | 355.493 | 260.531 | 0.000 | -0.000 | 0.000 |
| 244.517 | 260.531 | 355.492 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 63.463 | 0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 47.482 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | -0.000 | 47.481 |
| 371.565 | 244.511 | 244.510 | -0.000 | 0.000 | -0.000 |
| 244.519 | 355.494 | 260.531 | -0.000 | -0.000 | -0.000 |
| 244.518 | 260.532 | 355.493 | 0.000 | 0.000 | 0.000 |
| -0.000 | 0.000 | 0.000 | 63.463 | -0.000 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | 47.482 | -0.000 |
| 0.000 | 0.000 | 0.000 | 0.000 | -0.000 | 47.481 |
| 371.562 | 244.510 | 244.509 | -0.000 | 0.000 | 0.000 |
| 244.517 | 355.492 | 260.530 | 0.000 | 0.000 | 0.000 |
| 244.517 | 260.530 | 355.491 | -0.000 | 0.000 | 0.000 |
| 0.000 | -0.000 | -0.000 | 63.463 | -0.000 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 47.482 | 0.000 |
| -0.000 | -0.000 | -0.000 | -0.000 | 0.000 | 47.481 |
Free Surface Formation Energy Predictions
Static free surface formation energies are displayed for select crystals. The values displayed here are obtained by taking a perfect periodic bulk crystal, slicing along a crystallographic plane, and using force minimization to statically relax the surfaces. The free surface formation energy is computed by comparing the energy of the defect system to the bulk system and dividing by the total surface area created by the cut.
The calculation method used is available as the iprPy surface_energy_static calculation method.
Notes and Disclaimers:
- These values are meant to be guidelines for comparing potentials, not the absolute values for any potential's properties. Values listed here may change if the calculation methods are updated due to improvements/corrections. Variations in the values may occur for variations in calculation methods, simulation software and implementations of the interatomic potentials.
- The calculation only performs straight cuts along crystallographic planes and static relaxations. Lower energy configurations may exist that require atomic restructuring of the surfaces.
- Multiple values may be listed for a given plane followed by a number indicating different unique atomic planar cuts for the same theoretical plane.
- NIST disclaimer
Version Information:
- 2019-11-14. Calculations for the alternate #2 plane slices are now different from the #1 plane slices.
- 2019-08-07. Data added.
| Surface | γfs (mJ/m2) |
|---|---|
| (111) | 12711.14 |
| (332) | 14058.38 |
| (322) | 14245.92 |
| (221) | 14651.01 |
| (100) | 14898.72 |
| (211) | 15000.38 |
| (331) | 15129.05 |
| (311) | 15544.63 |
| (110) | 15607.29 |
| (321) | 15711.49 |
| (320) | 16371.62 |
| (310) | 16382.63 |
| (210) | 16539.96 |
| Surface | γfs (mJ/m2) |
|---|---|
| (0001) | 12805.41 |
| (1010) 1 | 13556.99 |
| (1011) 2 | 14104.05 |
| (2021) 1 | 14251.20 |
| (2130) 1 | 15355.34 |
| (1012) 2 | 15508.62 |
| (1012) 1 | 15600.61 |
| (2112) | 15652.94 |
| (1120) | 15685.54 |
| (2131) 1 | 15979.43 |
| (2132) 1 | 15979.76 |
| (2132) 2 | 15984.93 |
| (2131) 2 | 16026.03 |
| (2241) | 16211.50 |
| (1121) | 16374.24 |
| (2021) 2 | 16722.45 |
| (2130) 2 | 17333.58 |
| (1011) 1 | 18188.08 |
| (1010) 2 | 18652.35 |
| Surface | γfs (mJ/m2) |
|---|---|
| (331) 1 | 2379.36 |
| (221) | 3003.59 |
| (111) 1 | 3240.85 |
| (322) | 3250.77 |
| (332) | 3668.78 |
| (310) | 3704.50 |
| (311) 2 | 3784.31 |
| (210) | 4282.43 |
| (321) | 4717.14 |
| (111) 2 | 4783.06 |
| (320) | 4822.80 |
| (100) | 5089.23 |
| (211) | 5326.02 |
| (311) 1 | 5393.58 |
| (331) 2 | 5621.17 |
| (110) | 5766.47 |
| Surface | γfs (mJ/m2) |
|---|---|
| (332) | -113795.78 |
| (331) | -105509.40 |
| (322) | -100777.48 |
| (321) | -91824.10 |
| (320) | -87038.54 |
| (311) | -80726.69 |
| (310) | -76186.55 |
| (221) | -72683.29 |
| (211) | -59430.23 |
| (210) | -53452.92 |
| (111) | 421.64 |
| (100) | 527.44 |
| (110) | 580.17 |
| Surface | γfs (mJ/m2) |
|---|---|
| (331) | -82185.85 |
| (322) | -78144.15 |
| (321) | -71291.15 |
| (320) | -69017.52 |
| (311) | -65035.39 |
| (310) | -60330.42 |
| (221) | -57383.09 |
| (211) | -46476.29 |
| (210) | -42989.12 |
| (110) | 492.97 |
| (332) | 598.91 |
| (111) | 609.55 |
| (100) | 667.58 |
| Surface | γfs (mJ/m2) |
|---|---|
| (2112) | -43681.74 |
| (2132) 2 | -42564.08 |
| (1012) 2 | -41633.82 |
| (2132) 1 | -41230.58 |
| (1012) 1 | -39881.23 |
| (2021) 2 | -28960.06 |
| (2021) 1 | -28038.10 |
| (1011) 1 | -26369.39 |
| (1011) 2 | -25953.36 |
| (2241) | -17192.26 |
| (2130) 1 | -17103.97 |
| (2131) 1 | -16825.50 |
| (2131) 2 | -16816.21 |
| (1121) | -15998.11 |
| (2130) 2 | -14727.34 |
| (0001) | 421.08 |
| (1010) 1 | 506.04 |
| (1120) | 548.83 |
| (1010) 2 | 726.22 |
Stacking Fault Energy Predictions
Stacking fault energy plots and maps are displayed for select crystals. The values are computed by
- Starting with a bulk crystal system
- Creating a free surface along one of the system's periodic boundaries and using force minimization to relax it
- The system is sliced in half along a crystallographic plane parallel to the free surface. One half of the system is shifted relative to the other
- The atoms in the shifted system are allowed to relax only in the direction normal to the shifting plane
- The stacking fault energy for a given shift is computed by comparing the energy of the system before and after applying the shift, and dividing by the area of the fault plane
The calculation method used is available as the iprPy stacking_fault_map_2D calculation method.
Notes and Disclaimers:
- These values are meant to be guidelines for comparing potentials, not the absolute values for any potential's properties. Values listed here may change if the calculation methods are updated due to improvements/corrections. Variations in the values may occur for variations in calculation methods, simulation software and implementations of the interatomic potentials.
- Values between the measured points are interpolated and therefore may not perfectly capture minima and maxima.
- Multiple values may be listed for a given plane followed by a number indicating different unique atomic planar cuts for the same theoretical plane.
- NIST disclaimer
Version Information:
- 2022-06-28. Table added for intrinsic and unstable energies, and ideal shear stresses. Plots (and table) now ordered by the associated planes.
- 2019-11-14. Calculations for the alternate #2 plane slices are now different from the #1 plane slices. The 1D plotting vectors have been changed to avoid duplicates. Minor improvements to how the 2D plots are displayed.
- 2019-08-07. Plots added.
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/2 [0 -1 1] | 13954.21 |
| τ_ideal a/2 [0 -1 1] | 619.18 |
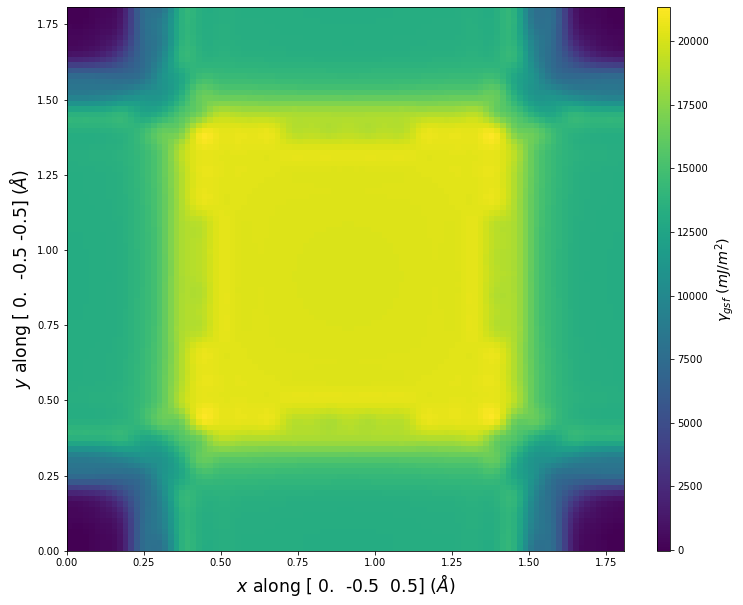
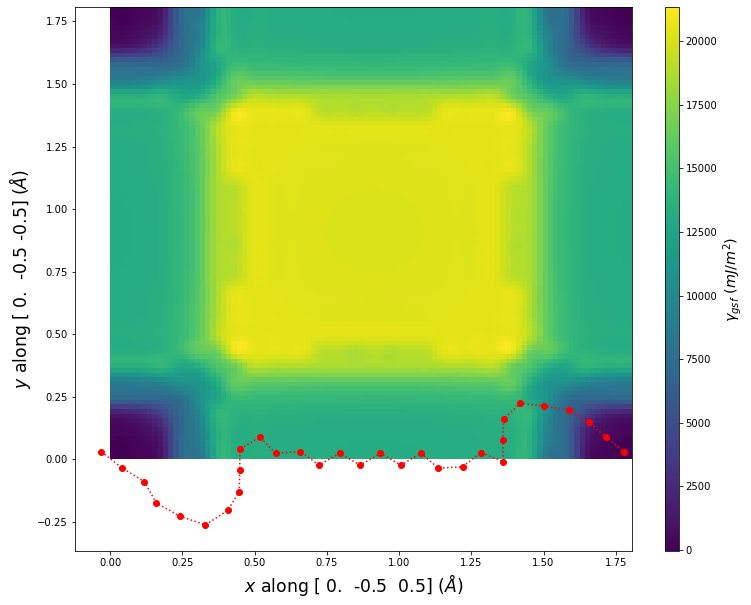
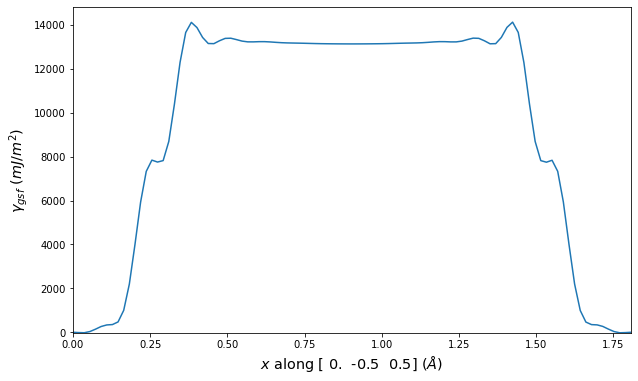
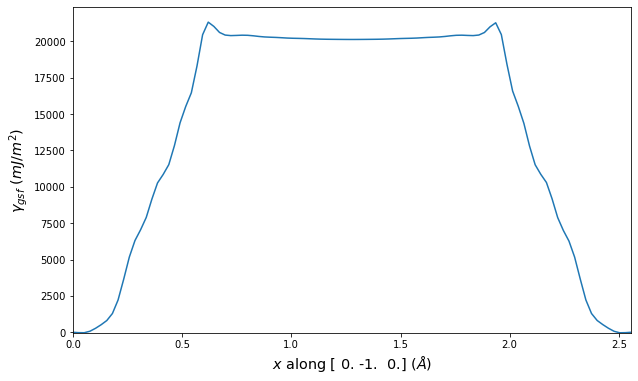
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_isf | -256.19 |
| E_usf a/2 [0 -1 1] | 7709.35 |
| τ_ideal a/2 [0 -1 1] | 740.72 |
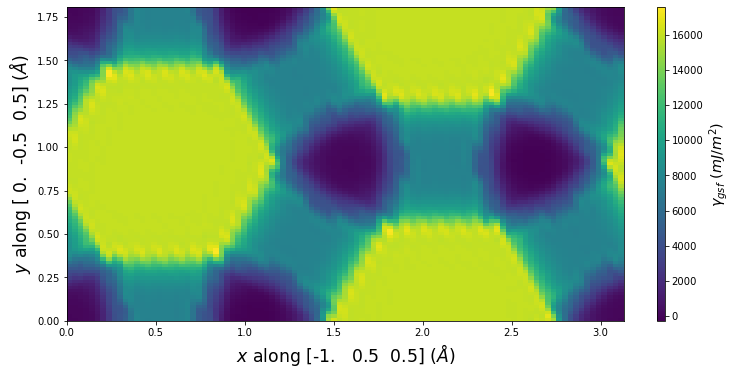
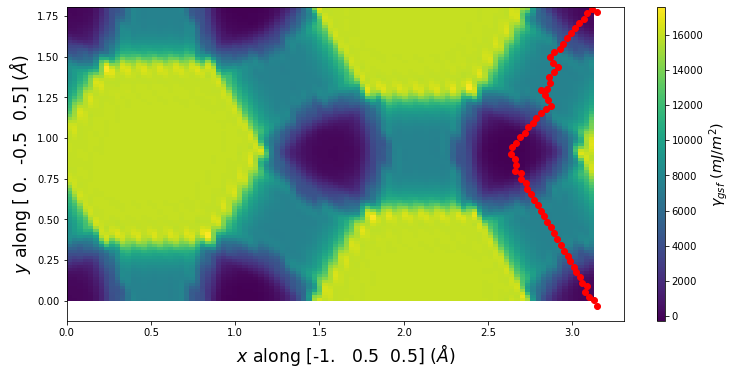
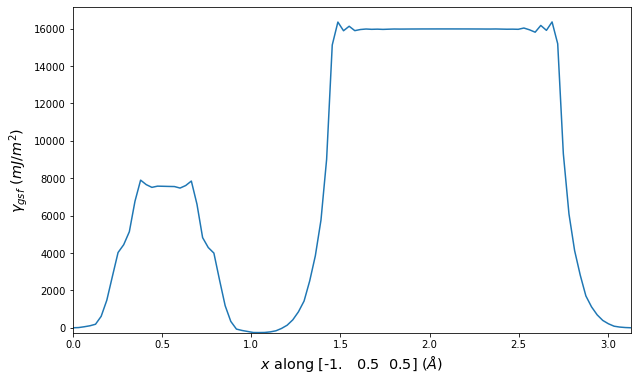
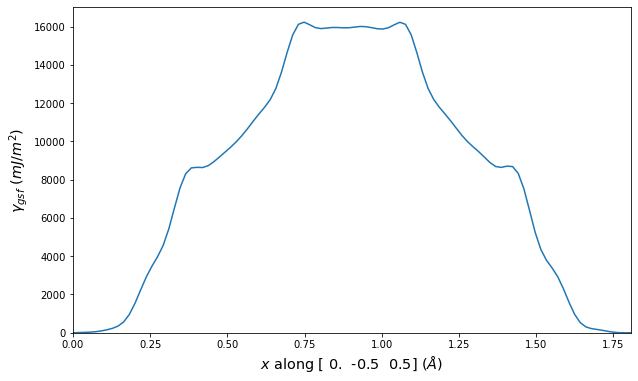
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_isf | 265.44 |
| E_usf a/3 [-2 1 1 0] | 7913.71 |
| τ_ideal a/3 [-2 1 1 0] | 711.95 |
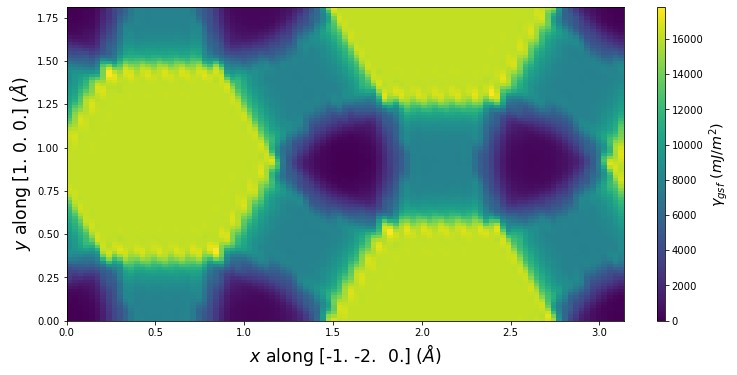
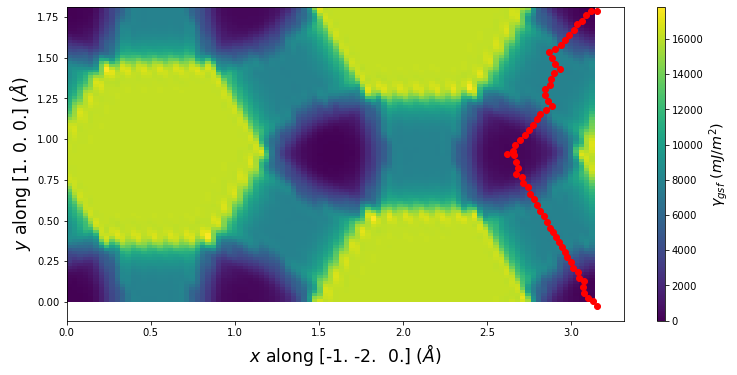
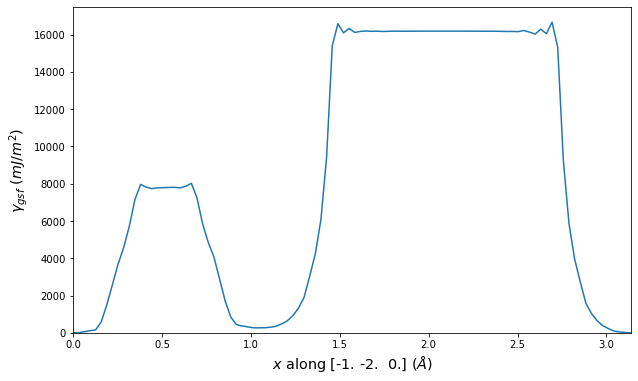
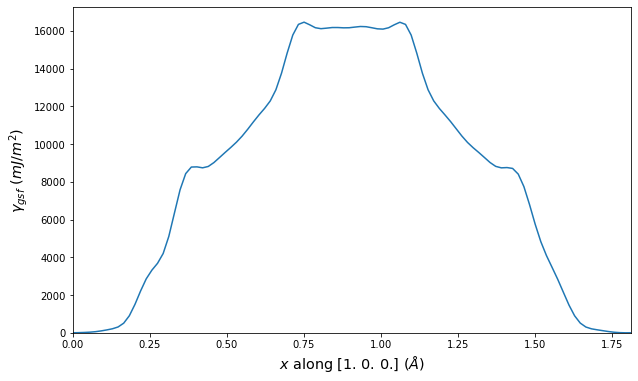
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-1 2 -1 0] | 10137.47 |
| τ_ideal a/3 [-1 2 -1 0] | 542.26 |
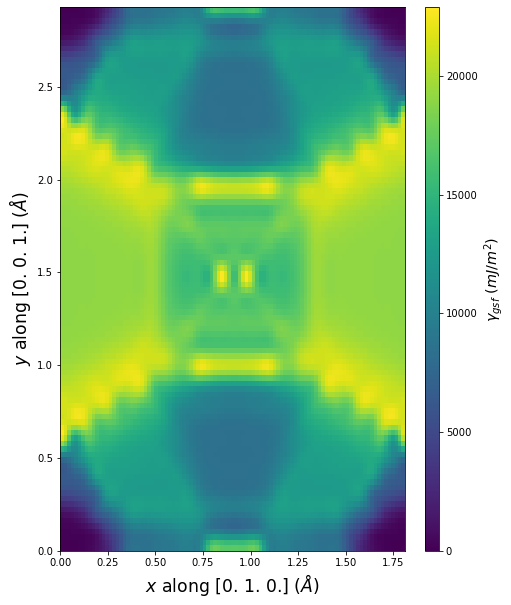
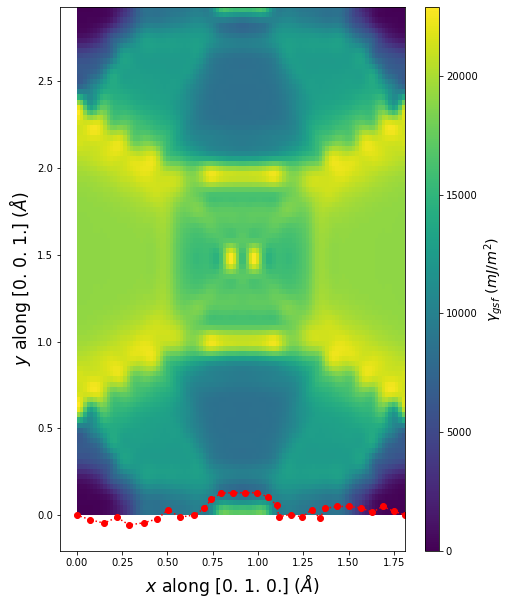
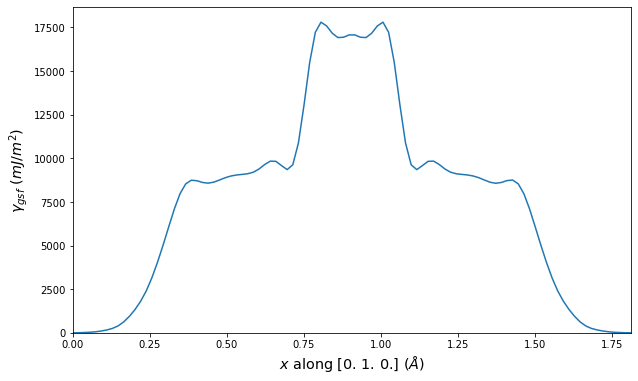
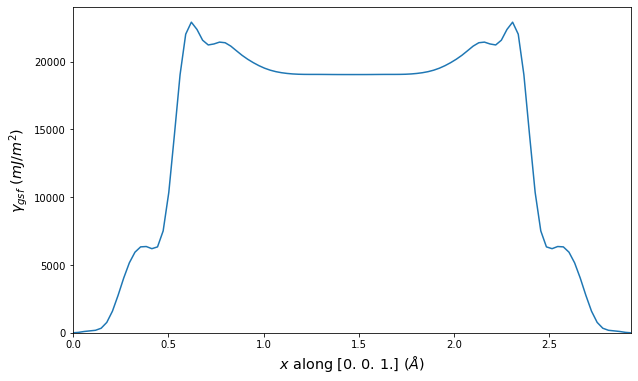
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-1 2 -1 0] | 16967.48 |
| τ_ideal a/3 [-1 2 -1 0] | 1300.94 |
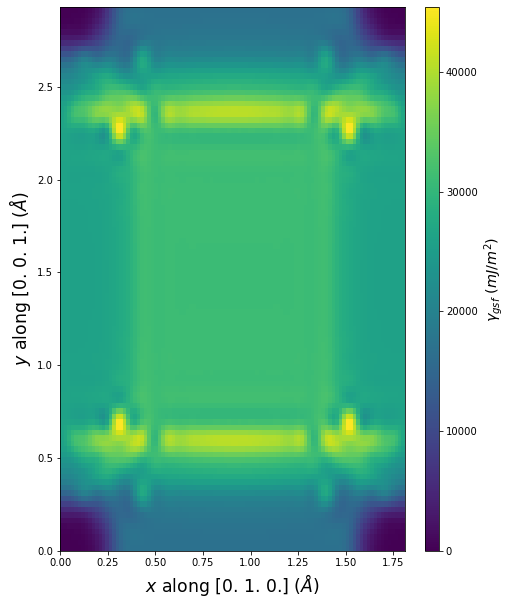
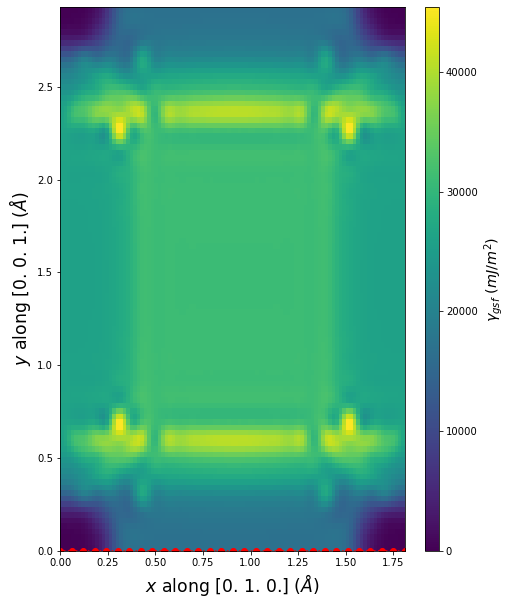
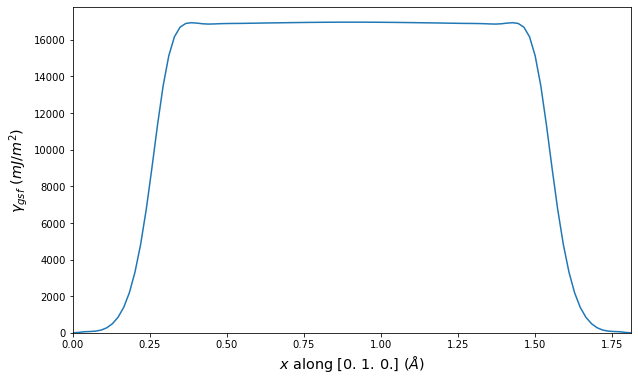
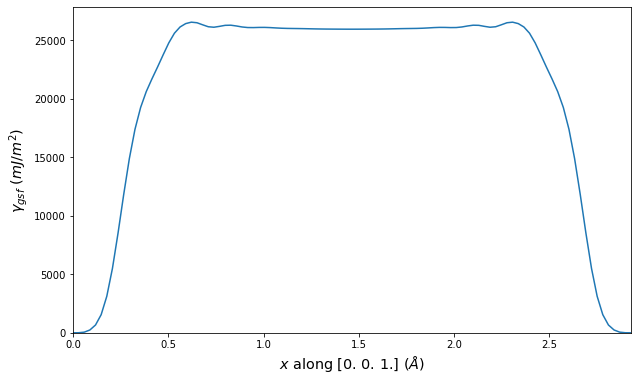
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-1 2 -1 0] | 11467.48 |
| E_usf a/3 [-2 1 1 3] | 15862.40 |
| τ_ideal a/3 [-1 2 -1 0] | 732.39 |
| τ_ideal a/3 [-2 1 1 3] | 714.30 |
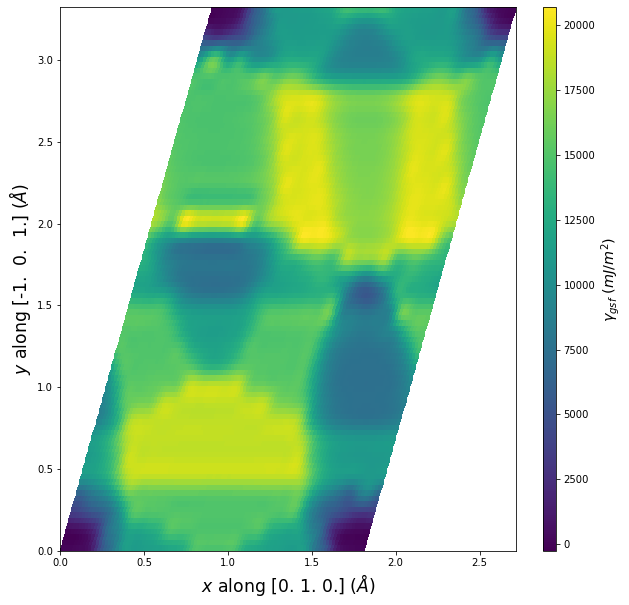
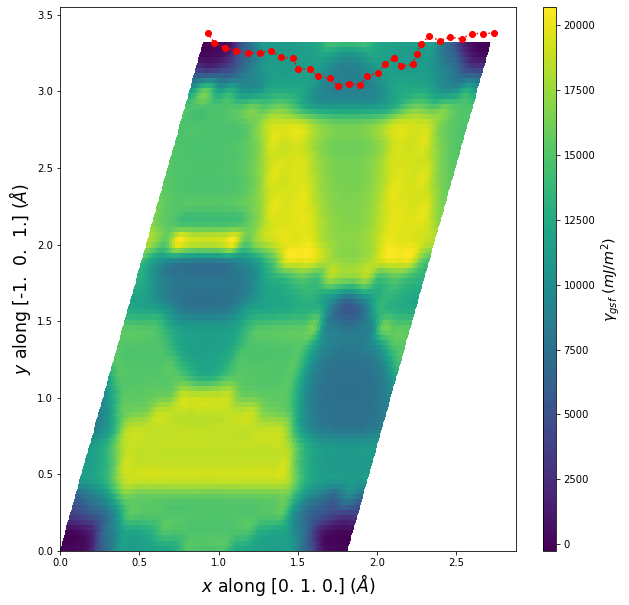
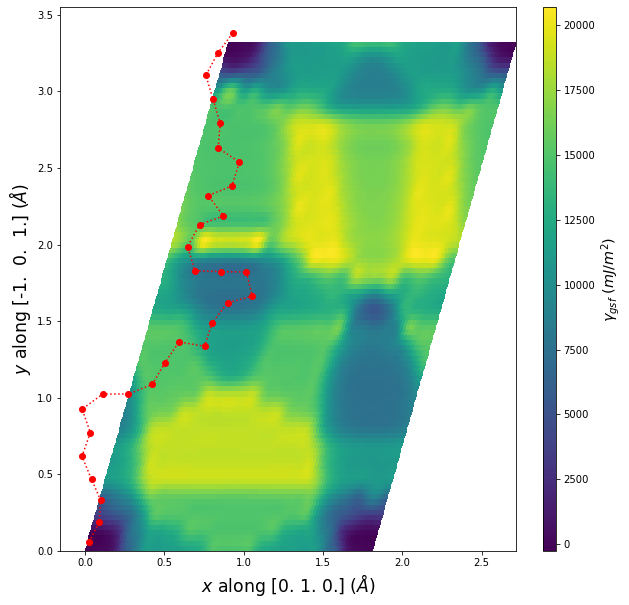
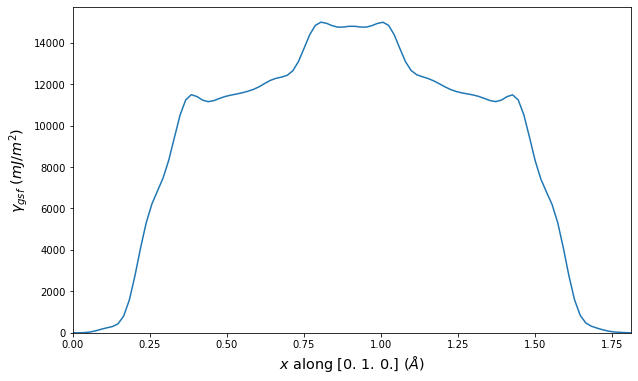
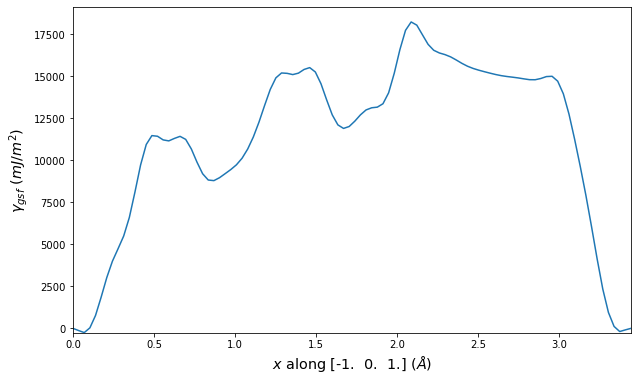
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-2 1 1 3] | 20391.87 |
| τ_ideal a/3 [-2 1 1 3] | 636.43 |
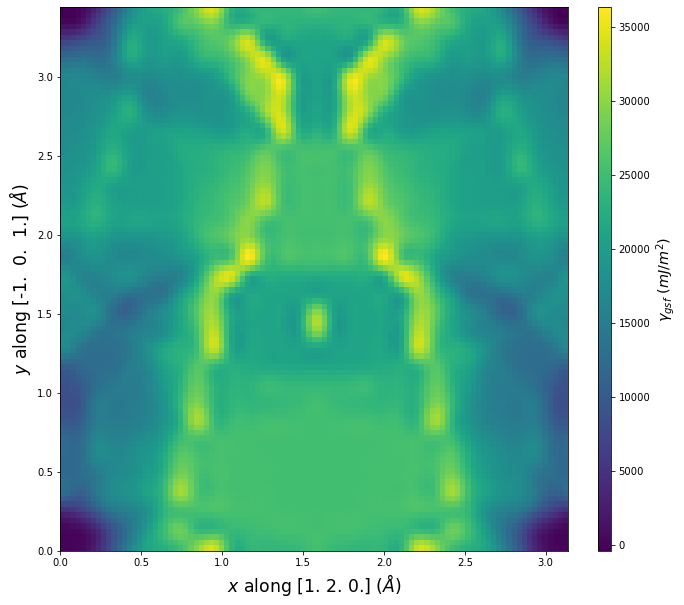
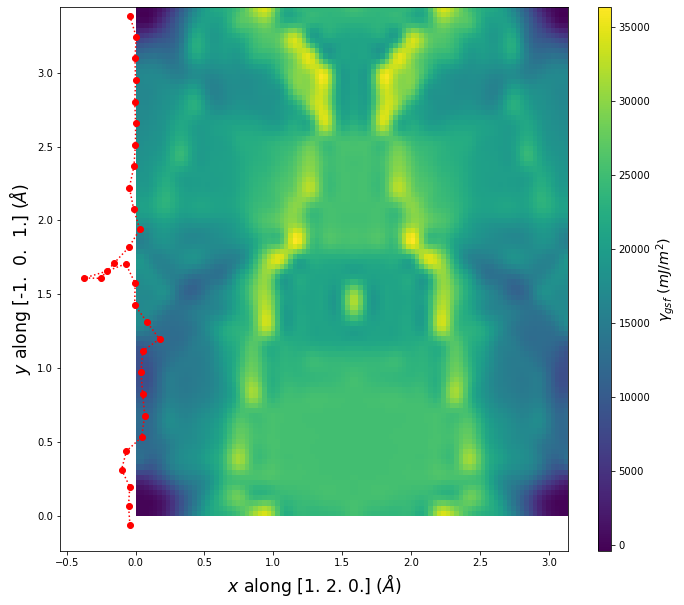
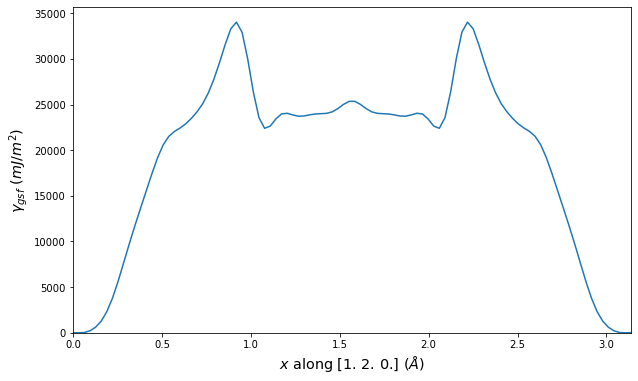
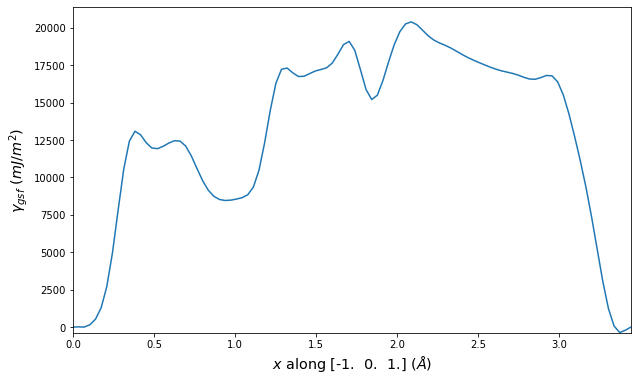
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/2 [0 -1 1] | 881.01 |
| τ_ideal a/2 [0 -1 1] | 1067.13 |
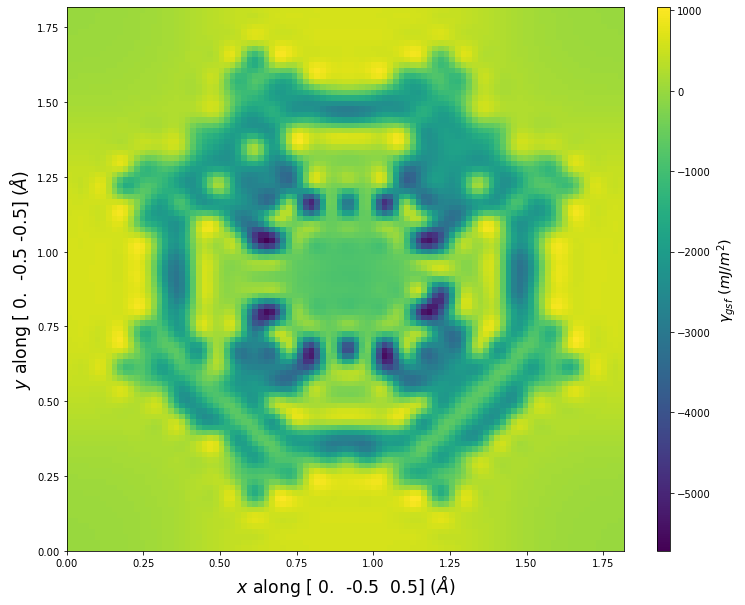

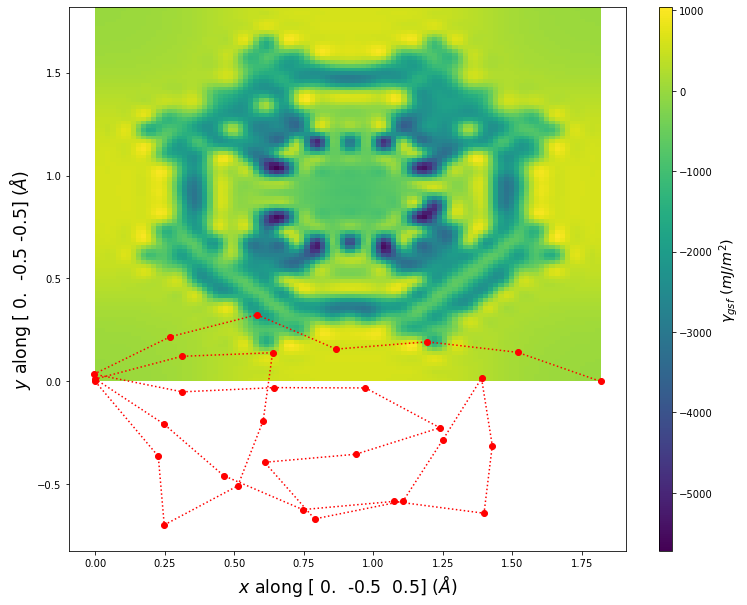
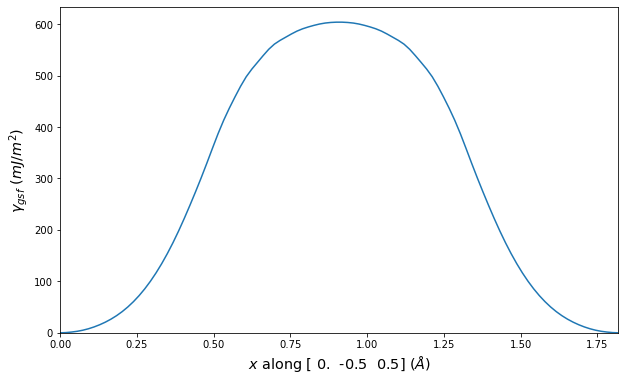
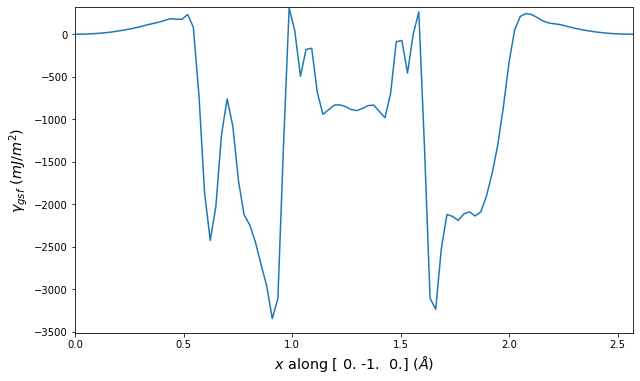
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_isf | -0.00 |
| E_usf a/2 [0 -1 1] | 67.56 |
| τ_ideal a/2 [0 -1 1] | 2.23 |


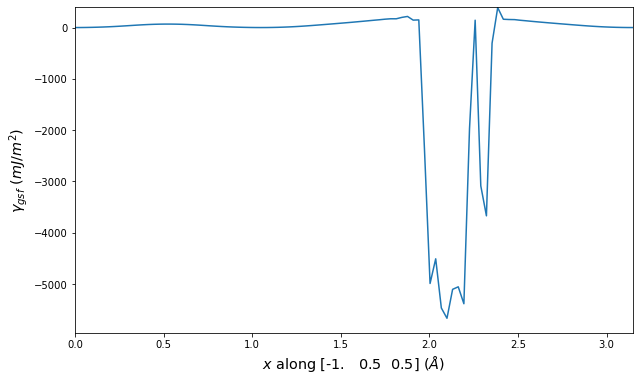
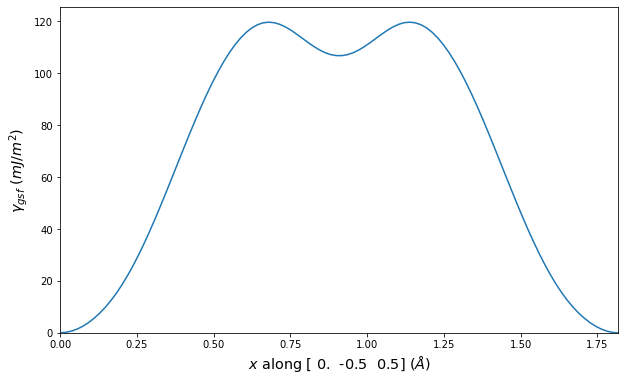
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/2 [-1 1 -1] | 365.67 |
| τ_ideal a/2 [-1 1 -1] | 6.69 |
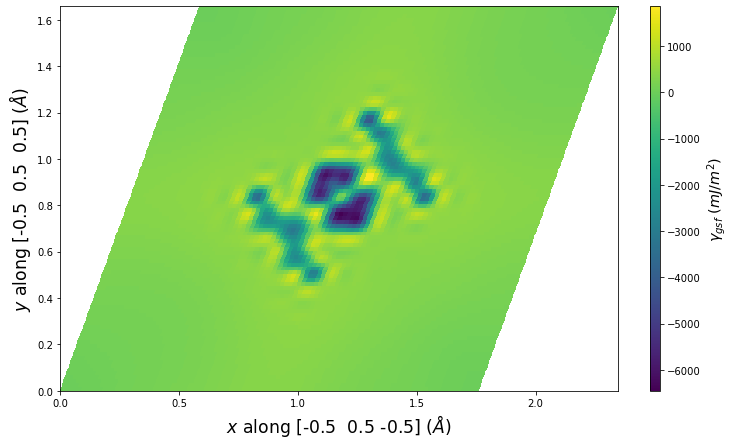

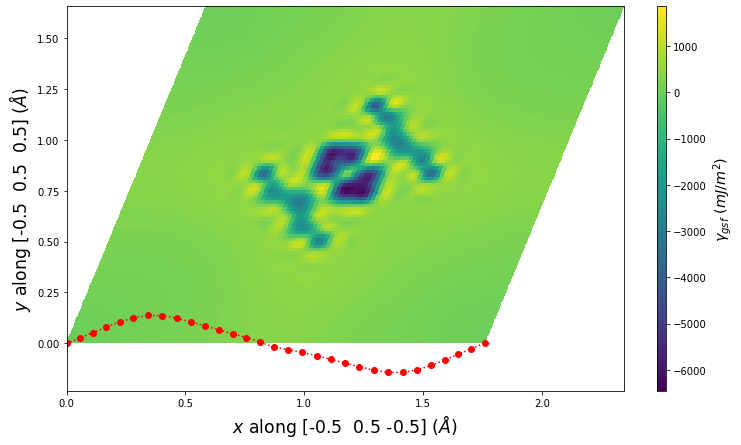
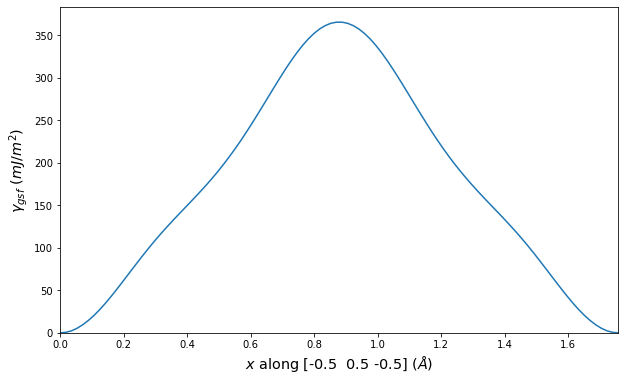
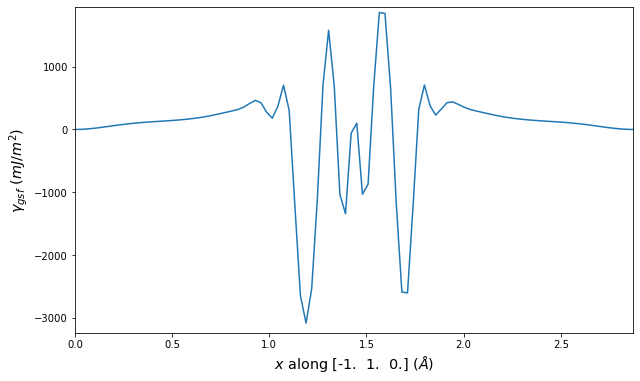
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/2 [-1 -1 1] | 423.43 |
| τ_ideal a/2 [-1 -1 1] | 7.15 |
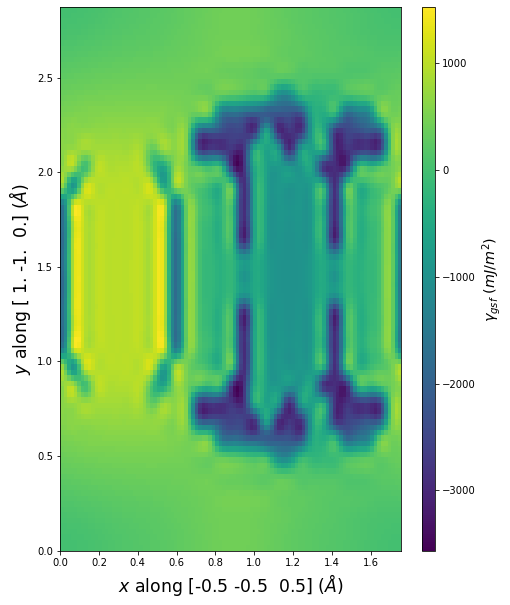
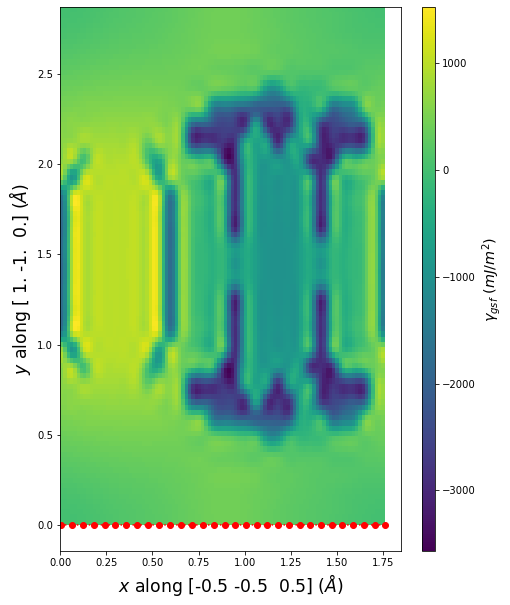
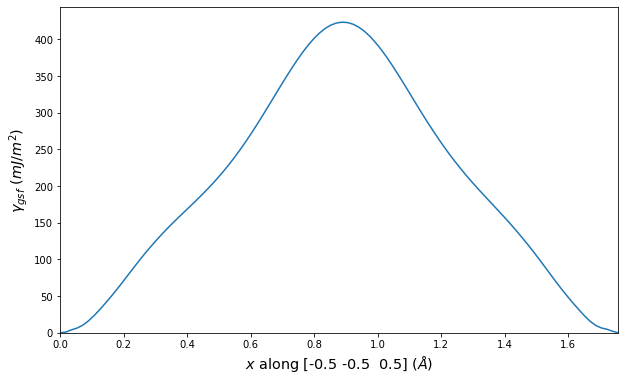
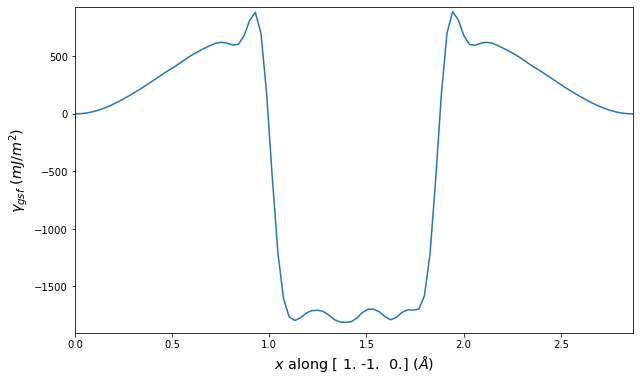
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_isf | 0.00 |
| E_usf a/3 [-2 1 1 0] | 80.51 |
| τ_ideal a/3 [-2 1 1 0] | 2.68 |
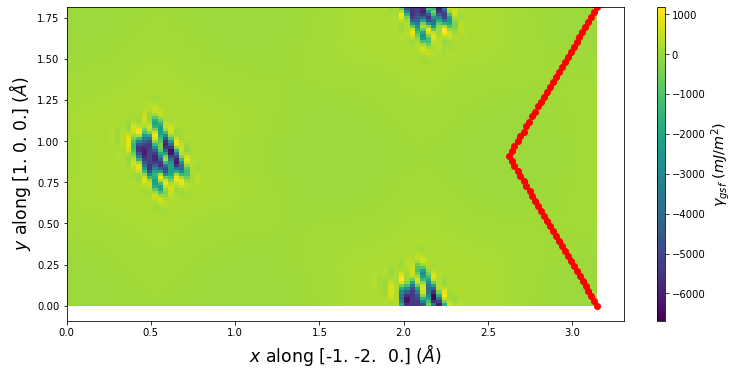
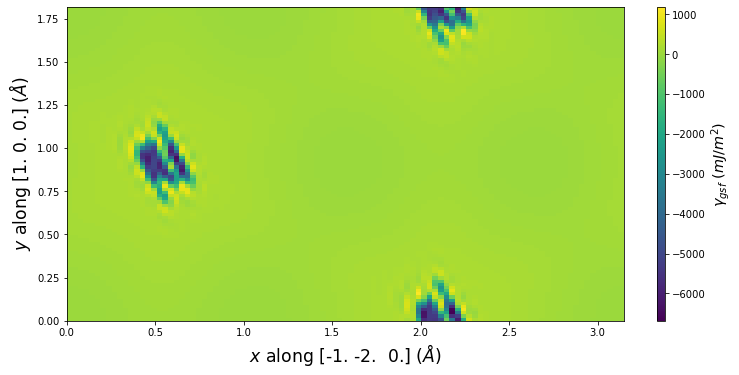

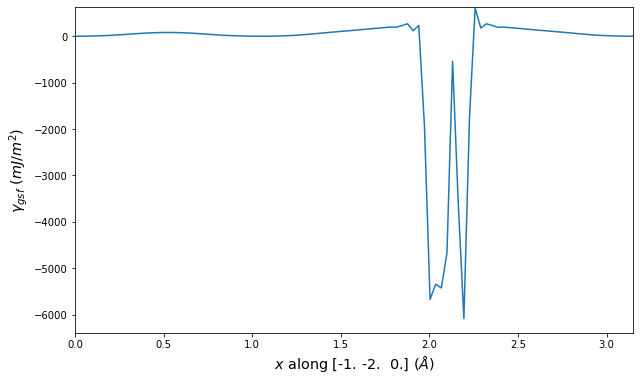
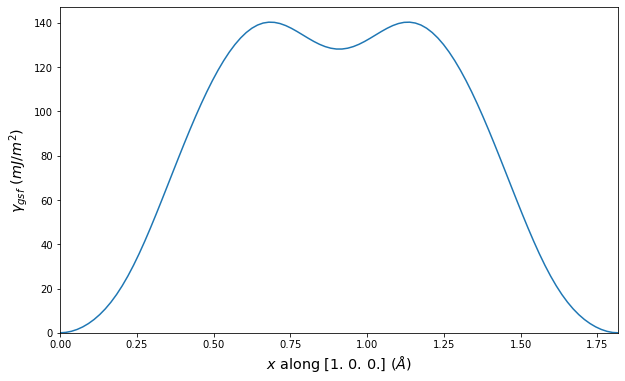
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-1 2 -1 0] | 116.92 |
| τ_ideal a/3 [-1 2 -1 0] | 3.36 |
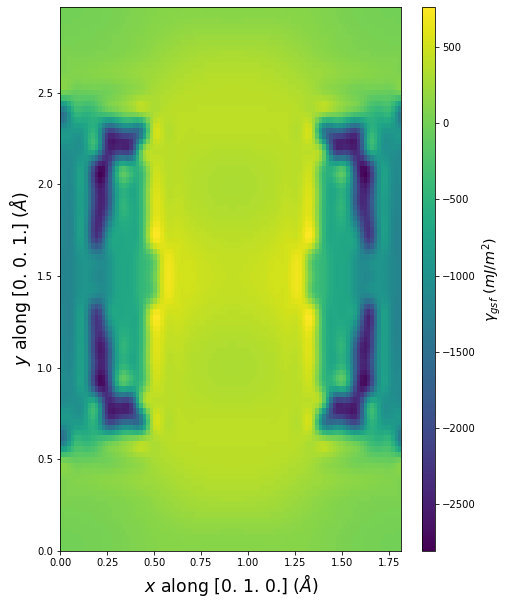
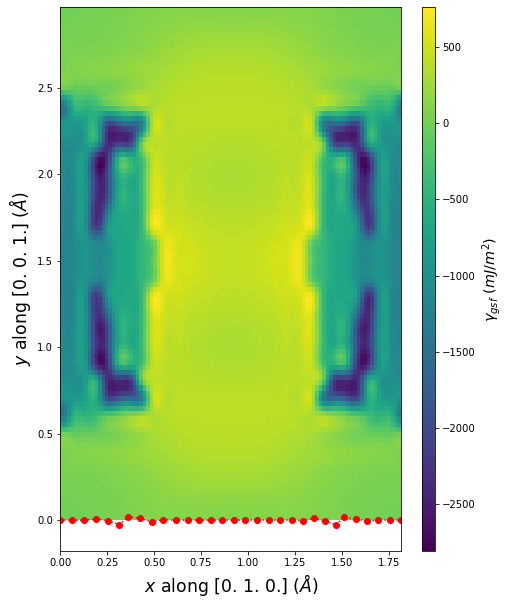
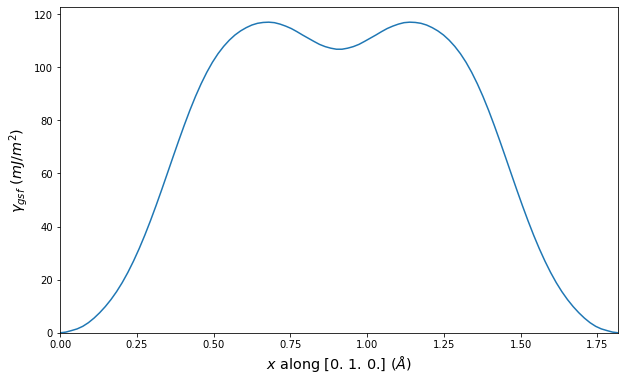
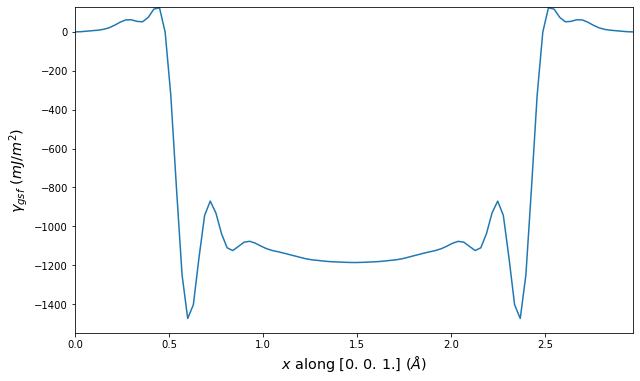
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-1 2 -1 0] | 832.34 |
| τ_ideal a/3 [-1 2 -1 0] | 18.82 |
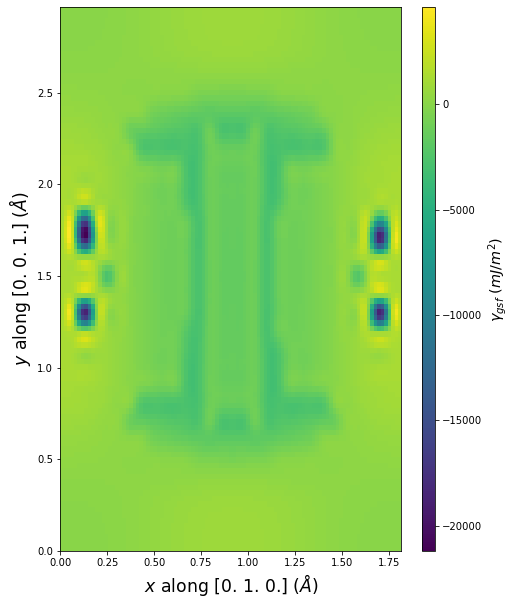

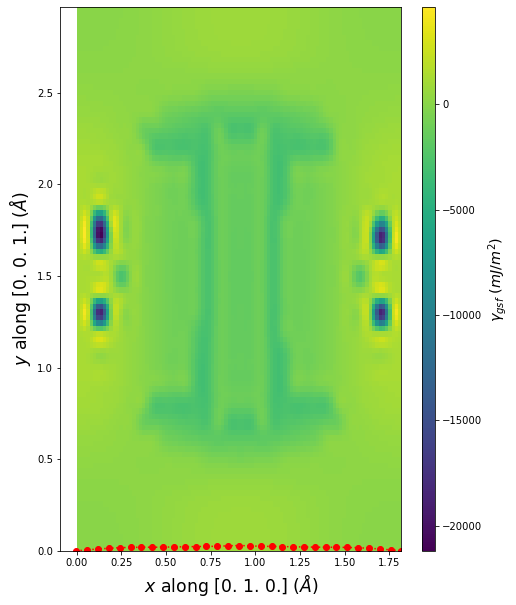
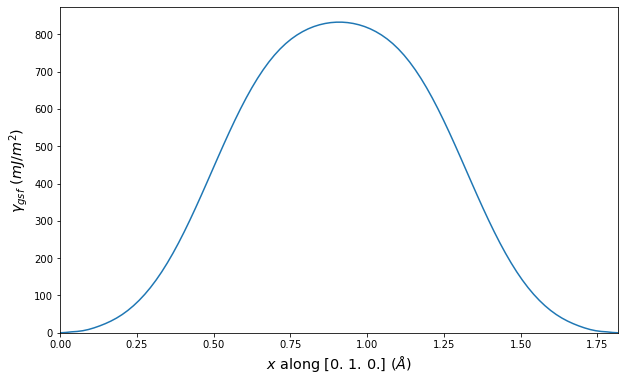
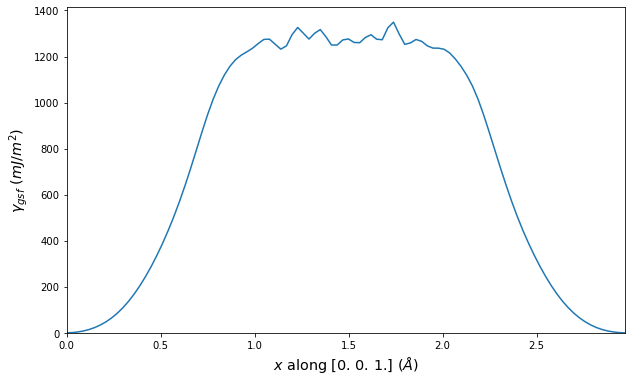
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-1 2 -1 0] | 177.75 |
| E_usf a/3 [-2 1 1 3] | 132.87 |
| τ_ideal a/3 [-1 2 -1 0] | 796.71 |
| τ_ideal a/3 [-2 1 1 3] | 259.29 |
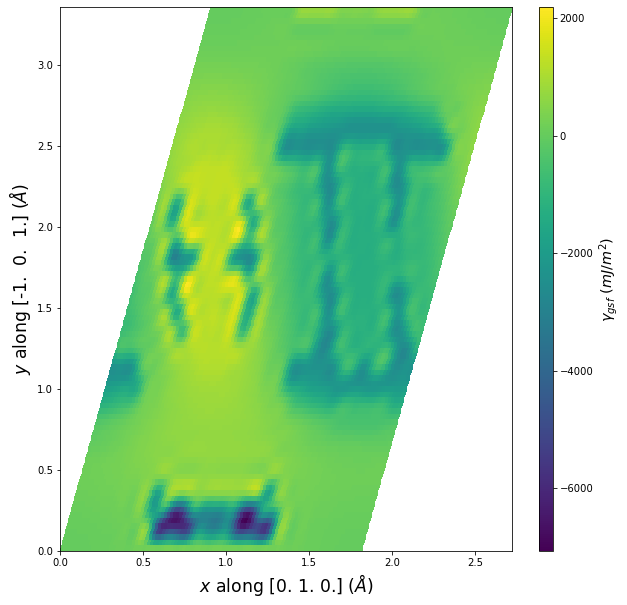

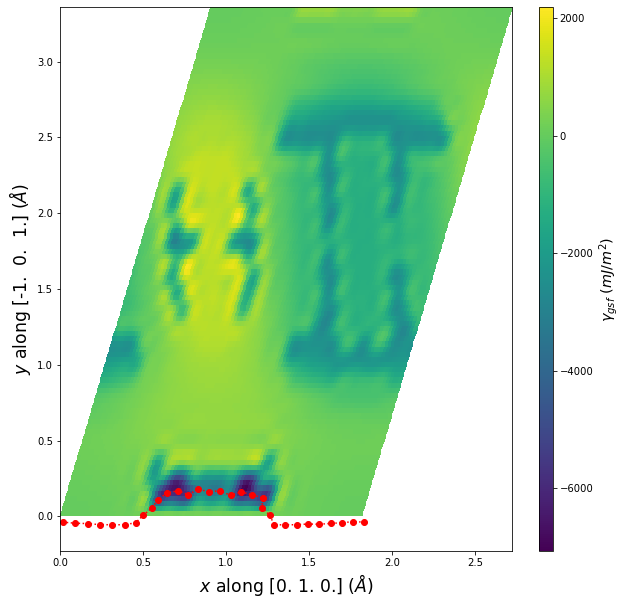
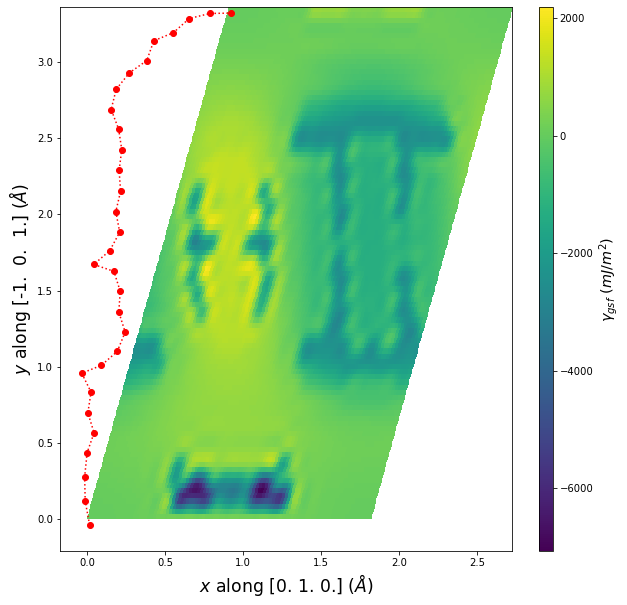
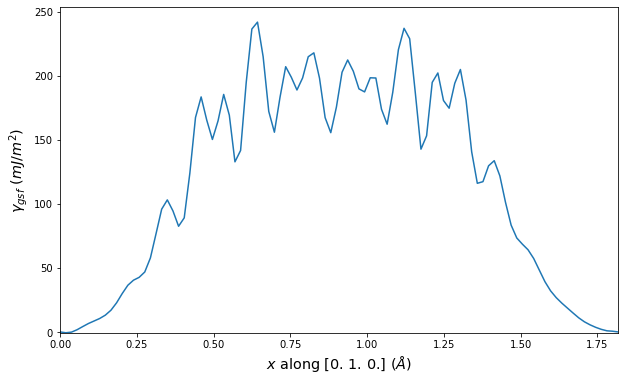
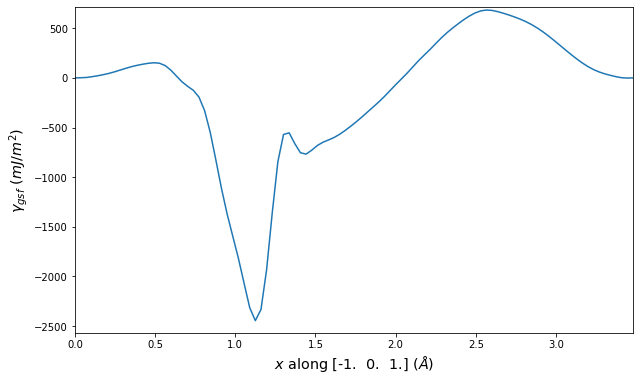
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-1 2 -1 0] | 601.80 |
| E_usf a/3 [-2 1 1 3] | 673.56 |
| τ_ideal a/3 [-1 2 -1 0] | 9729.15 |
| τ_ideal a/3 [-2 1 1 3] | 5078.01 |

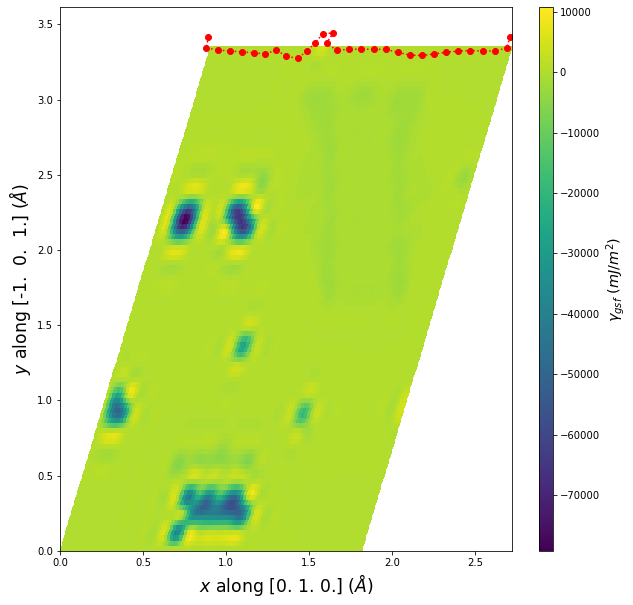
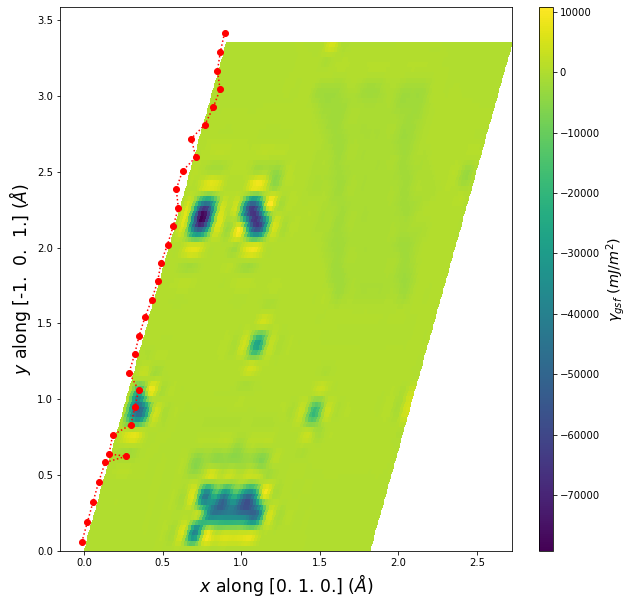
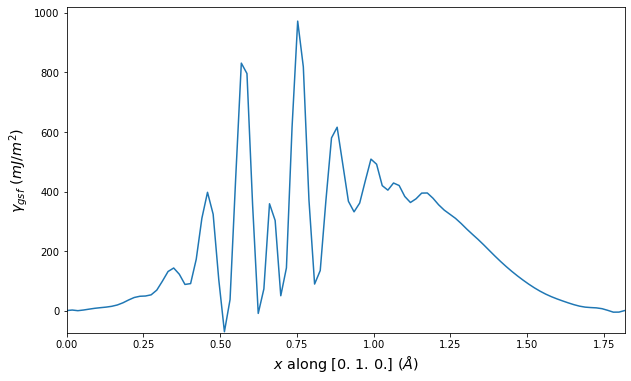
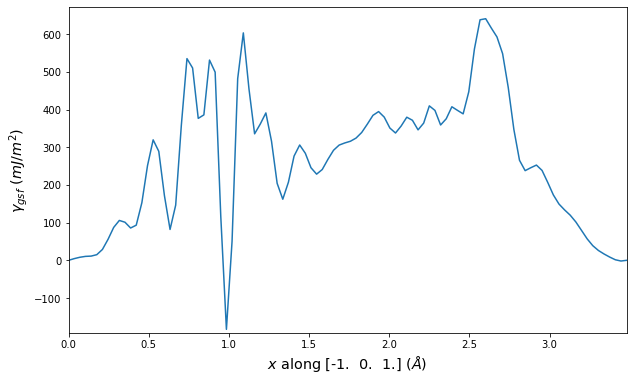
Stacking fault energies in mJ/m2 and ideal shears in GPa
| E_usf a/3 [-2 1 1 3] | 760.81 |
| τ_ideal a/3 [-2 1 1 3] | 639.61 |
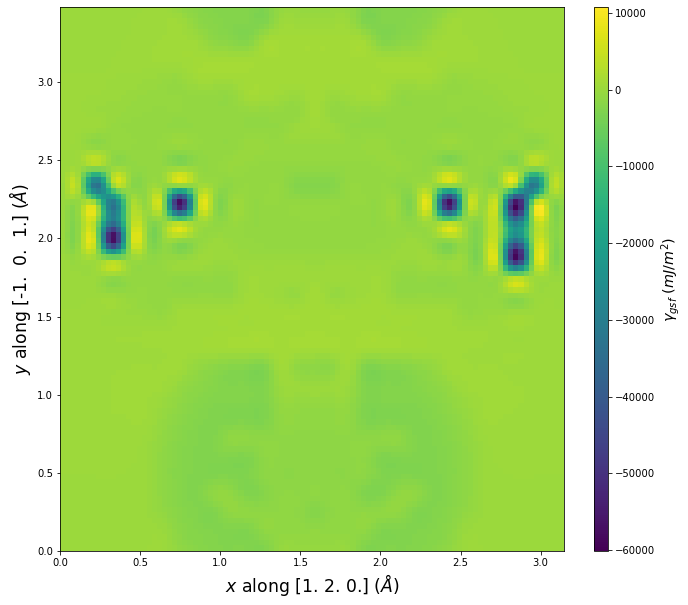

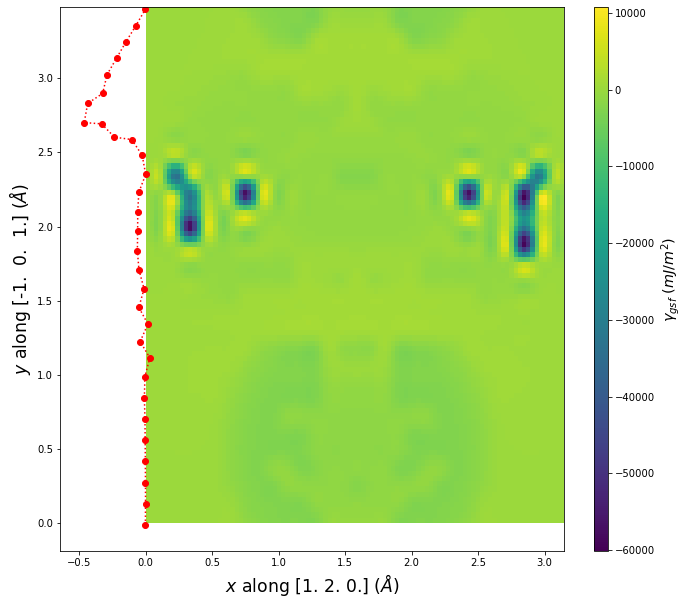
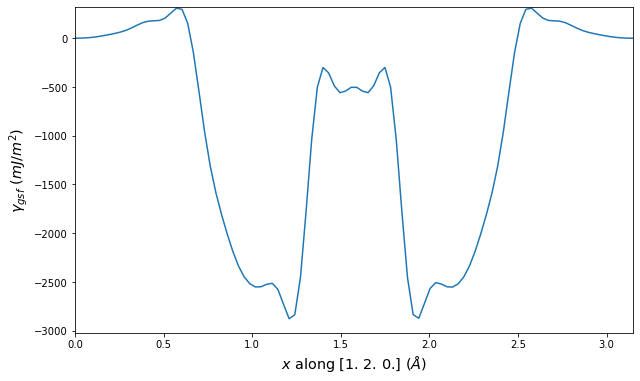
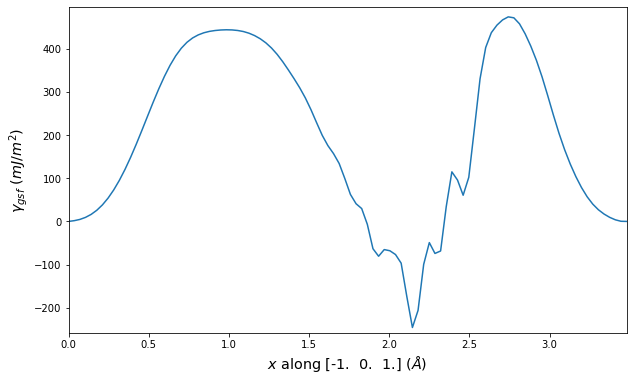
Dislocation Monopole Structures
Plots of the atomic structure of dislocation cores are shown here. The dislocations were constructed using dislocation monopole configurations in which a straight dislocation is created in the center of the system using anisotropic elasticity and the elastic constants computed above. A cylindrical region centered around the dislocation is allowed to relax while the remaining atoms are held fixed. The relaxation consists of 1 million NVT steps at 10 K followed by an energy/force minimization. The final plots show the atomic configuration near the center of the monopole system, and display differential displacement arrows and Nye tensor color maps that are both taken with respect to the Burgers vector direction.
These plots show a wide range of final structures, some corresponding to expected dislocation structures and others not. While the characterizations are dislocation-type dependent, some general classification guidelines are listed here.
- Planar dislocations exist on a single slip plane, which results in a grouping of large differential displacement arrows between two atomic rows. All edge dislocations and some screw dislocations may be planar dislocations.
- Non-planar dislocations are screw dislocations that are positioned at the intersection of more than one slip plane. This results in the differential displacement arrows circling the position of the dislocation core. These core structures often have rotational symmetry due to the intersecting planes.
- The plotted atomic positions are also useful for characterizing the final configuration. For screw dislocations, the atomic projections should match projections in the bulk crystal if the crystal has not transformed. For edge dislocations, the projections should be the bulk projections with the extra half-plane of atoms directly above the dislocation core.
- Images in which the atomic positions look correct but no large differential displacement arrows are observed are likely the result of the dislocation gliding out of view during relaxation.
- Phase transformations are identified by large differential displacement arrows spread out in both plot directions, often across the whole image. Additionally, the atomic position projections will likely not correspond to what is expected.
- Twins typically appear as a long line of large differential displacement arrows along a single plane. Note that twins can nucleate phase transformations so the distinction between the two is not always clear.
The calculation method used is available as the iprPy dislocation_monopole calculation method.
Links are provided above the images allowing for users to download the final defect configuration and a corresponding defect-free configuration if they wish to further analyze the structures or use them for subsequent simulations.
Notes and Disclaimers:
- While the general dislocation monopole method is universal to all dislocations, these calculations rely on dislocation-type specific atomic configuration sizes and differential displacement/Nye tensor settings. For the system size, the specific super cell multipliers vary based on the crystal and crystal vector orientations, but the sizes are always selected to minimize the length along the dislocation line and attempt to make the two other box dimensions similar. For the structure analysis tools, both are sensitive to the cutoff distance used to identify neighbors, and both have some method-specific parameters. Where known, the settings used here correspond to the most common ones used in literature.
- For bcc dislocations, the differential displacement vectors shown here are "wrapped" in that if their magnitudes exceed b/2 then a value of b is added/subtracted to wrap the vector back around. This convention reveals the dislocation's structure and location without showing the slip history.
- NIST disclaimer
Version Information:
- 2025-08-07. Plots first added.
MD Solid Property Predictions
Plots of lattice and elastic constants are shown as a function of temperature. The 0K points were taken from the Crystal Structure Predictions and the Elastic Constants Predictions sections above for the unique crystal structures relaxed with the "dynamic" method. Starting from the 0 K relaxed crystal unit cells, supercell systems are created by replicating all three dimensions by the same multiplier to achieve at least 4000 atoms. The systems are then relaxed at 50 K and zero pressure using 1 million NPT steps. Lattice constants are estimated by averaging the measured box dimensions. Temperatures are iteratively increased by 50 K, with each subsequent relaxation calculation starting from the final atomic configuration at the previous temperature and relaxing for another 1 million steps.
The elastic constants are calculated using the deformation–fluctuation hybrid method. Starting from the final atomic configurations of the dynamic relaxations, the system is allowed to evolve at constant volume with a Langevin thermostat. The Born matrix is computed during this run by evaluating how the atomic forces would vary due to applied linear strain fields. The elastic constants can then be estimated using the averaged Born matrix values and the averaged stresses on the system.
The calculation methods used are available as the iprPy relax_dynamic and elastic_constants_dynamic calculation methods.
Clicking on the image of a plot will open an interactive version of it in a new tab. The underlying data for the plots can be downloaded by clicking on the links above each plot.
Notes and Disclaimers:
- The maximum temperature shown for a crystal either corresponds to the maximum value that has been computed so far, or the maximum value that the structure is believed to have remained untransformed. The unstable/transformation temperature identification is mostly automated and can miss transitions not associated with large discontinuities in property values with changing temperature.
- For succinctness, the elastic constants displayed here are averaged according to the 0 K structure's crystal symmetry family. If the structures deviate from the expected symmetry at higher temperatures, the values may not be valid. Raw results are available for verification if requested.
- NIST disclaimer
Version Information:
- 2025-08-07. Plots added.
MD Liquid Property Predictions
Plots of radial distribution functions at different temperatures and the temperature-dependent diffusion and viscosity are shown here for elemental liquids. Melt phases are constructed by starting with a 10x10x10 fcc super cell and relaxing with 50,000 NPT steps at zero pressure and a high melting temperature (3000 K for most potentials and elements). Following the melt, liquid structures are relaxed by running NPT for 10,000 steps over which the temperature is dropped by 50 K, then 50,000 NPT at the target temperature to estimate the equilibrium volume, followed by 20,000 NVT steps at the averaged equilibrium volume and target temperature. Structures from the NVT run are sampled to compute the radial distribution function of the liquid at the corresponding temperature.
The final relaxed configurations at each temperature explored are used as the initial configurations for the diffusion and viscosity calculations. For diffusion, the system is integrated for 100 runs of 2,000 NVT steps during which both the mean squared displacement (MSD) and the velocity auto correlation function (VACF) are computed. From these simulations, three estimates of diffusion are computed: one from the MSD of the full 200,000 step run, one in which the MSD is reset for each short run and then averaged, and one from the VACF computed for each short run and averaged. For viscosity, the Green-Kubo method is used which is evaluated during a 1 million step NVT run.
The calculation methods used are available as the iprPy relax_liquid_redo, diffusion_liquid, and viscosity_green_kubo calculation methods.
Clicking on the image of a plot will open an interactive version of it in a new tab. The underlying data for the plots can be downloaded by clicking on the links above each plot.
Notes and Disclaimers:
- The current liquid relaxation has some known issues, and therefore may be recalculated with an improved method later. Most notably, the unconstrained NPT relaxation may lead to one of the periodic dimensions becoming quite small which can negatively affect the simulations and property calculations. Additionally, the relaxations likely need to be longer to reduce the error associated with the measured volume and energy values.
- Values are only displayed down to the last temperature currently calculated, or the last temperature at which the system is believed to remain liquid until the end of all simulations. The transformation temperature is estimated by looking at discontinuities in the measured property values and may incorrectly predict a phase to still be liquid if the discontinuities are small.
- The three diffusion estimates provide a range of prediction values. For MSD, the longer the run the more likely it is that a small number of atoms will diffuse much farther than the rest, which skews the squared displacement higher.
- NIST disclaimer
Version Information:
- 2025-08-07. Plots added. Data should be thought of as preliminary as results may be recalculated later.
MD Thermodynamic Predictions
Plots of internal energy, Gibbs free energy, entropy, heat capacity and volume are shown here as a function of temperature for various crystal structures and liquid phases. The included crystal structures correspond to those in the Solid Structures vs. Temperature section, and the liquid phases to those in the Liquid Properties section. Internal energy and volume are taken from the associated structure relaxations mentioned in those previous sections. Constant pressure heat capacity is estimated using 3-point numerical derivatives of enthalpy versus temperature. Note that since all simulations done here are at 0 pressure, internal energy and enthalpy are equivalent.
The Gibbs energies of the phases are estimated using thermodynamic integration between the interatomic potential in question and a simpler model with known Gibbs free energy values. For solids, the reference model is an Einstein solid, while for liquids it is the Uhlenbeck-Ford potential. Besides a short run at the start of the solid calculations to estimate Einstein model spring constants, the two calculations proceed similarly. Starting with the final relaxation configurations, the systems are stabilized for 25,000 steps. Then, over the next 50,000 steps the potential is swapped out for the reference potential. The system is then stabilized for another 25,000 steps with the reference model before a reverse swap of 50,000 steps is performed. The simulation ends with one final 25,000 step stabilization period. From the simulation, the irreversible work of transformation is estimated and used to compute the absolute Gibbs free energy of the target phase and potential relative to the reference potential. Entropy is estimated as the difference in enthalpy and Gibbs free energy and divided by temperature.
The calculation methods used are available as the iprPy relax_dynamic, relax_liquid_redo, free_energy, and free_energy_liquid calculation methods.
Clicking on the image of a plot will open an interactive version of it in a new tab. The underlying data for the plots can be downloaded by clicking on the links above each plot.
Notes and Disclaimers:
- Good free energy estimates require that the initial and final phases before and after the thermodynamic integration remain the same. This is not an issue with the liquid calculations as the Uhlenbeck-Ford model always predicts a liquid, but may be an issue for the solid phase estimates. This is likely to be an issue for non-compact phases which the Einstein model is poor at representing.
- NIST disclaimer
Version Information:
- 2025-08-07. Plots added.